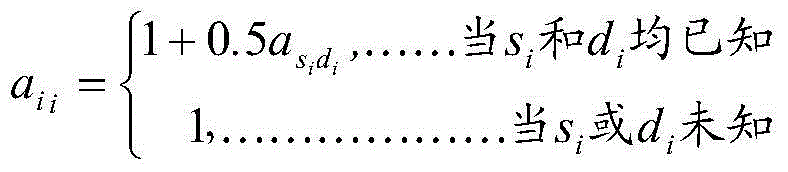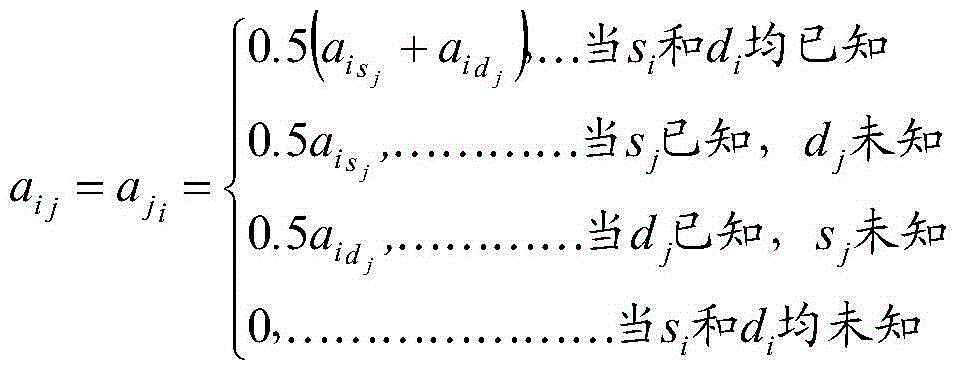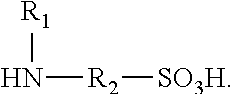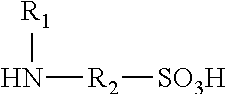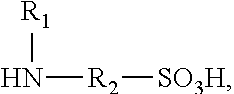Patents
Literature
Hiro is an intelligent assistant for R&D personnel, combined with Patent DNA, to facilitate innovative research.
61results about How to "Advantage" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Single crystal acoustic resonator and bulk acoustic wave filter
ActiveUS20160028367A1AdvantageSimple and cost-effectivePiezoelectric/electrostrictive device manufacture/assemblyPiezoelectric/electrostriction/magnetostriction machinesMetallic materialsCarbide
A method of wafer scale packaging acoustic resonator devices and an apparatus therefor. The method including providing a partially completed semiconductor substrate comprising a plurality of single crystal acoustic resonator devices provided on a silicon and carbide bearing material, each having a first electrode member, a second electrode member, and an overlying passivation material. At least one of the devices to be configured with an external connection, a repassivation material overlying the passivation material, an under metal material overlying the repassivation material. Copper pillar interconnect structures are then configured overlying the electrode members, and solder bump structures are form overlying the copper pillar interconnect structures.
Owner:AKOUSTIS INC
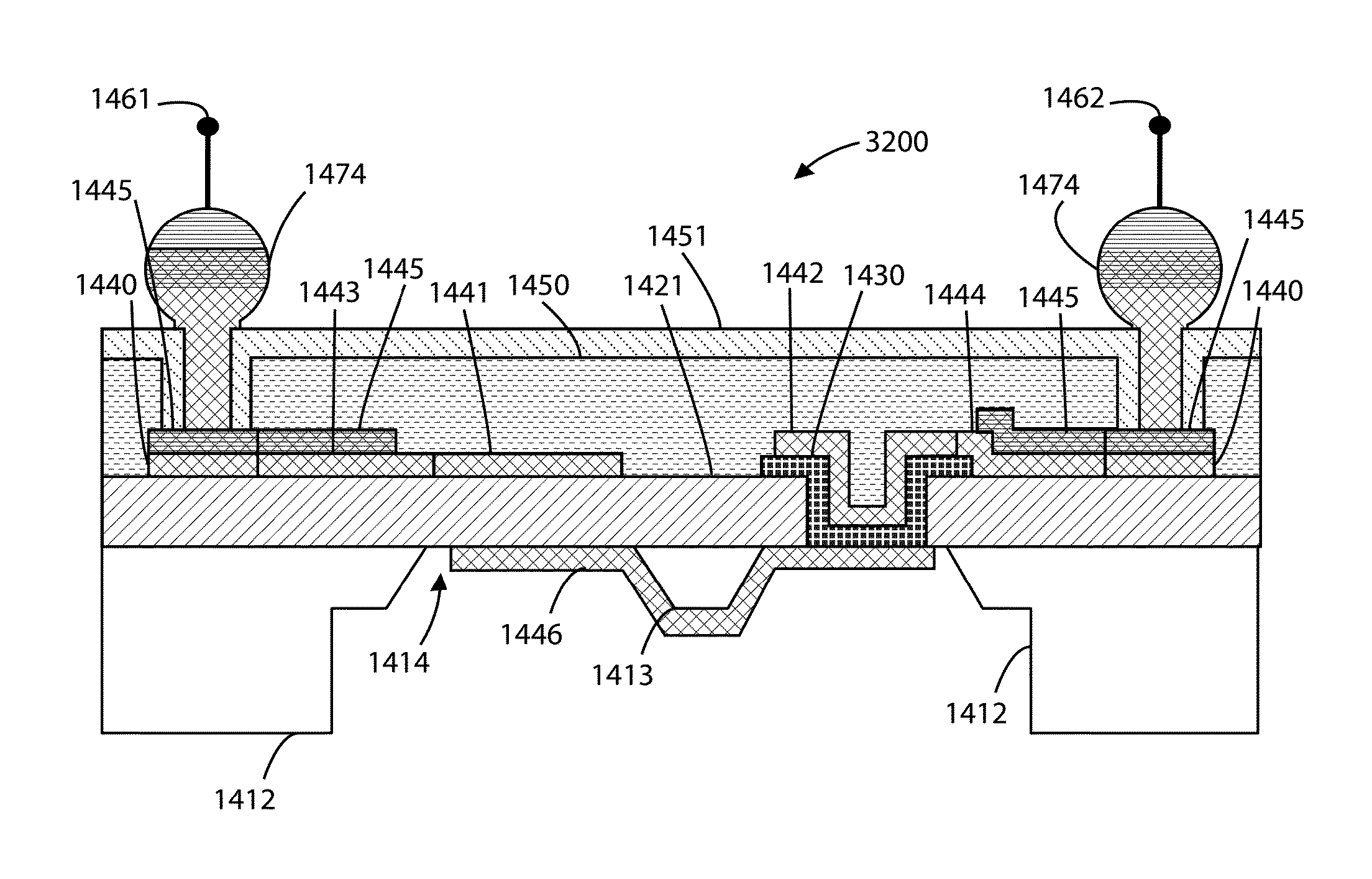
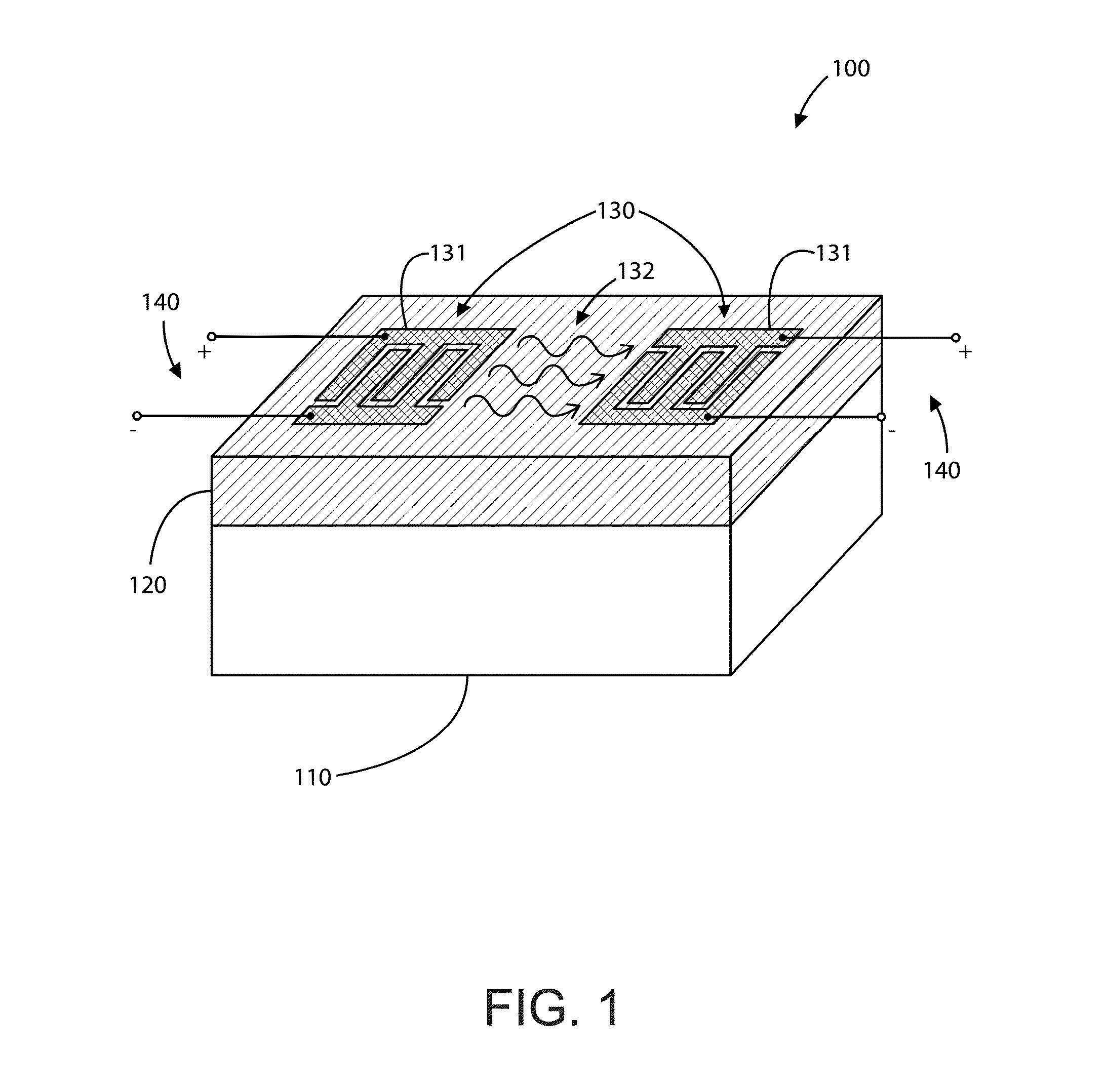
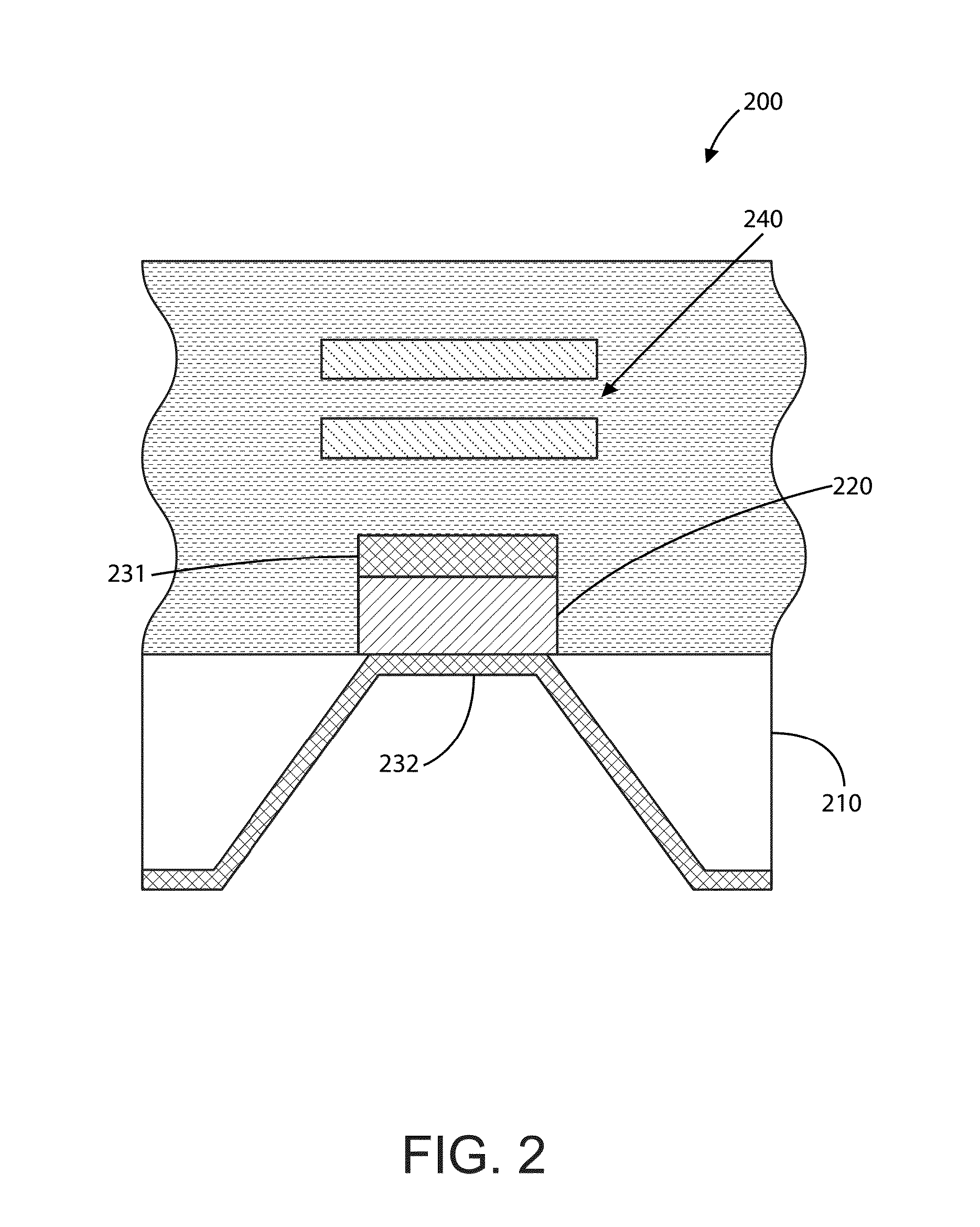
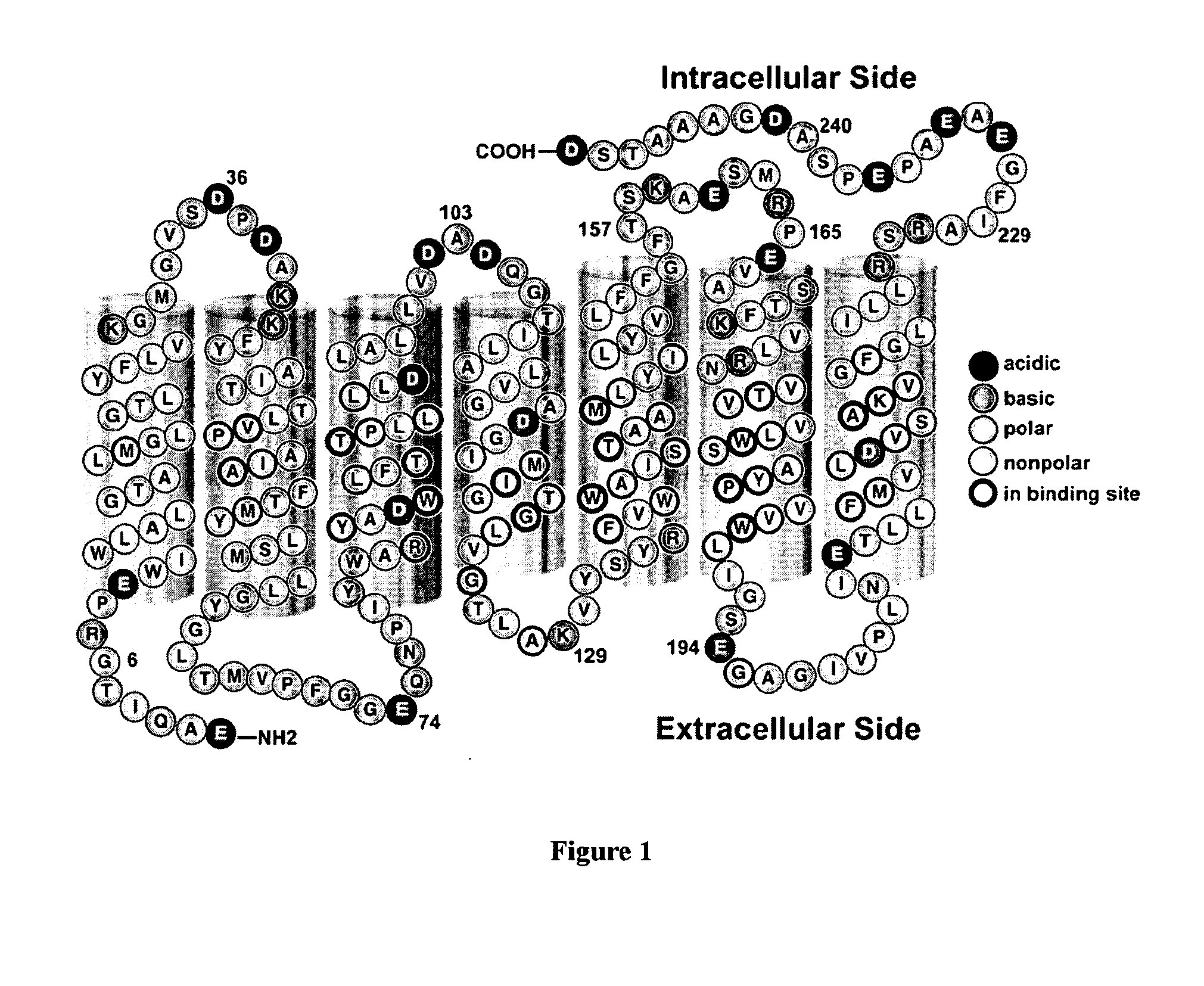
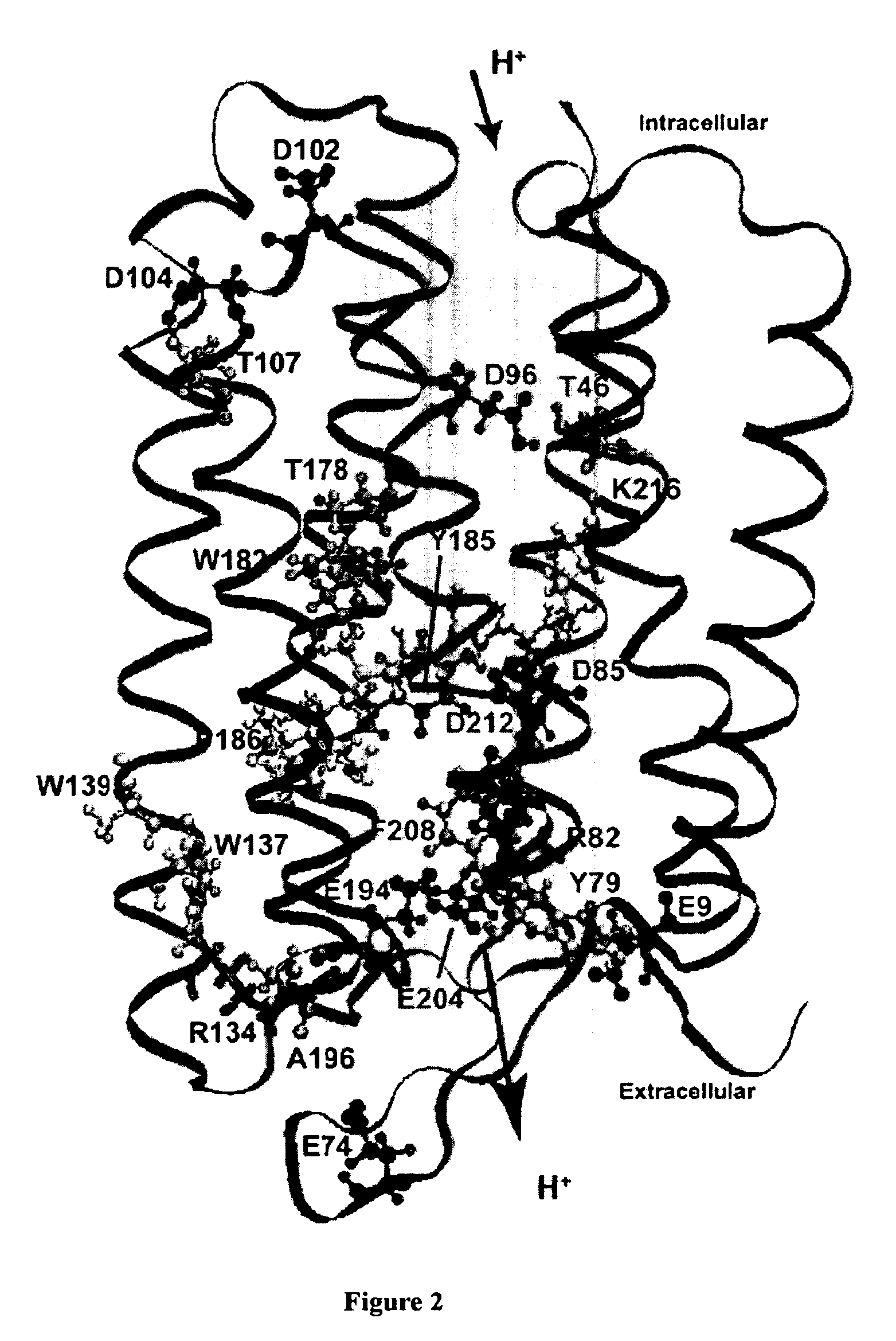
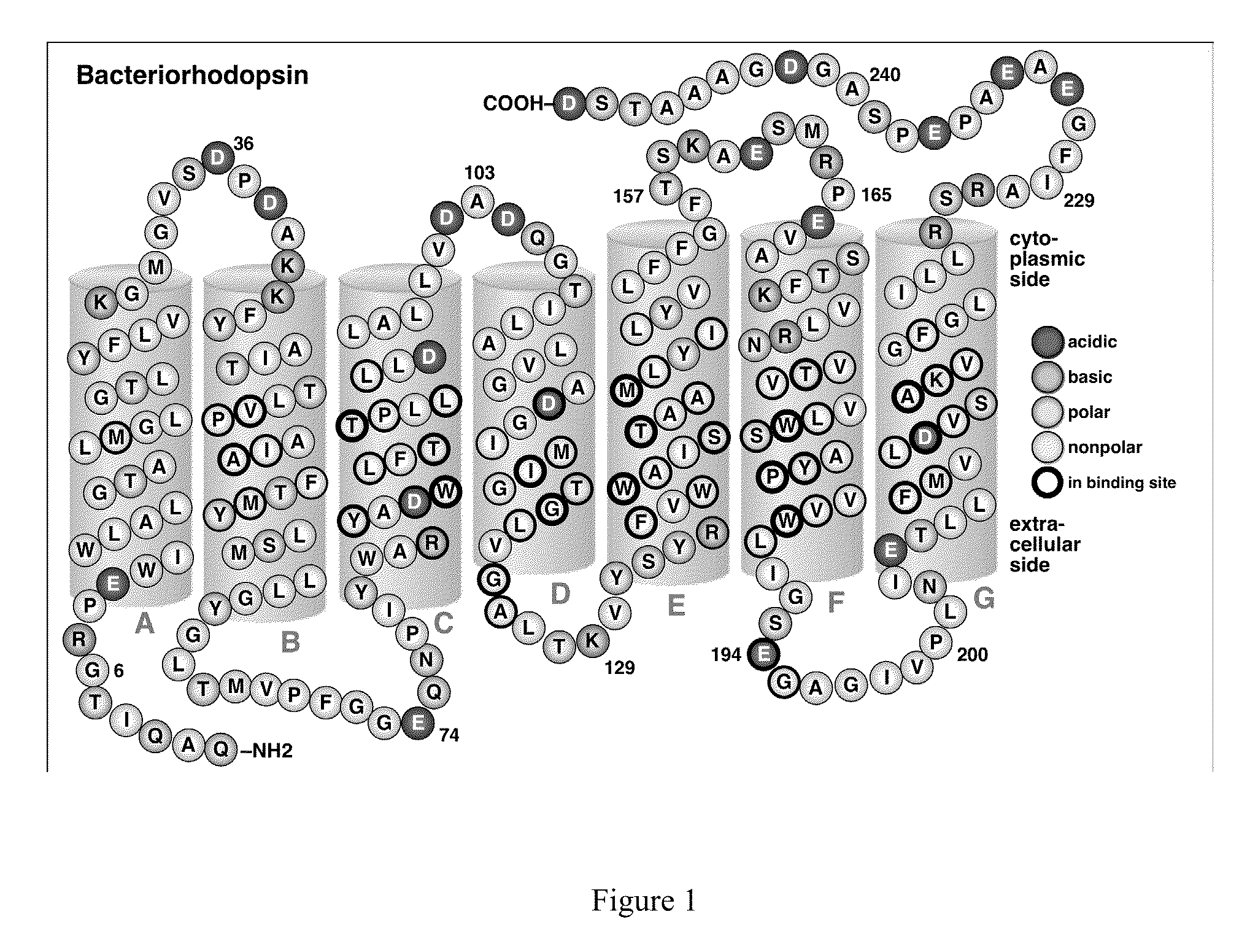
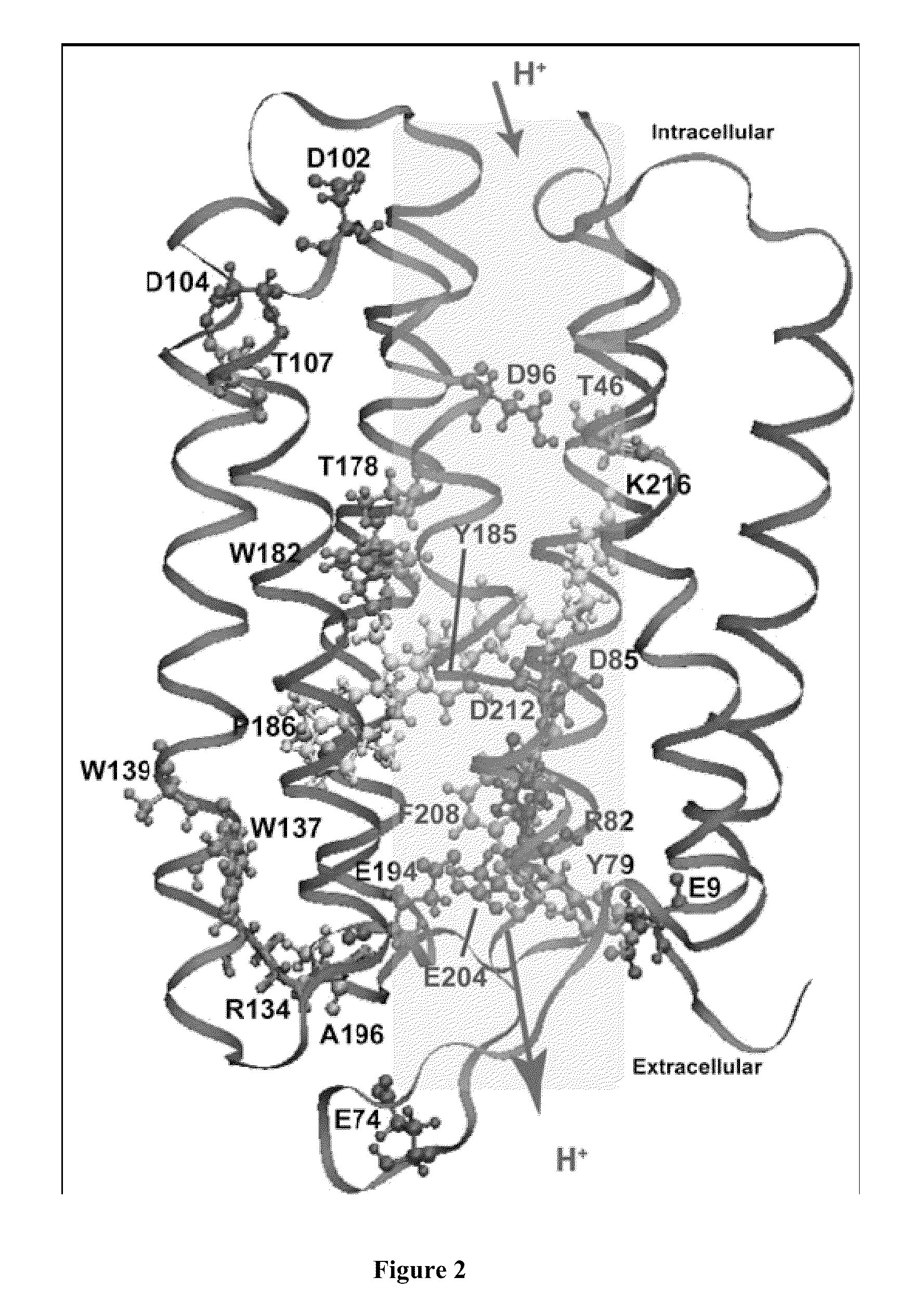
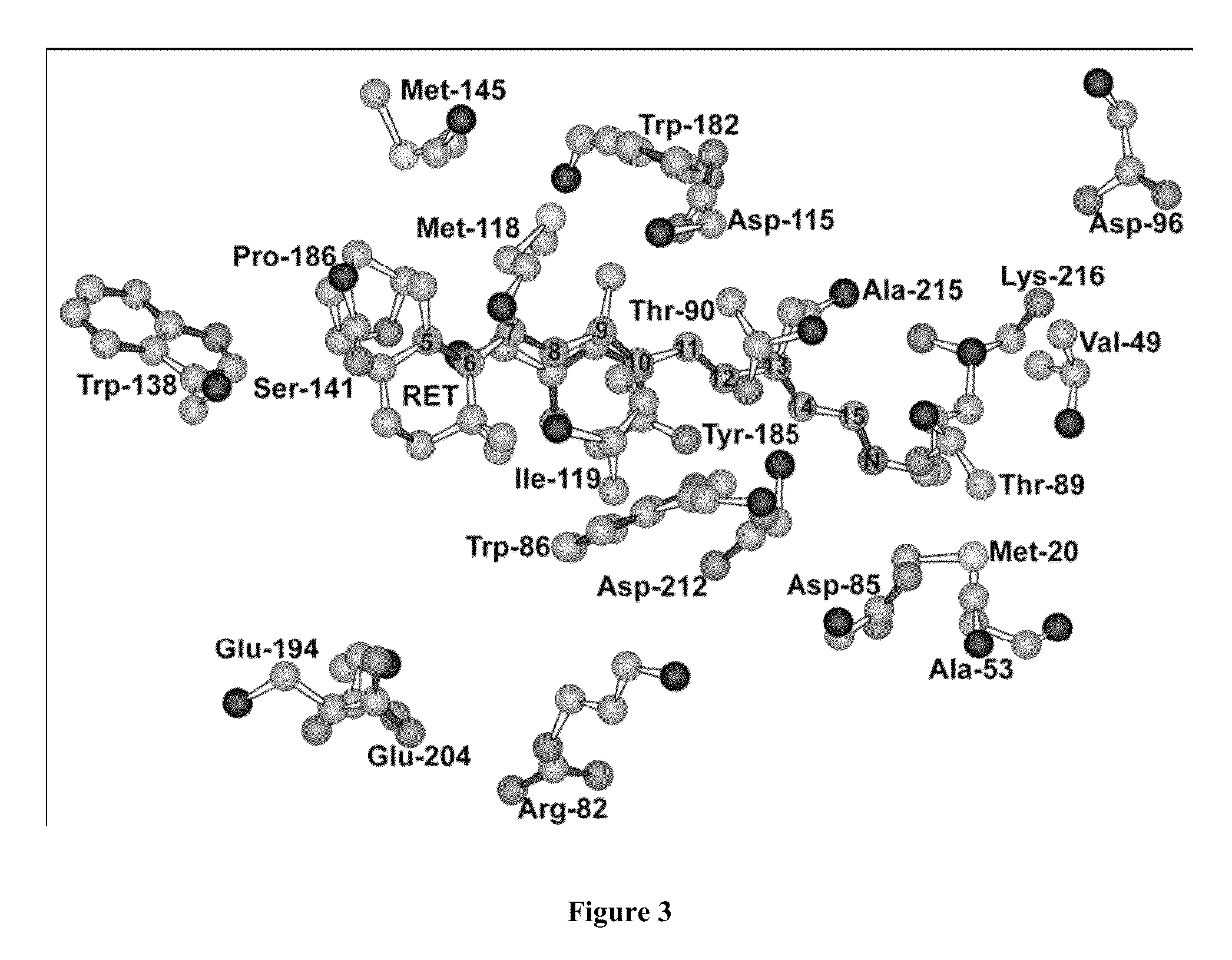
Bacteriorhodopsin Protein Variants and Methods of Use for Long Term Data Storage
ActiveUS20090268511A1High through-put storageEnhanced protein performanceNanoinformaticsPeptide preparation methodsLong term dataWild type
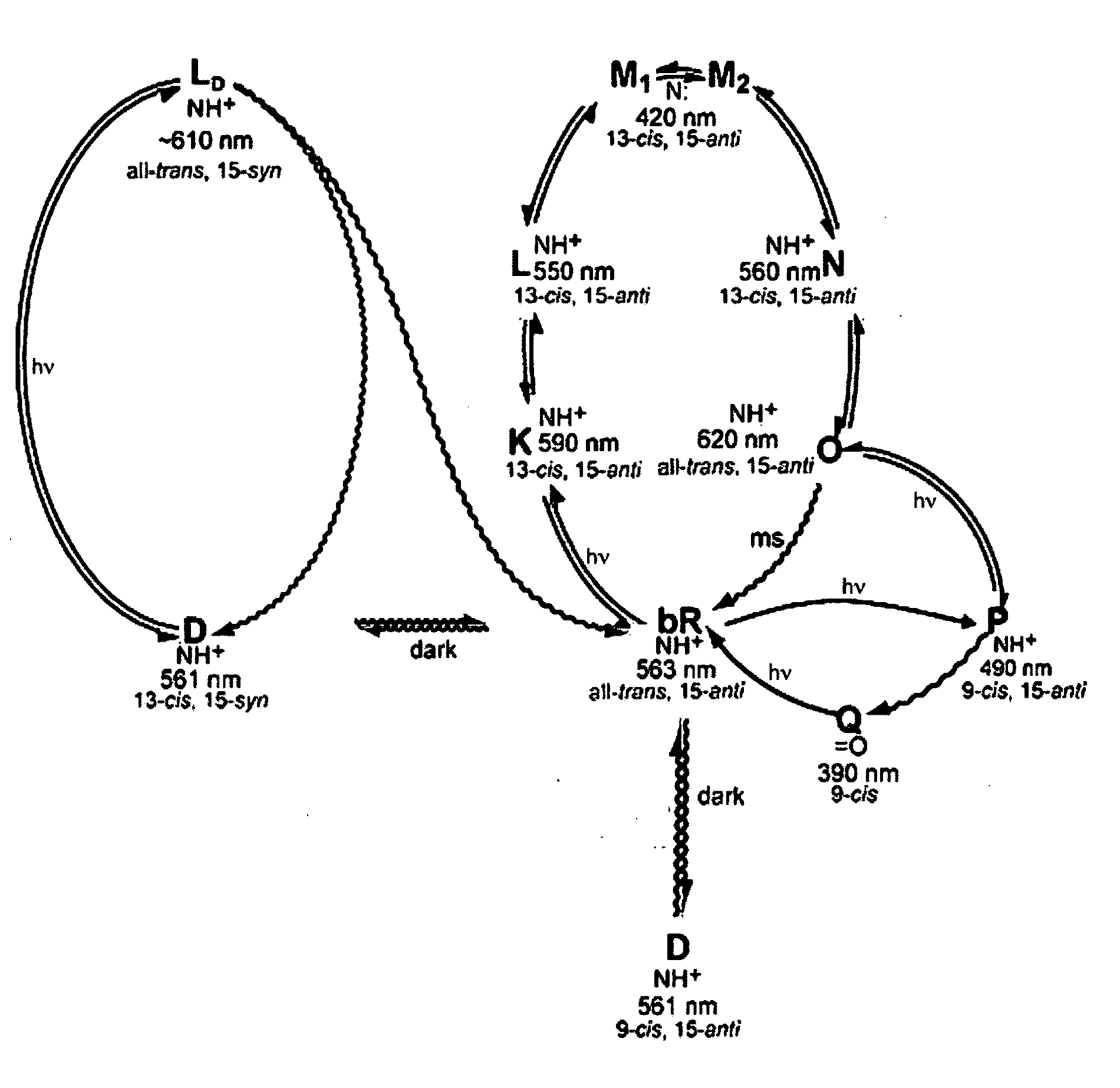
Bacteriorhodopsin protein variants and methods using the bacteriorhodopsin variants for performance in holographic and three-dimensional (3D) memory storage devices are described. The amino acid and chemical modifications of bacteriorhodopsin provided herein achieve greatly enhanced protein performance. The memory storage devices write, read and erase data proficiently. The bacteriorhodopsin protein variants are useful in optical memory storage and associative processor systems. Irradiation of the light-sensitive protein with light of known wavelength causes the protein to switch between different states. The variants enter the branched photocycle via a single or a two photon process and form the permanent ‘Q’ state more efficiently than the wild-type bacteriorhodopsin protein. This branching photocycle of the variants is exploited in the fabrication of 3D memory storage devices.
Owner:UNIV OF CONNECTICUT
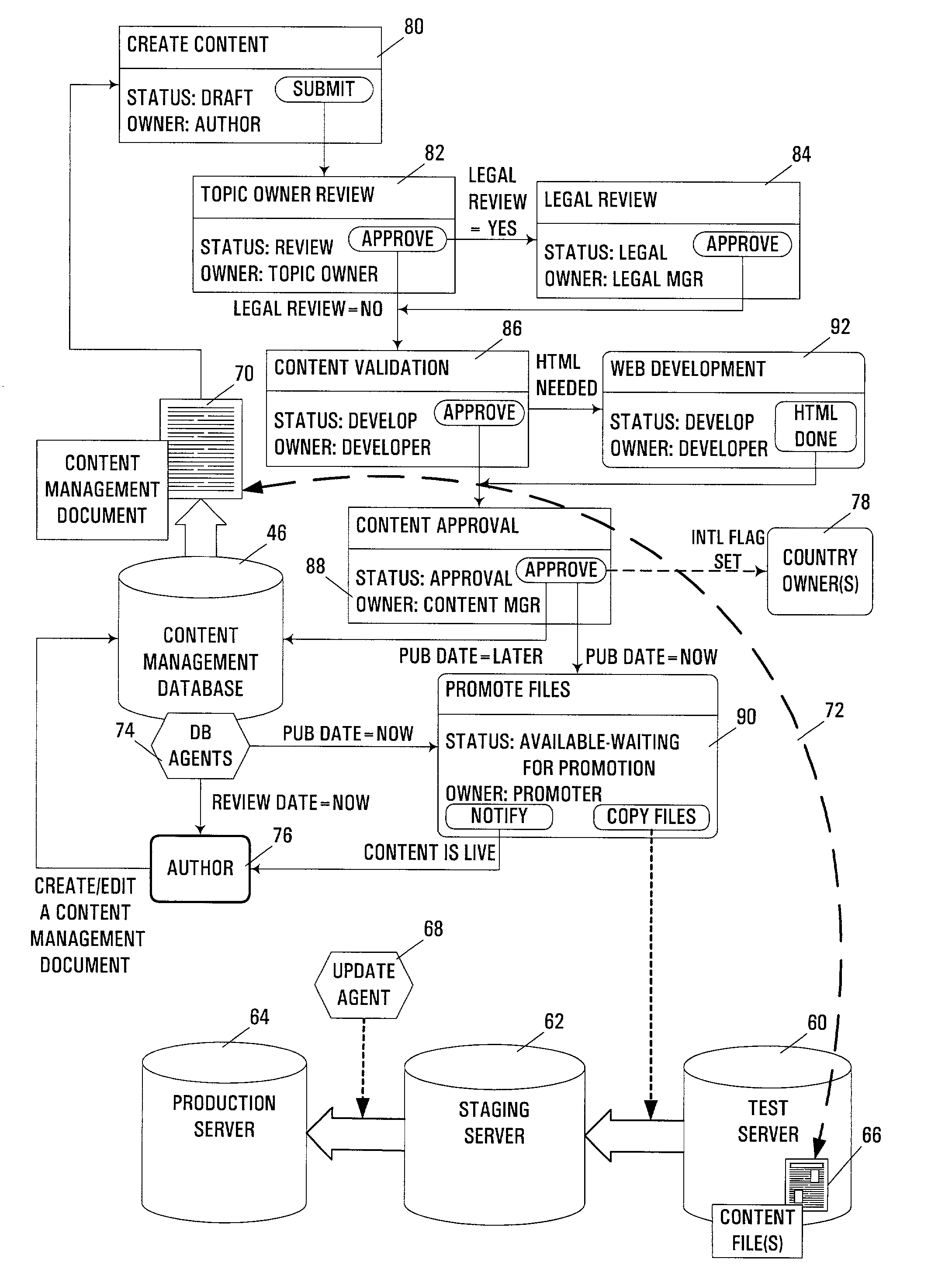
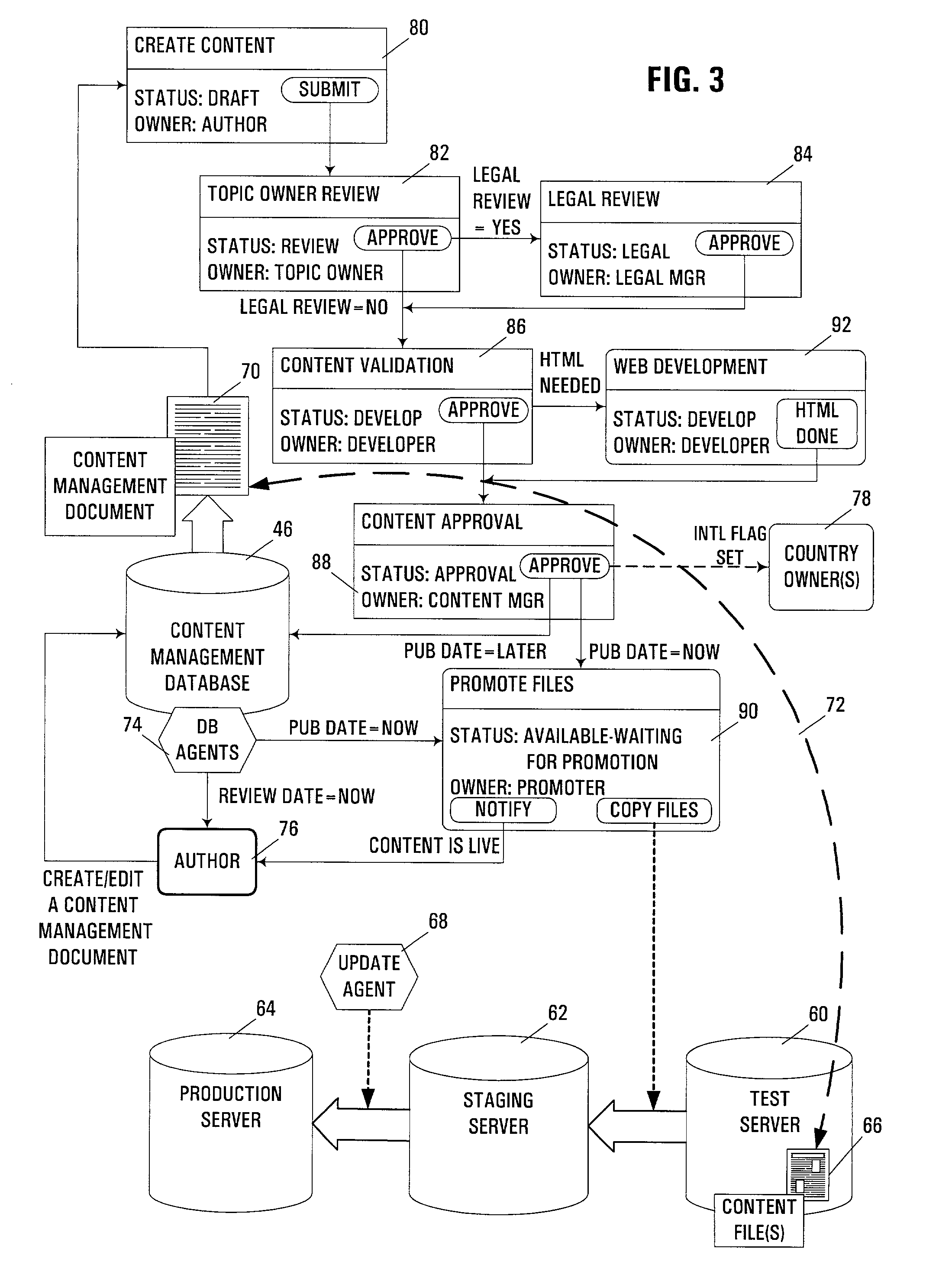
Automated management of internet and/or web site content
InactiveUS7325193B2Good flexibilityEnhanced interactionData processing applicationsDigital computer detailsMessage passingContent management system
An apparatus, program product, and method manage content from a content-controlled database (e.g., web pages or other files maintained in a web site) using a content management record linked to each content-controlled content item in the database. Each content management record is utilized in conjunction with a multi-stage content management process, where at least one stage is a review stage during which approval of an associated content item for a content management record is obtained. As a result of receiving appropriate approval, such an associated content item may be promoted and made available to users of the content-controlled database, with the content management record updated to reflect such a status of the associated content item. Content management information for a content item may be maintained in a content management record that is separate from the content item. Moreover, content management records may be maintained in a groupware-type environment, whereby collaborative tools such as document sharing and messaging may be used to facilitate the interaction among members of a content creation, development and management team during the various stages of a content management process. Furthermore, content management records may be monitored over time to provide for periodic review of promoted content items to ensure that content in a database is maintained current and accurate. In addition, multiple language and / or country versions of a content item may be linked together, such that changes made to one language / country version of a content item may automatically prompt a review of other versions to ensure that the changes are propagated to the other versions when necessary.
Owner:IBM CORP
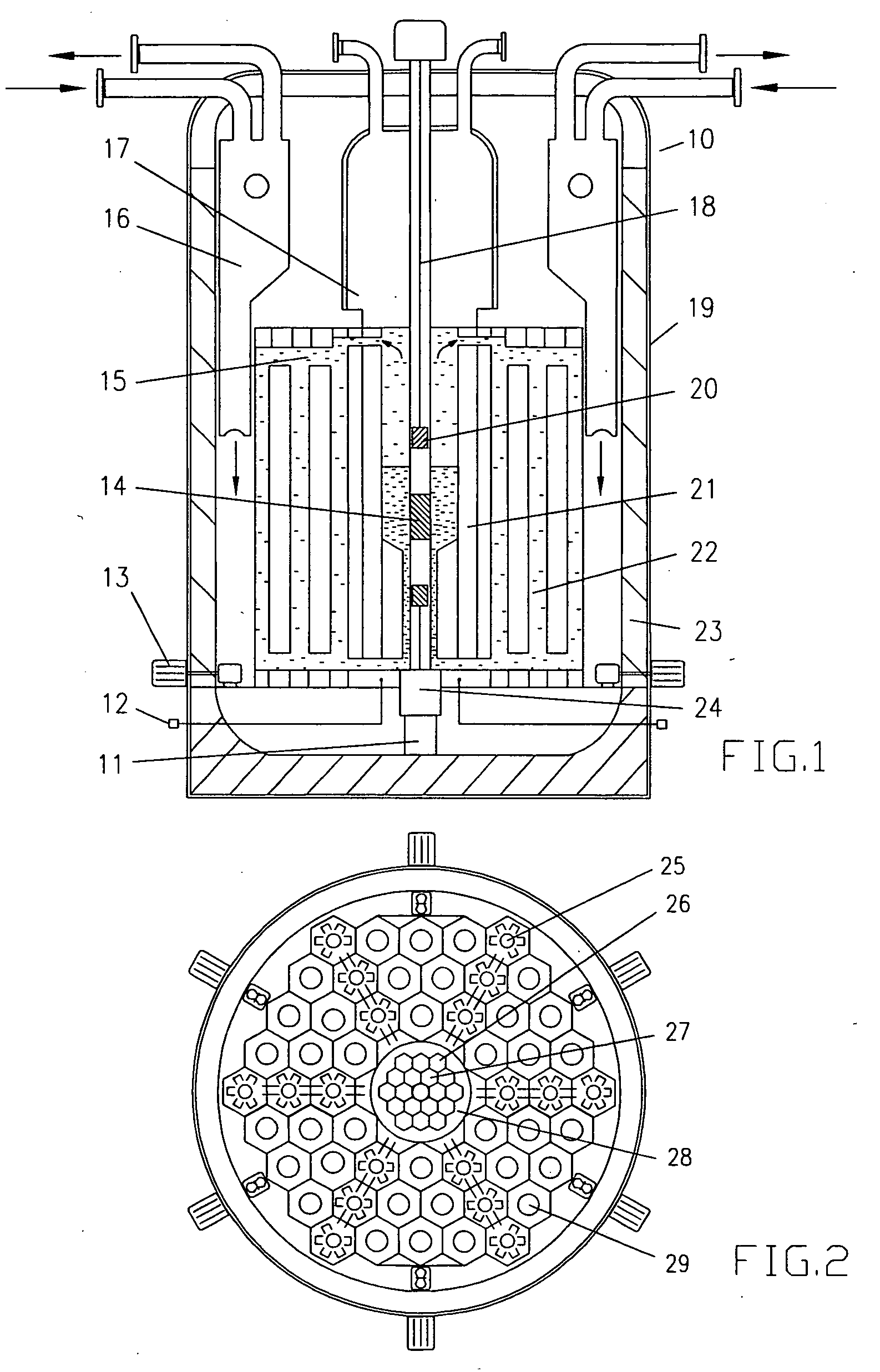
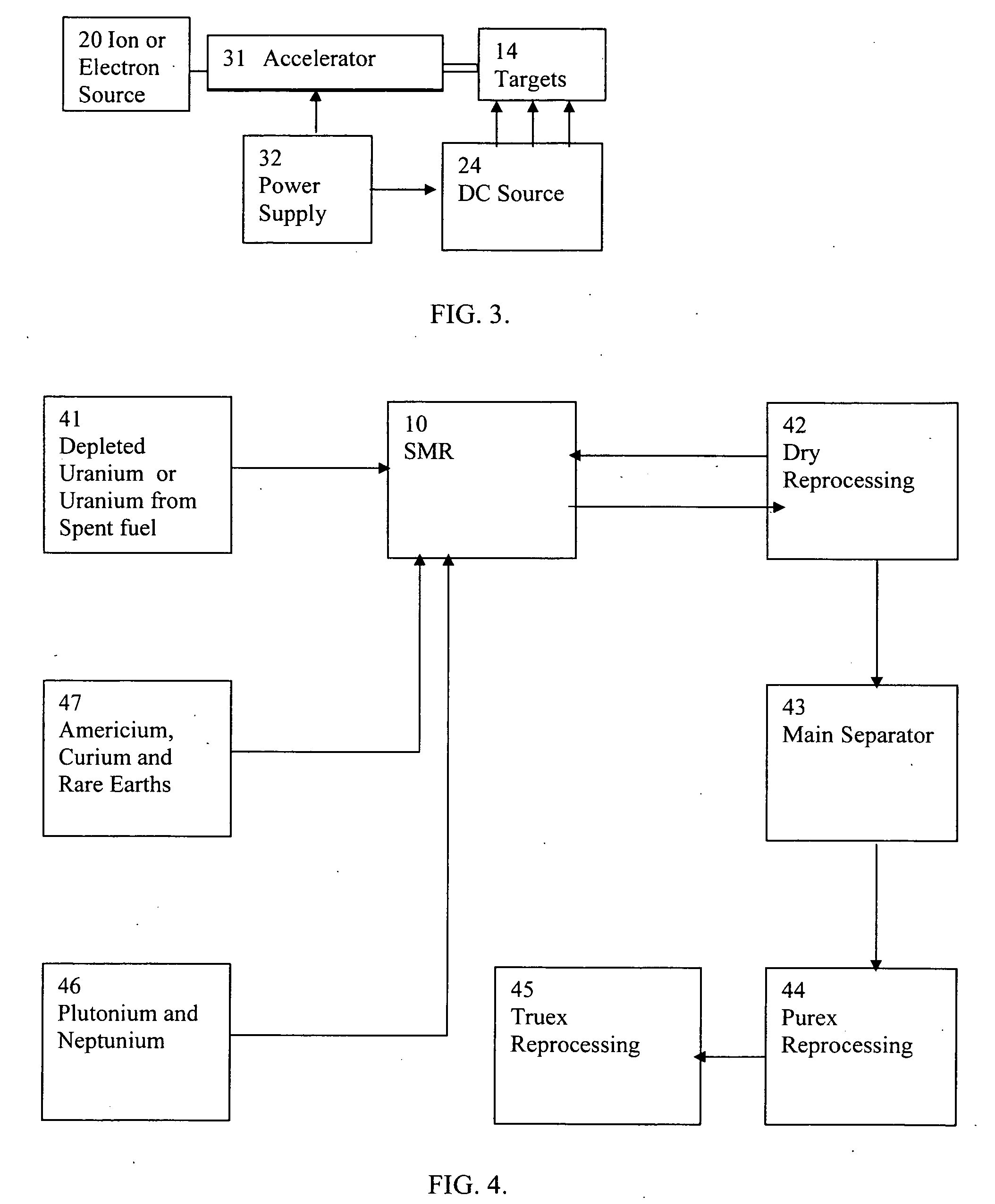
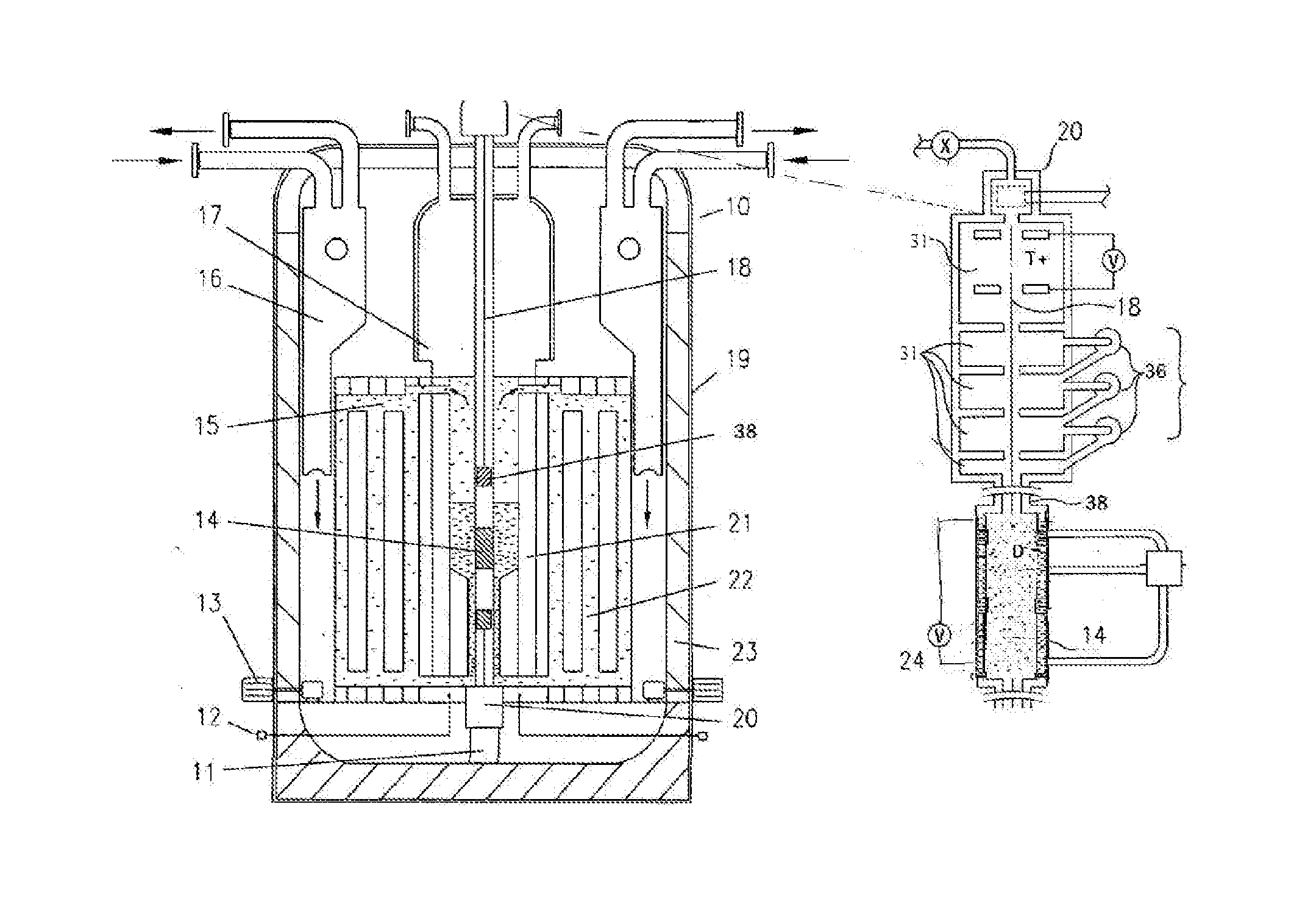
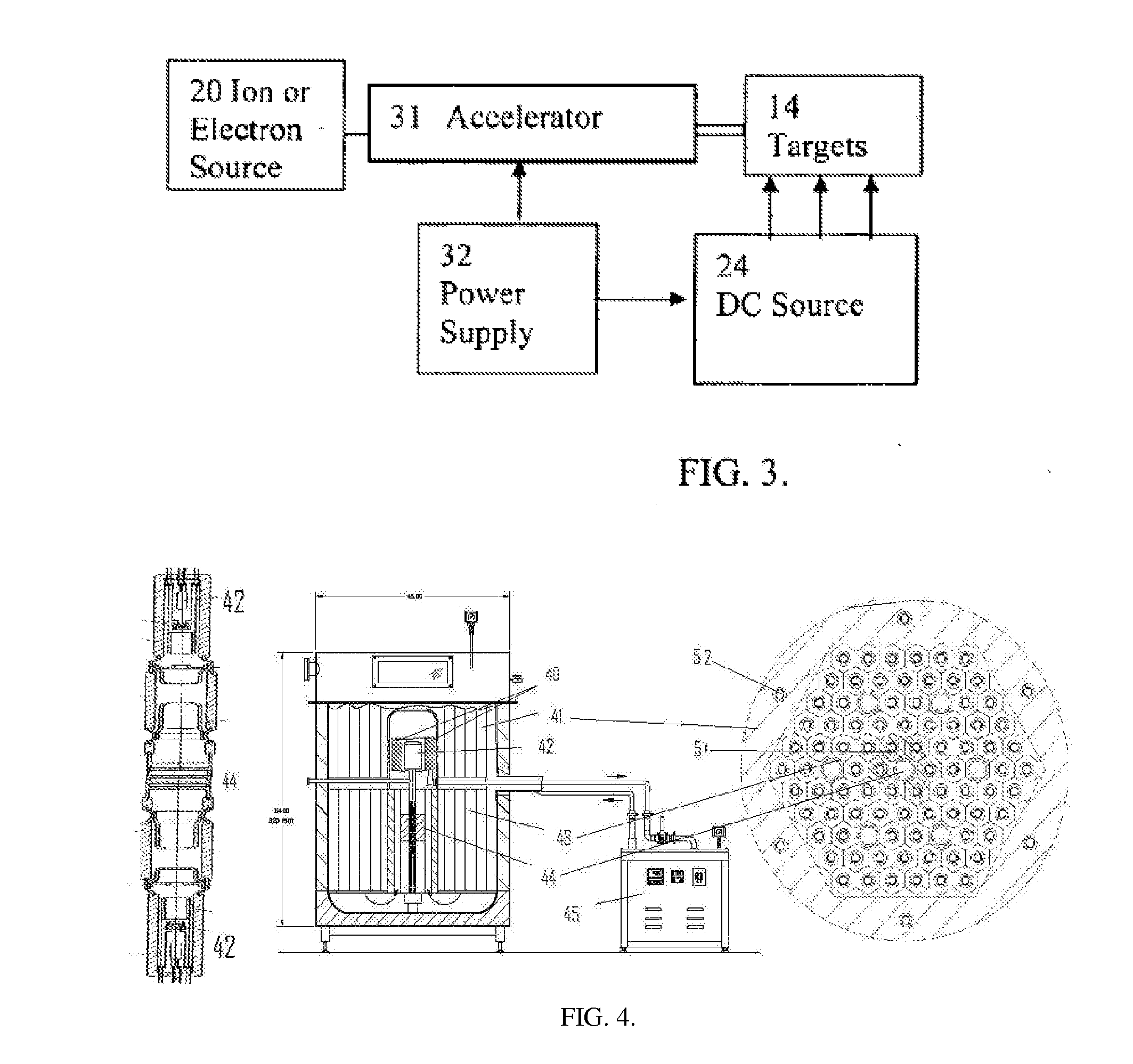
High flux sub-critical reactor for nuclear waste transmulation
InactiveUS20080232533A1Improve distributionImprove economyConversion outside reactor/acceleratorsNuclear energy generationHigh fluxPu element
A process to safely convert about 95% of the nuclear waste into a usable fuel source is disclosed. The process, involving a sub-critical power reactor and a proliferation-resistant fuel cycle, consumes depleted uranium or thorium fuel with fissionable fuel, including reactor or weapons-grade plutonium. The reactor is comprised of coaxial neutron and energy-amplifying regions separated by moderating and thermal neutron absorbing layers. Control of the water or gas-cooled reactor is provided by plutonium-helium loops with a variable volume flow rate and an external source of neutrons that quickly reacts to any fluctuations of the reactor parameters. A second embodiment of the invention is a compact sub-critical propulsion reactor utilizing fission electric cell and thermo-acoustic technology for electrical power generation.
Owner:BLANOVSKY ANATOLY
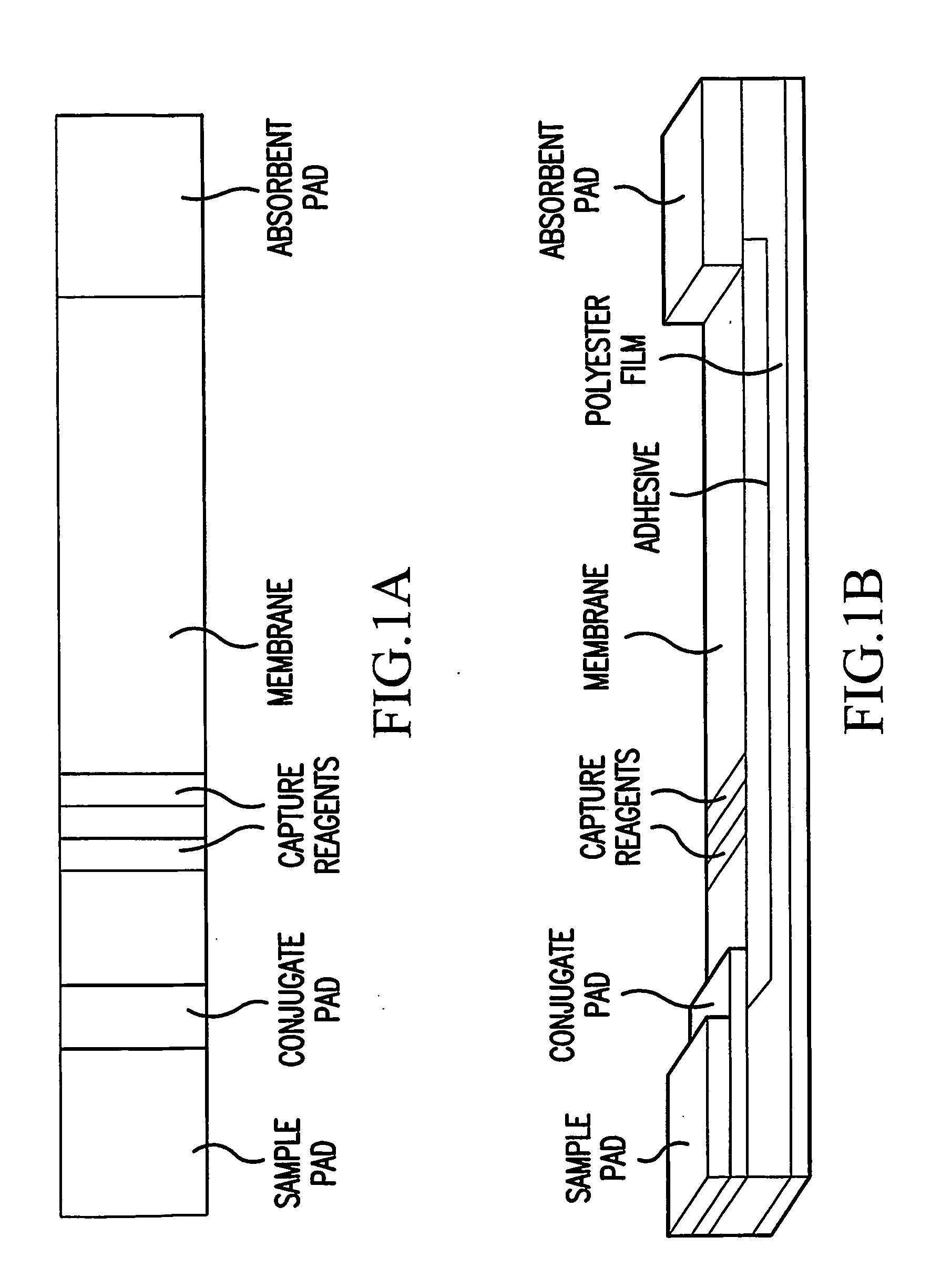
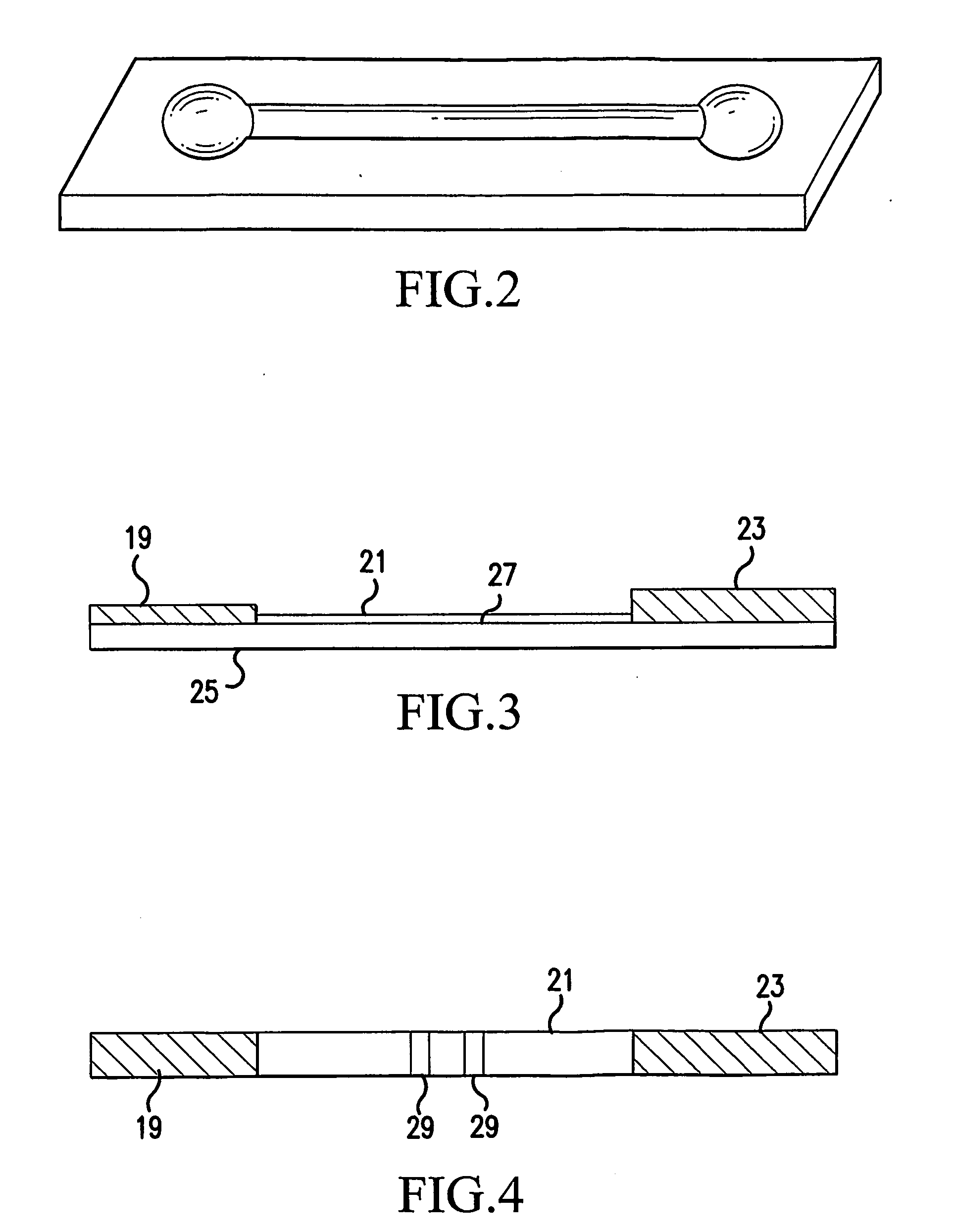
Molecularly imprinted polymer and use thereof in diagnostic devices
InactiveUS20100068820A1AdvantageImprove accuracyMaterial analysis by observing effect on chemical indicatorPharmaceutical non-active ingredientsCross-linkFunctional monomer
An adhesive is provided containing at least one synthetic polymer with receptor sites that enable the selective capture or release of a target molecule. A polymer is synthesized by polymerizing and cross-linking a functional monomer or functional copolymers in the presence of a target or template molecule allowing for reversible interactions between the polymer and the target molecule. The target molecule may be extracted from the polymer creating receptor sites complimentary to the target molecule. Alternatively, the target molecule may remain in the polymer network and be controllably released. The molecularly imprinted polymer is formulated into an adhesive. The adhesive can be used as a component in an in-vitro diagnostic device to release template molecules or to capture target molecules in vacated receptor sites in the synthetic polymer.
Owner:ADHESIVES RES
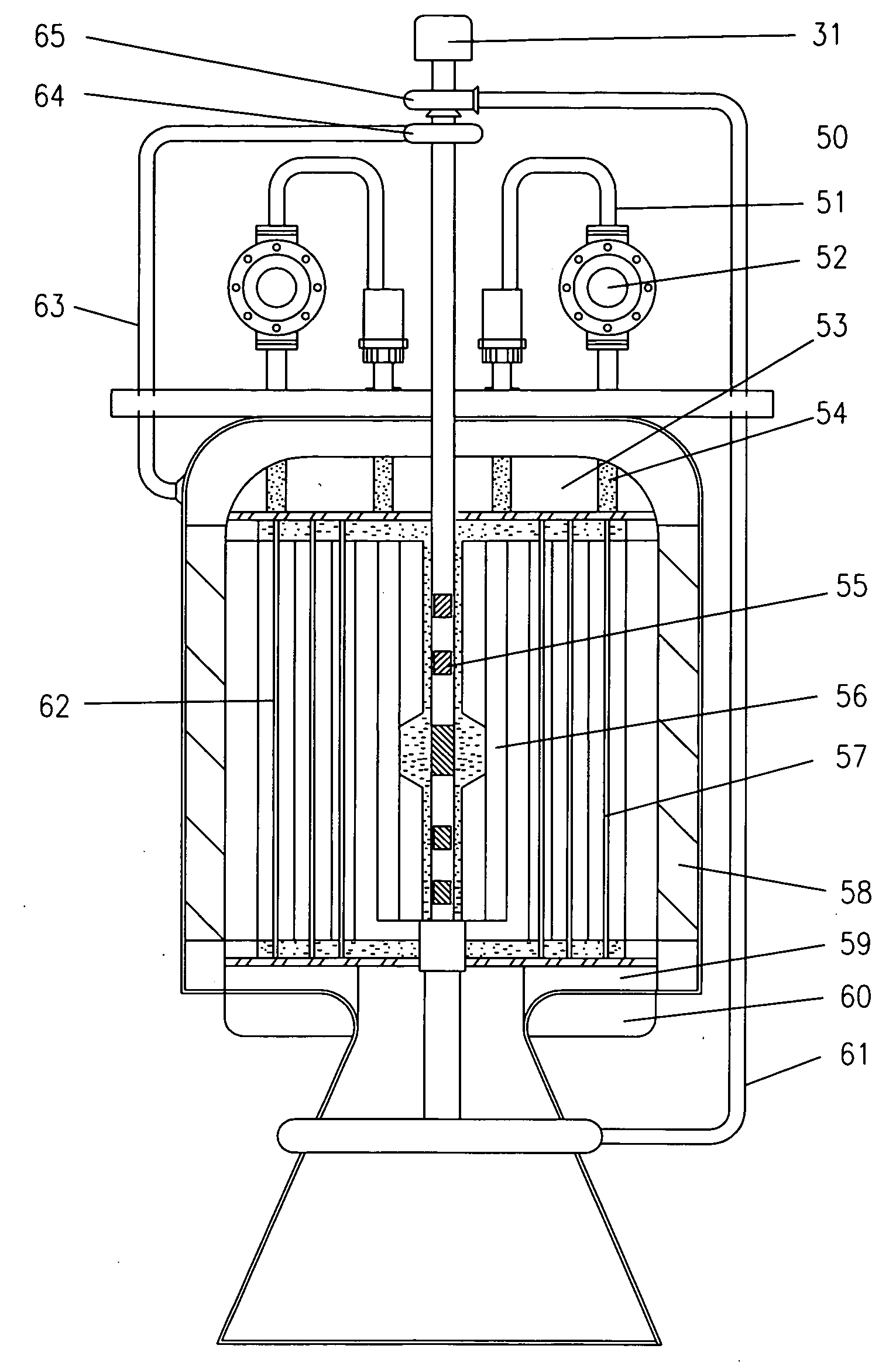
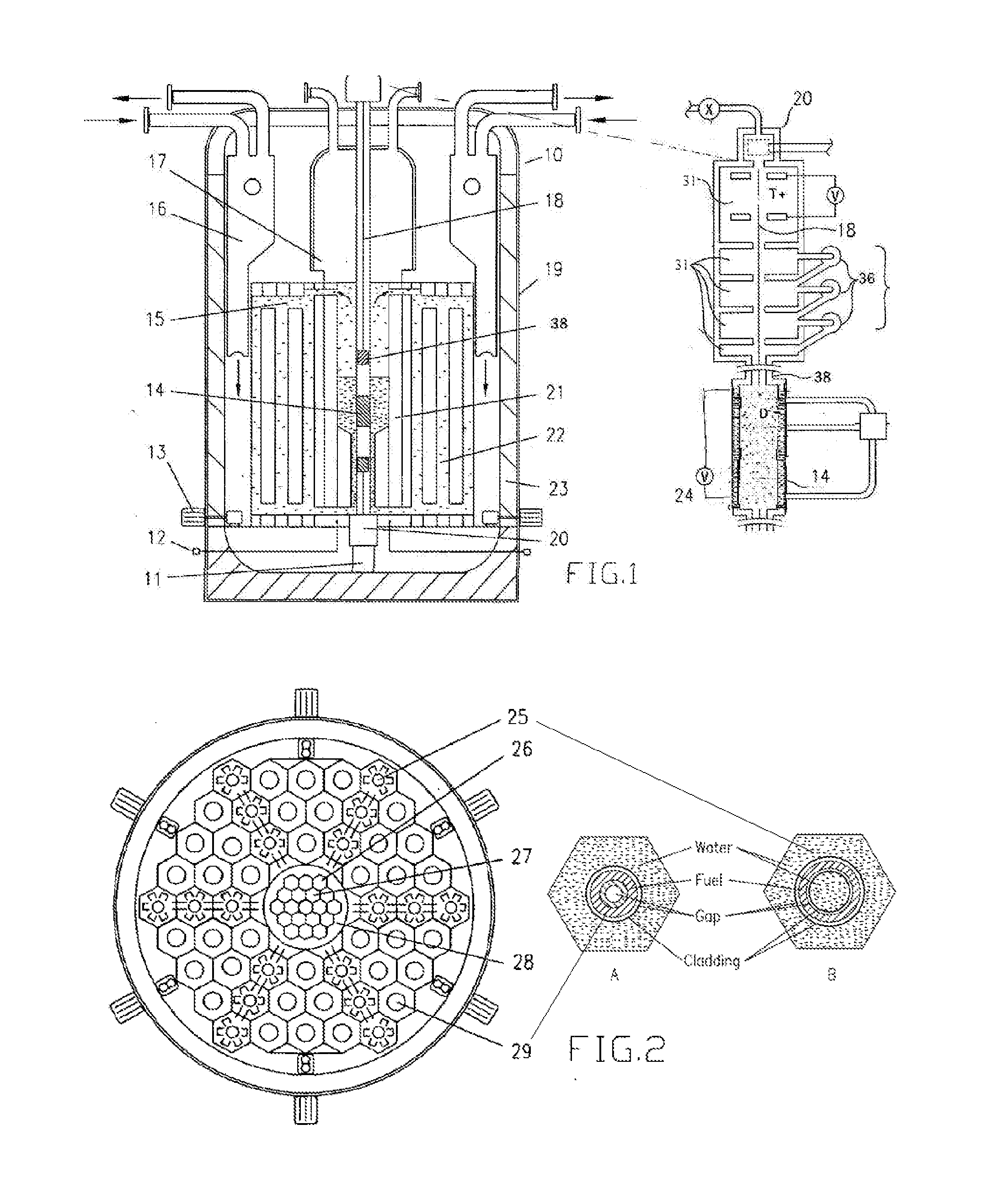
Sustainable Modular Transmutation Reactor
InactiveUS20150098544A1Improve distributionImprove economyIntegral reactorsConversion outside reactor/acceleratorsFission fusionDelayed neutron
A light water reactor to safely convert depleted uranium into a fuel source that could be used as a sustainable source of energy for centuries. The reactor is a type of breed-burn reactor uniquely combined with a proliferation-resistant fuel cycle with no uranium enrichment and no plutonium isolation. It is comprised of a compact factory-produced fast region and a thermal region that produces about 95% of the core power and contains the passageways for transports of delayed-neutron emitters to the fast region, where they can provide additional neutrons (source-based mode) or all the necessary excitation without an external neutron source (self-regulating mode). A second embodiment of the invention is a small unit driven by a neutron source with beam recycling for propulsion, electrical power or radioisotope production. It could also serve as a demonstration facility for the transmutation reactor with fission-fusion fuel.
Owner:BLANOVSKY ANATOLY
Systems and methods for retrieving data
InactiveUS20050108024A1Easy retrievalEasy to updateRelational databasesResourcesInformation retrievalRequest for information
Owner:ADVENT SOFTWARE
Method to split data operational function among system layers
InactiveUS20180041477A1AdvantagePlatform integrity maintainanceSoftware simulation/interpretation/emulationOperating systemData operations
A method for accelerating data transfer and improving data storage in a computing environment is provided that includes splitting a function into two layers of an operating system to generate two separate sets of outcomes from the two layers. A set of outcomes from the two layers are combined so as to be imperceptible to a user save for faster operation. The splitting of the function is evaluated and optimized based on machine performance feedback. A computing system for communicating with a network is provided that performs this method. A bifurcated operating system affords additional advantages in performing function splitting.
Owner:TOUCANH LLC
Biodegradable absorbents and methods of preparation
InactiveUS7309498B2Promote healingEasy peelabilityAntibacterial agentsAntimycoticsHydrophilic polymersWater vapor
A biodegradable microfiber absorbent comprises a substantially homogeneous mixture of at least one hydrophilic polymer and at least one biodegradable polymer. The absorbent can be prepared by an electro hydrodynamic spinning of a substantially homogeneous polymer mixture. Medical dressings for burns and wounds, cavity dressings, drug delivery patches, face masks, implants, drug carriers that comprises at least one microfiber electrospun from a polymer mixture are provided. The dressings can have variable water vapor penetration characteristics and variable biodegradation times.
Owner:MAKSIM BELENKIY
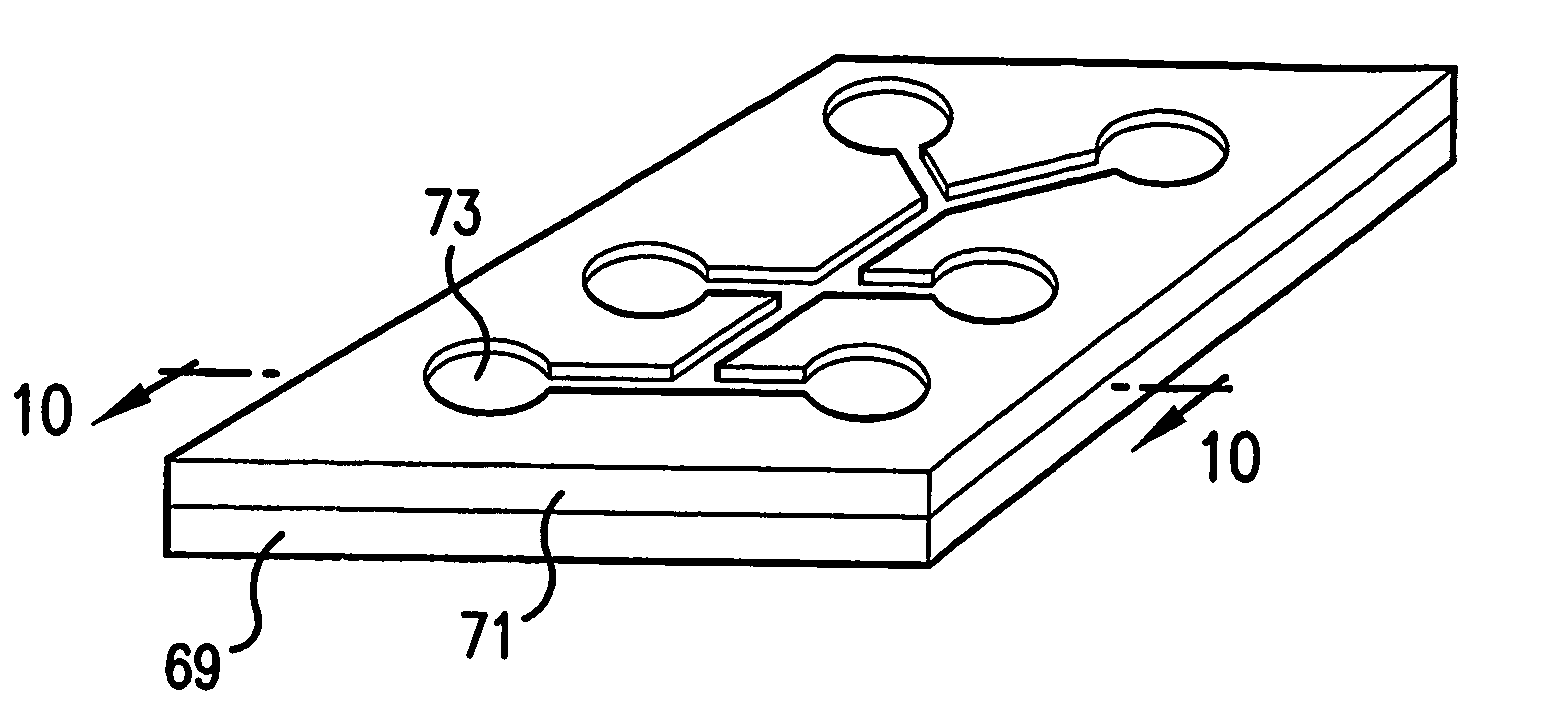
Artery visualization device and artery imaging device
ActiveUS20150133791A1Efficiently incidentAdvantageDiagnostics using spectroscopyIntravenous devicesMedicineSkin surface
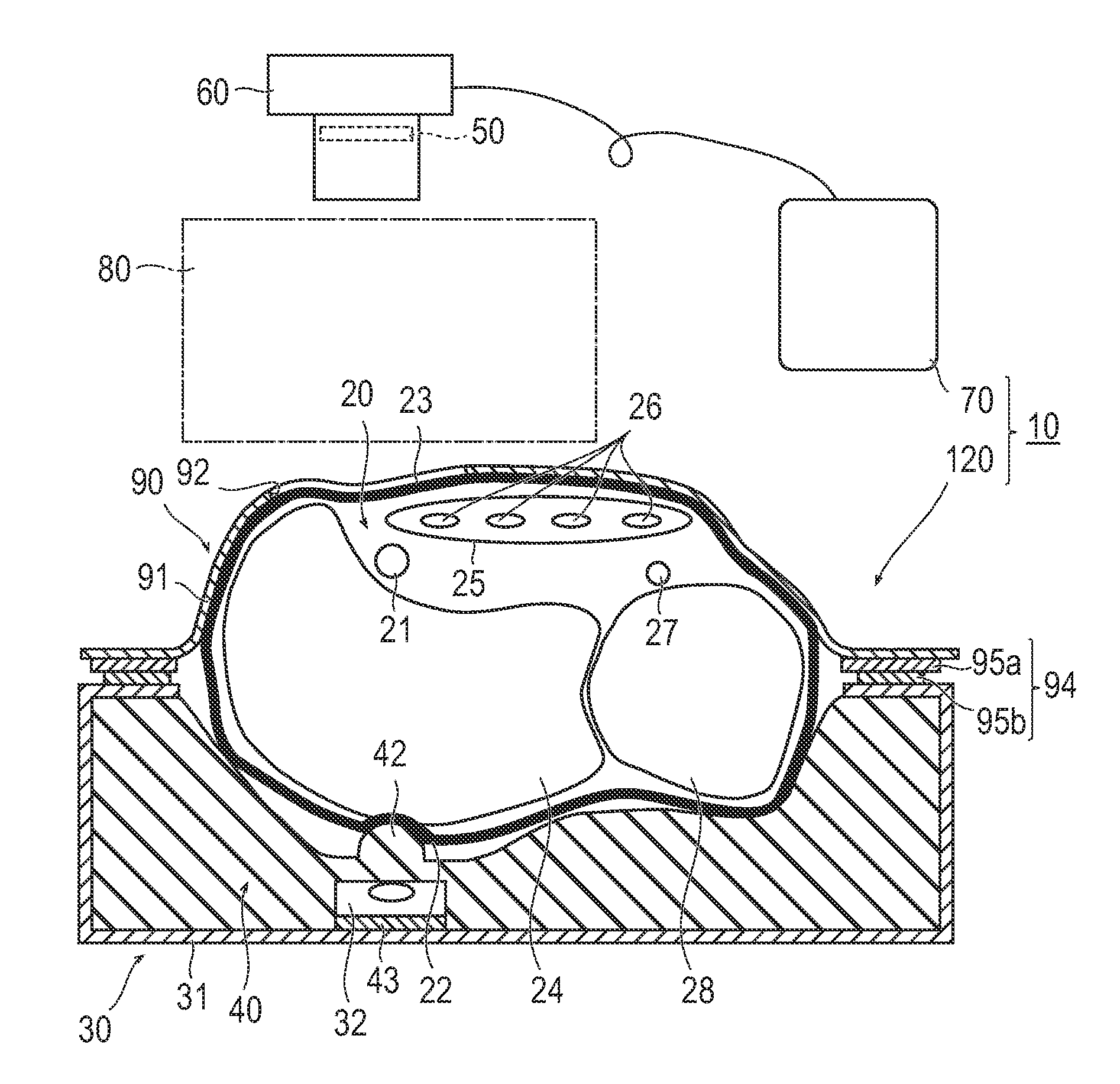
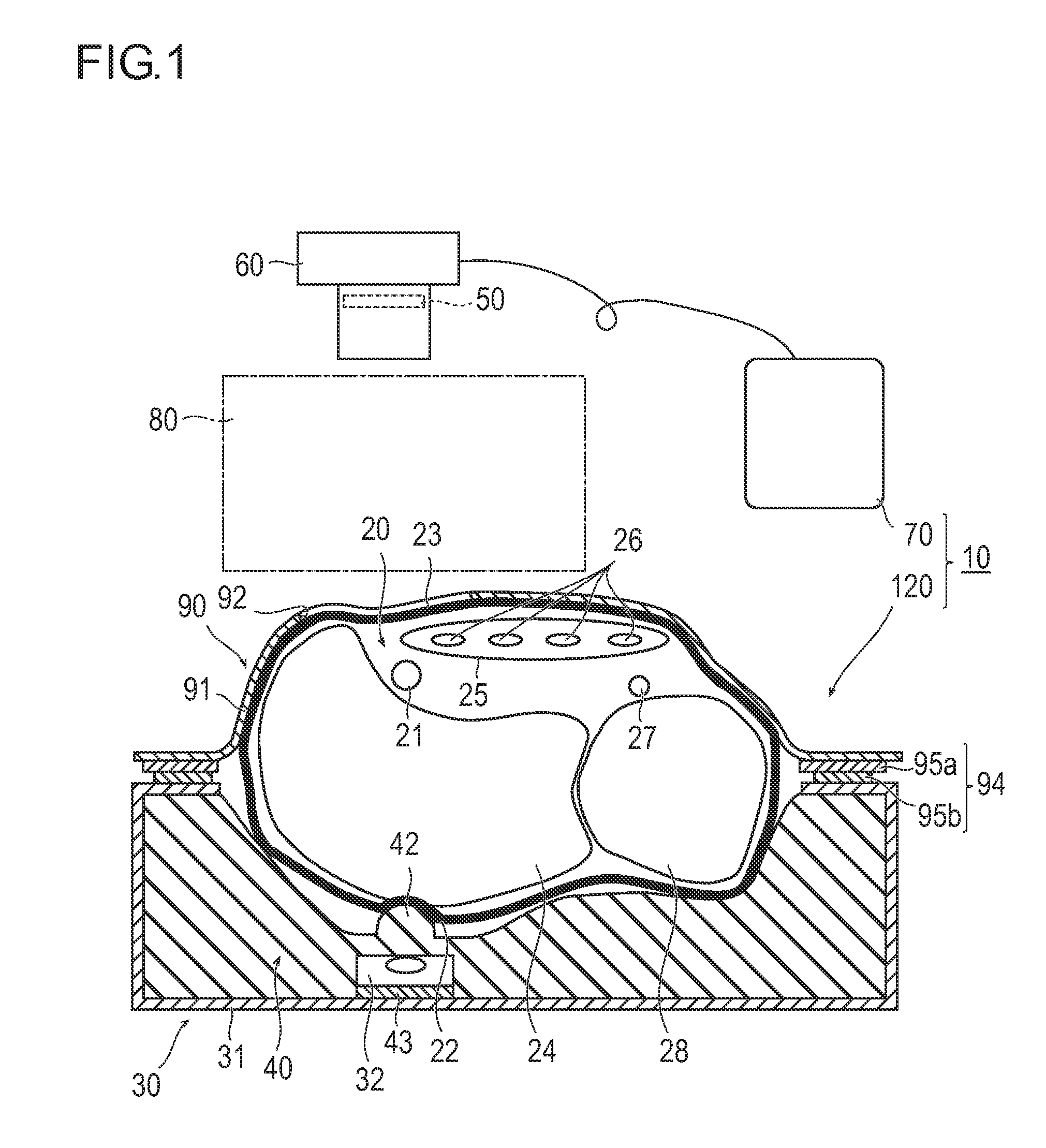
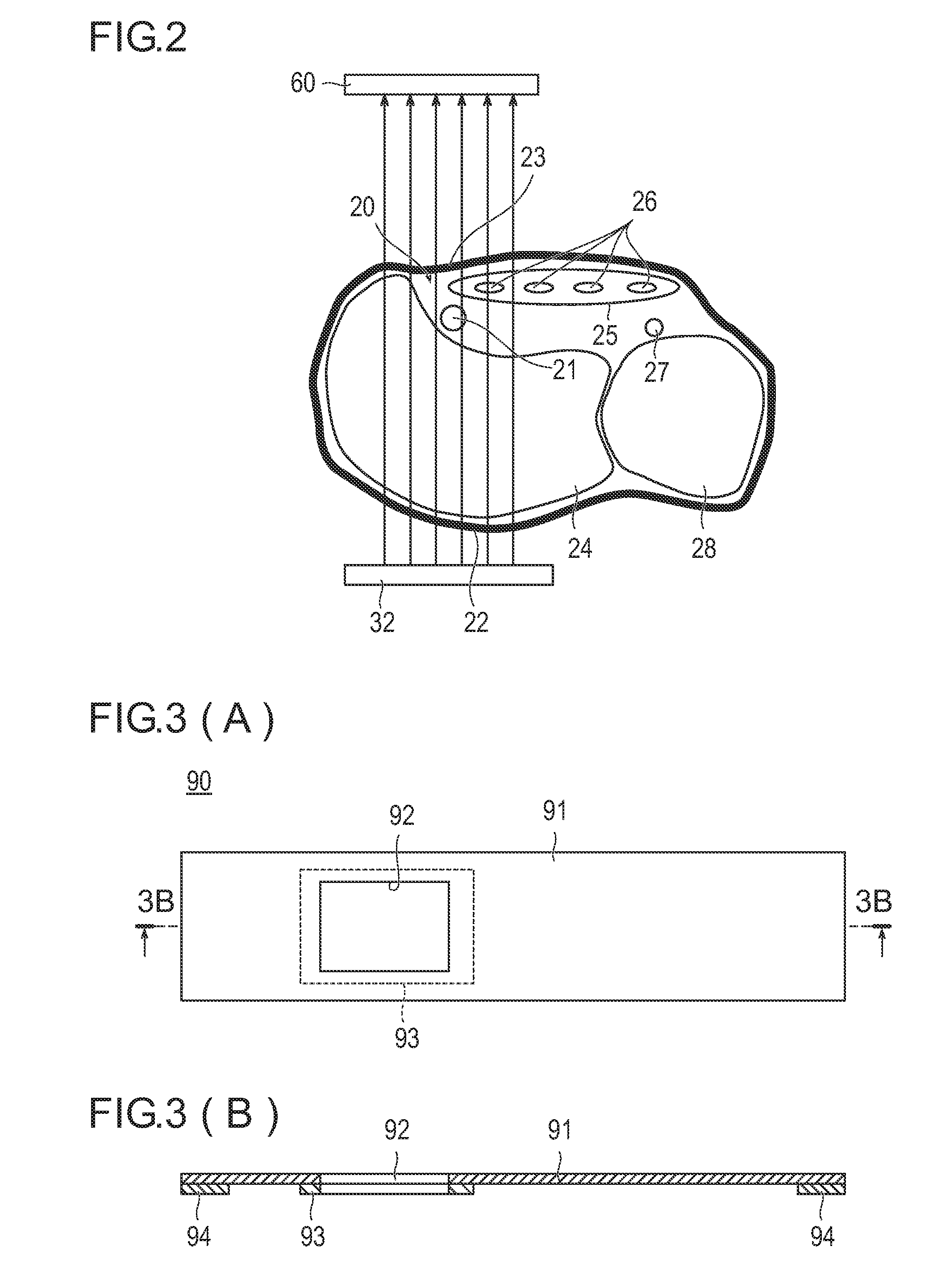
[Problem] To provide an artery visualization device capable of very appropriately visualizing a to-be-punctured artery and an artery imaging device used for the artery visualization device.[Solution] An artery visualization device (10) includes en irradiation unit (30) which irradiates the near-infrared light emitted from a light source (32) toward a back-side skin surface (22) at a visualization site (20) where a to-be-punctured artery (21) is running, a ht guiding part (40) which encapsulates the light source and is pressed against the back-side skin surface and which is formed with a material of transmitting the near-infrared light emitted from the light, source and suppressing reflection of the near-infrared light on the surface of the back-side skin surface, an optical filter (50) which blocks visible light and transmits the near-infrared light passing through a front-side skin surface (23) at the visualization site, a camera (60) (an imaging unit) which receives the near-infrared light passing through the optical filter to capture an image of the visualization site (20), and a monitor (70) (a display unit) which displays the image captured by the camera.
Owner:NAT UNIV CORP KOCHI UNIV
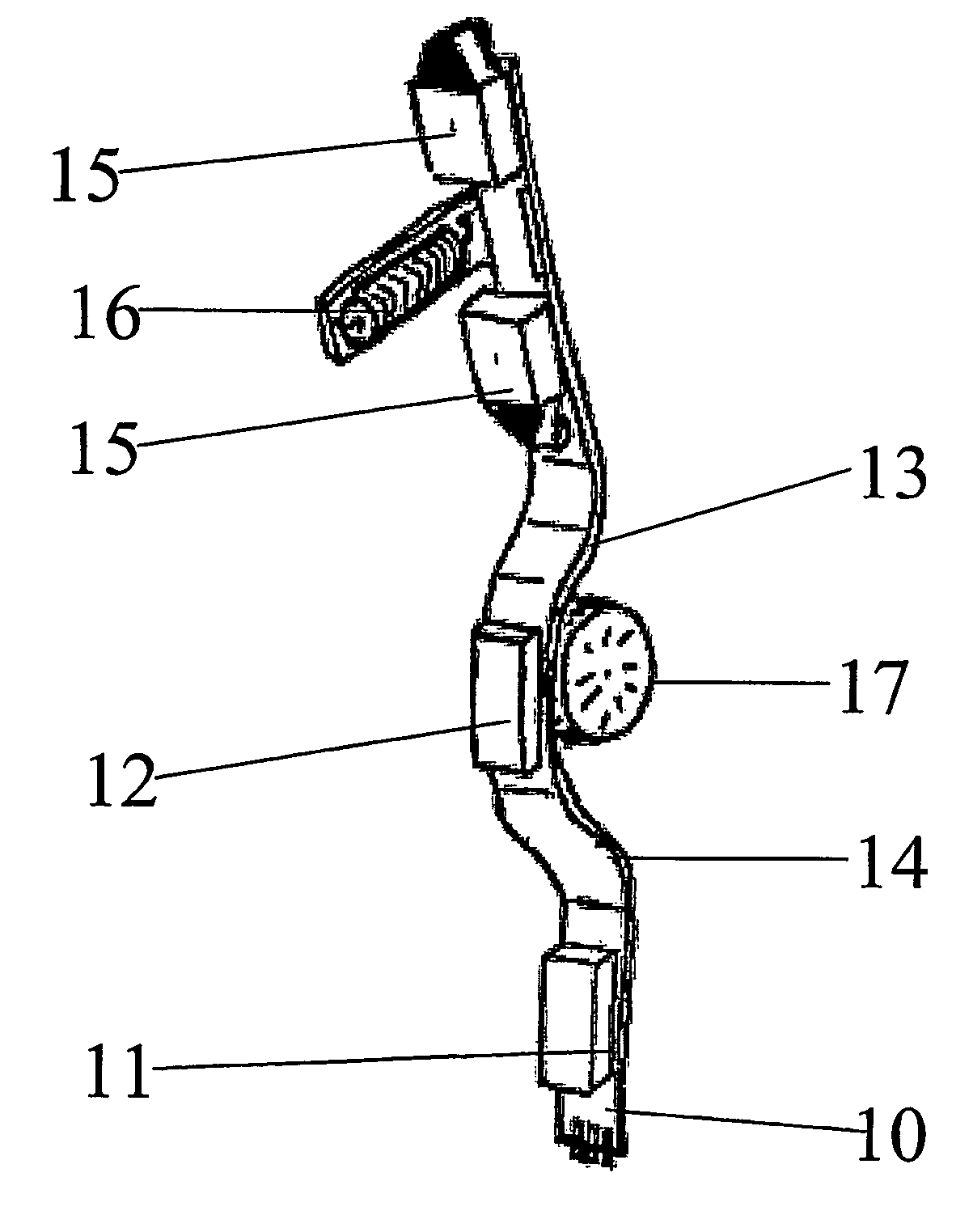
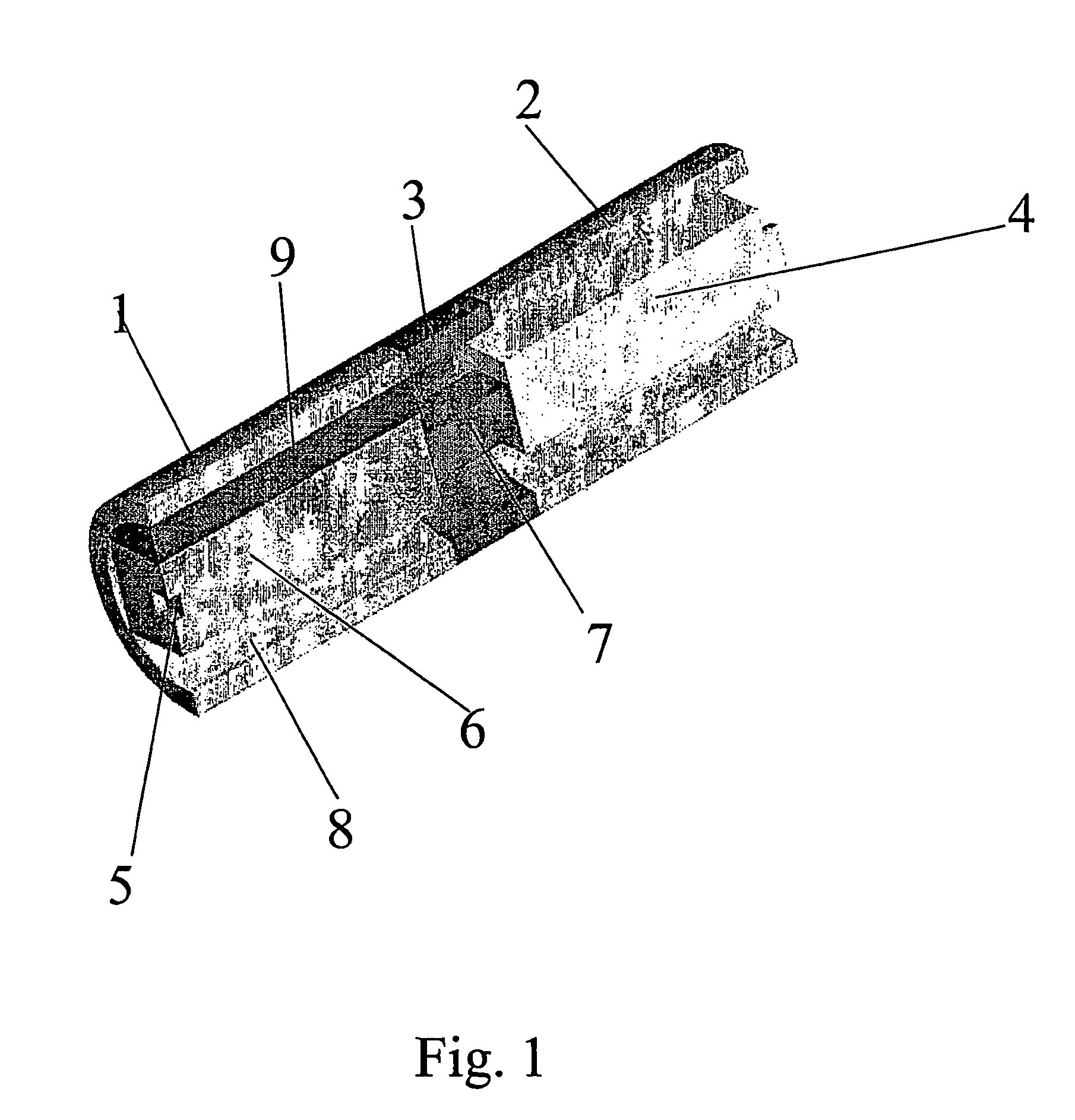
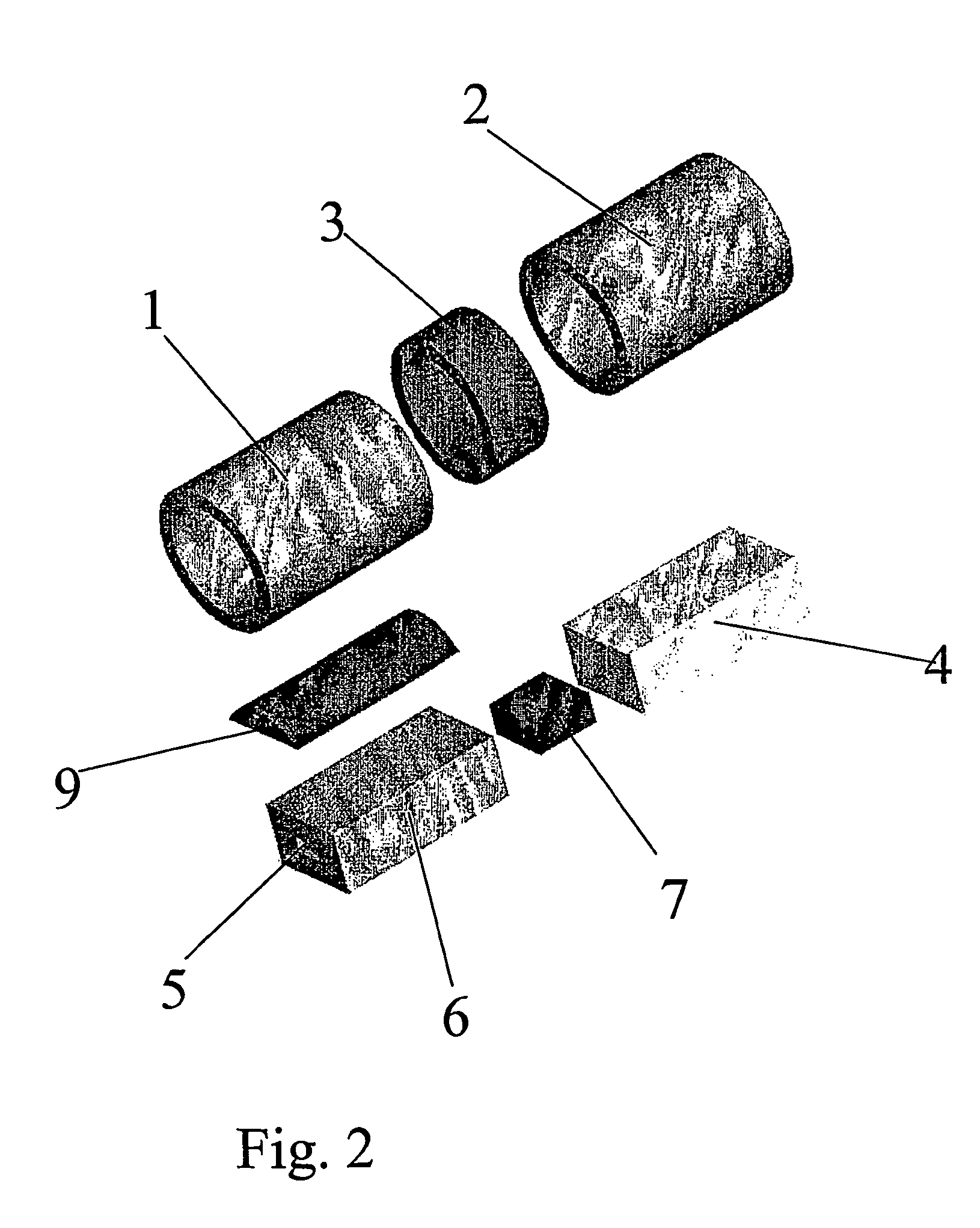
Hearing aid, headset or similar device for delivering a sound signal at the vicinity of the tympanic membrane
InactiveUS7454027B2Increase the use of spaceEasy to usePrimary cell to battery groupingDeaf-aid setsEngineeringHearing aid
A hearing aid or headset for delivering a sound signal to the vicinity of the tympanic membrane includes at least a signal capture device, a signal processing unit, a receiver and an energy storage unit for storage and supply of electric energy, the energy storage unit including two or more sections which are interconnected in a manner which allows angulation of one section with respect to the neighboring section / sections.
Owner:OTICON
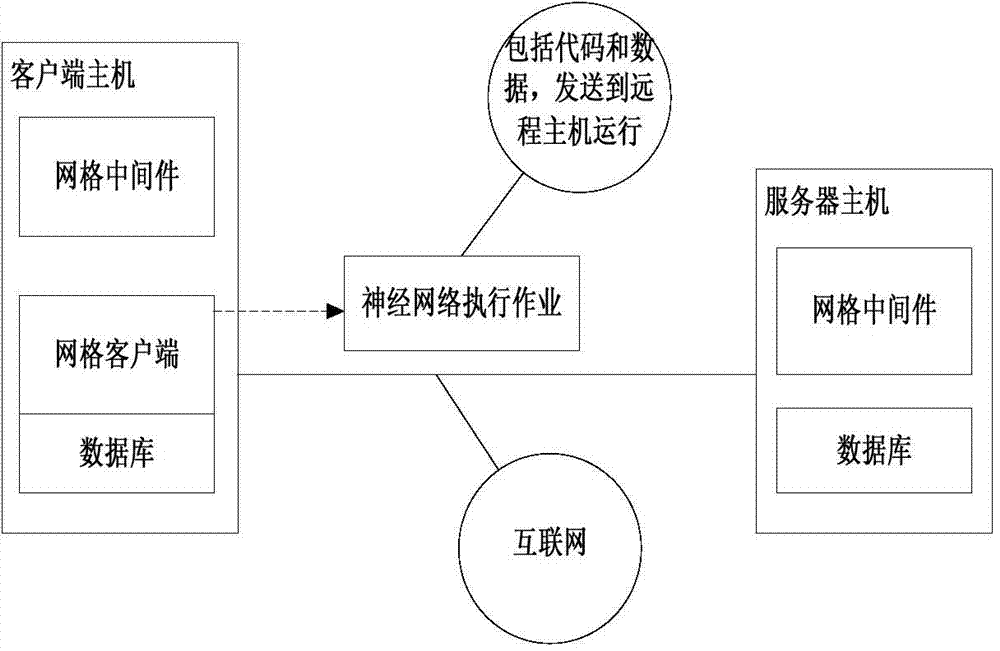
Intelligent computer
InactiveCN104732274AAdvantageGive full play to comprehensive advantagesBiological neural network modelsNeural network systemSoftware engineering
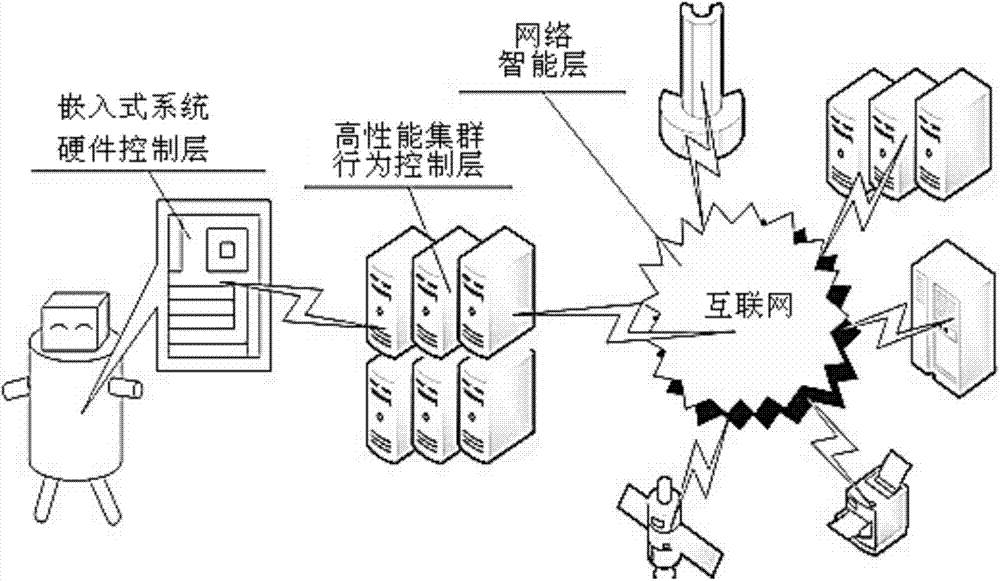
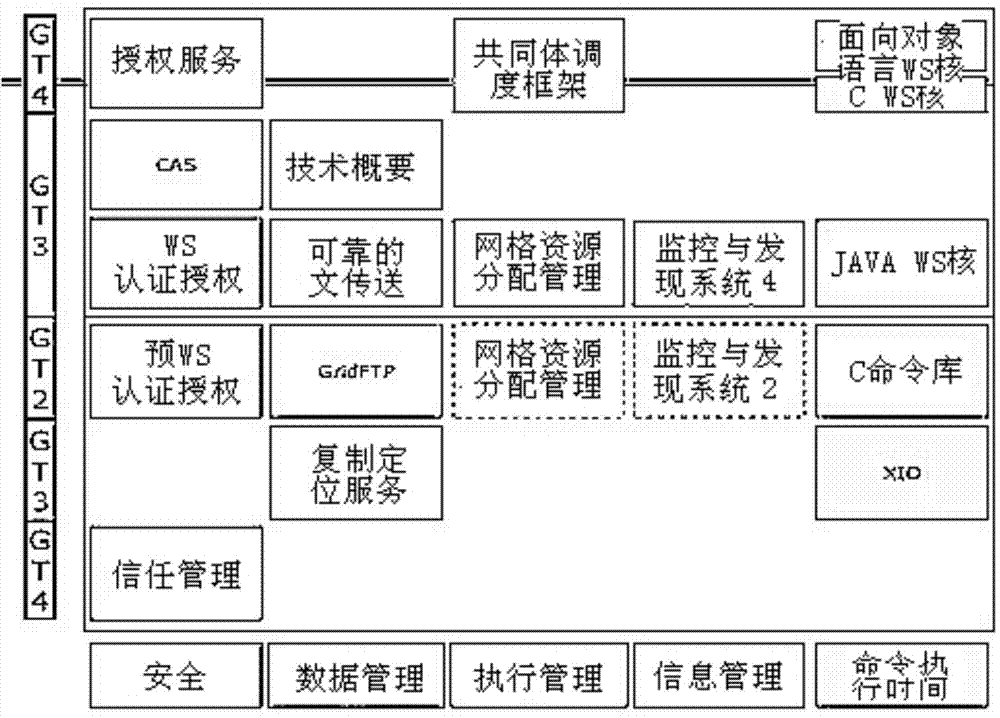
The invention discloses an intelligent computer. The intelligent computer comprises a hybrid neural network application layer, an HnetCP interface layer, a LabGrid middleware layer and a software and hardware resource layer. In the hybrid neural network application layer, a customer realizes a hybrid neural network system and develops a visual hybrid neural network application development environment through an interface of the HnetCP interface layer. The HnetCP interface layer defines various interfaces for operating a hybrid neural network, a top-layer application is an application in the hybrid neural network application layer, and bottom-layer network middleware is the LabGrid middleware layer. The LabGrid middleware layer provides a grid operation environment for the upper-layer application. The software and hardware resource layer is located at the bottom, and software resources comprise kinds of software supporting the upper-layer application. According to the intelligent computer, knowledge stored in the computer and experiential knowledge of people are integrated, and the overall advantages of a computer system are exerted.
Owner:SOUTH CHINA UNIV OF TECH
Multipurpose exercise training device
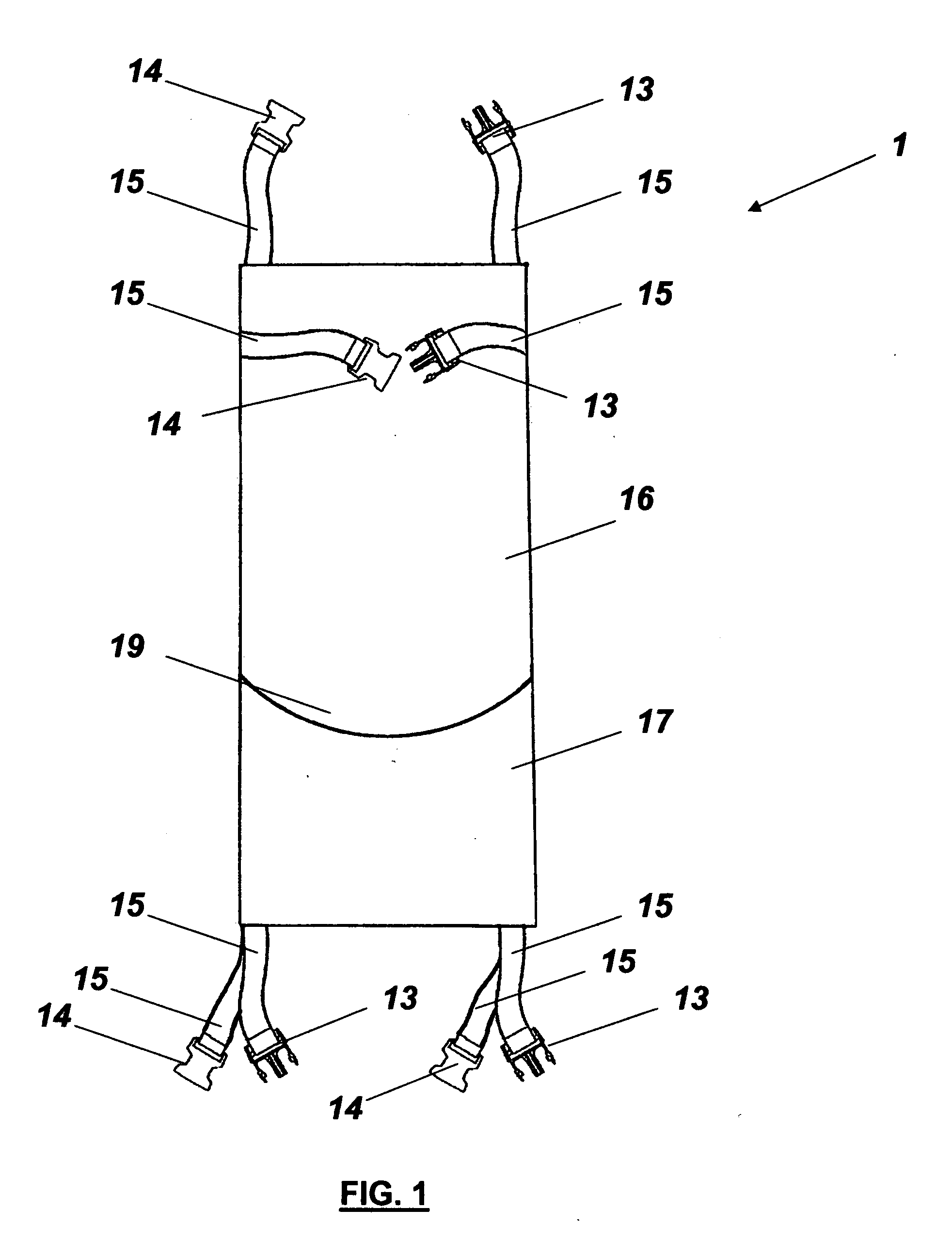
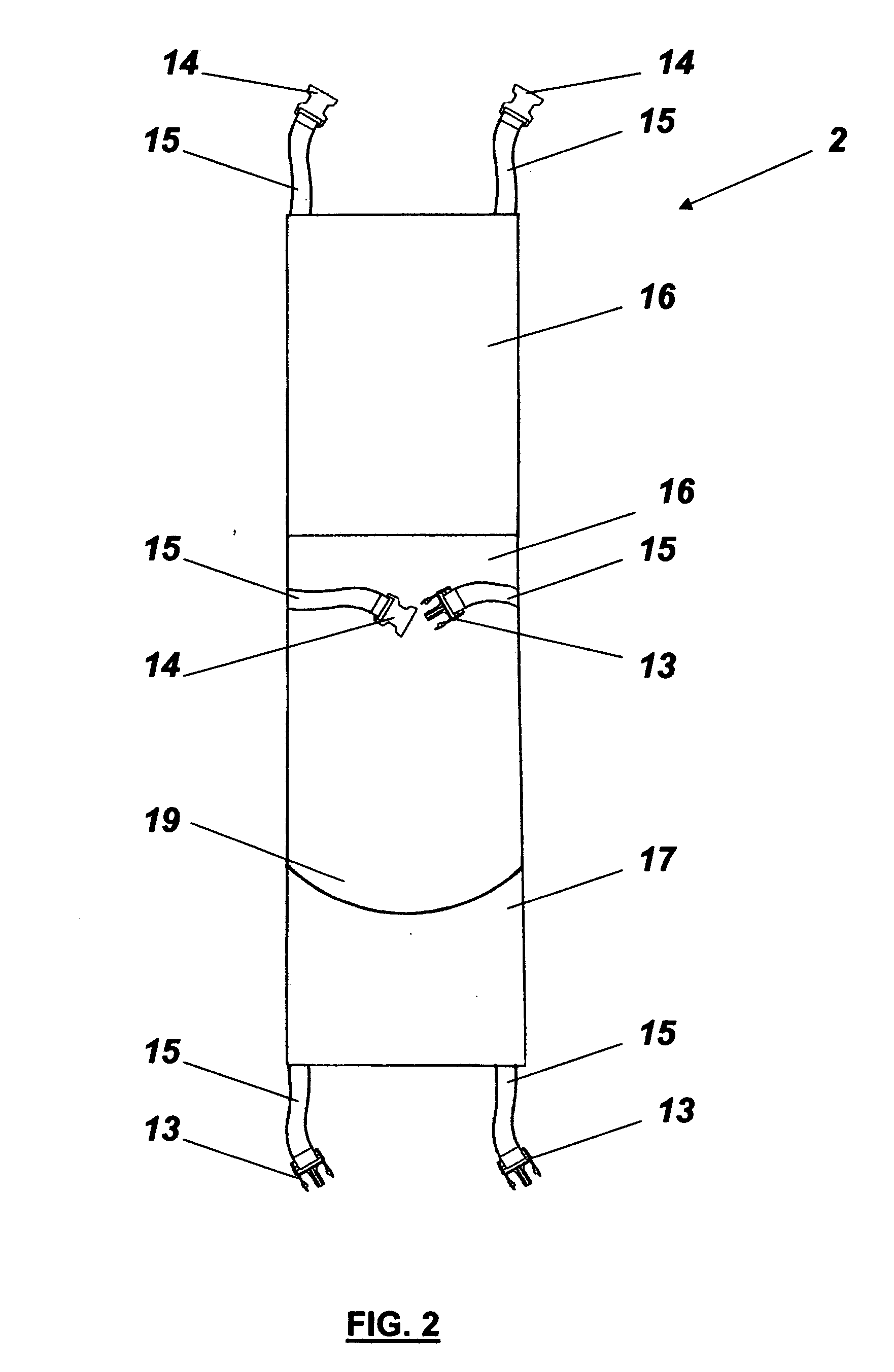
InactiveUS9757604B2Easily transformed in configurationEasy to transformDumb-bellsSkipping-ropesMuscle strengthJumping rope
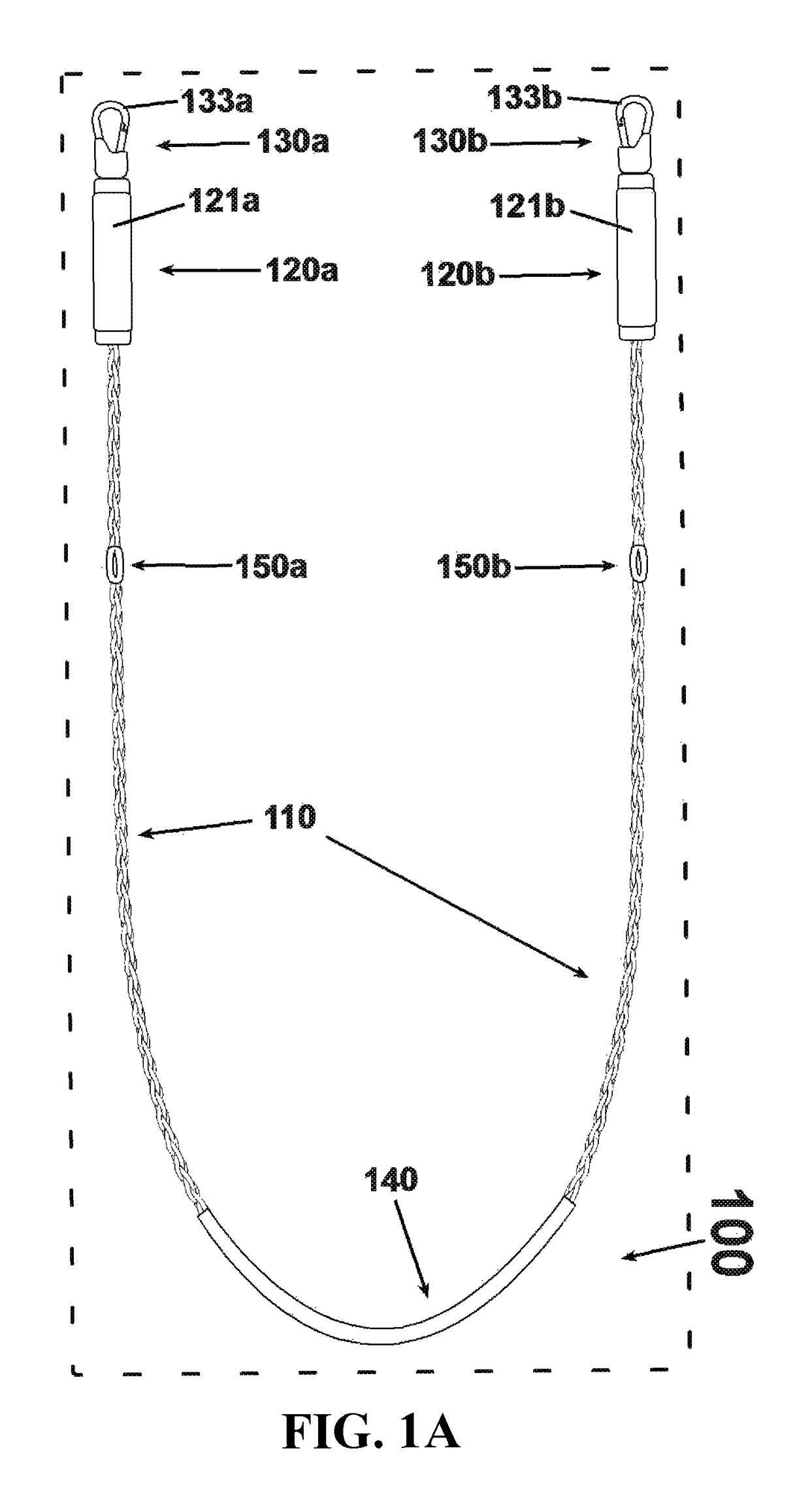
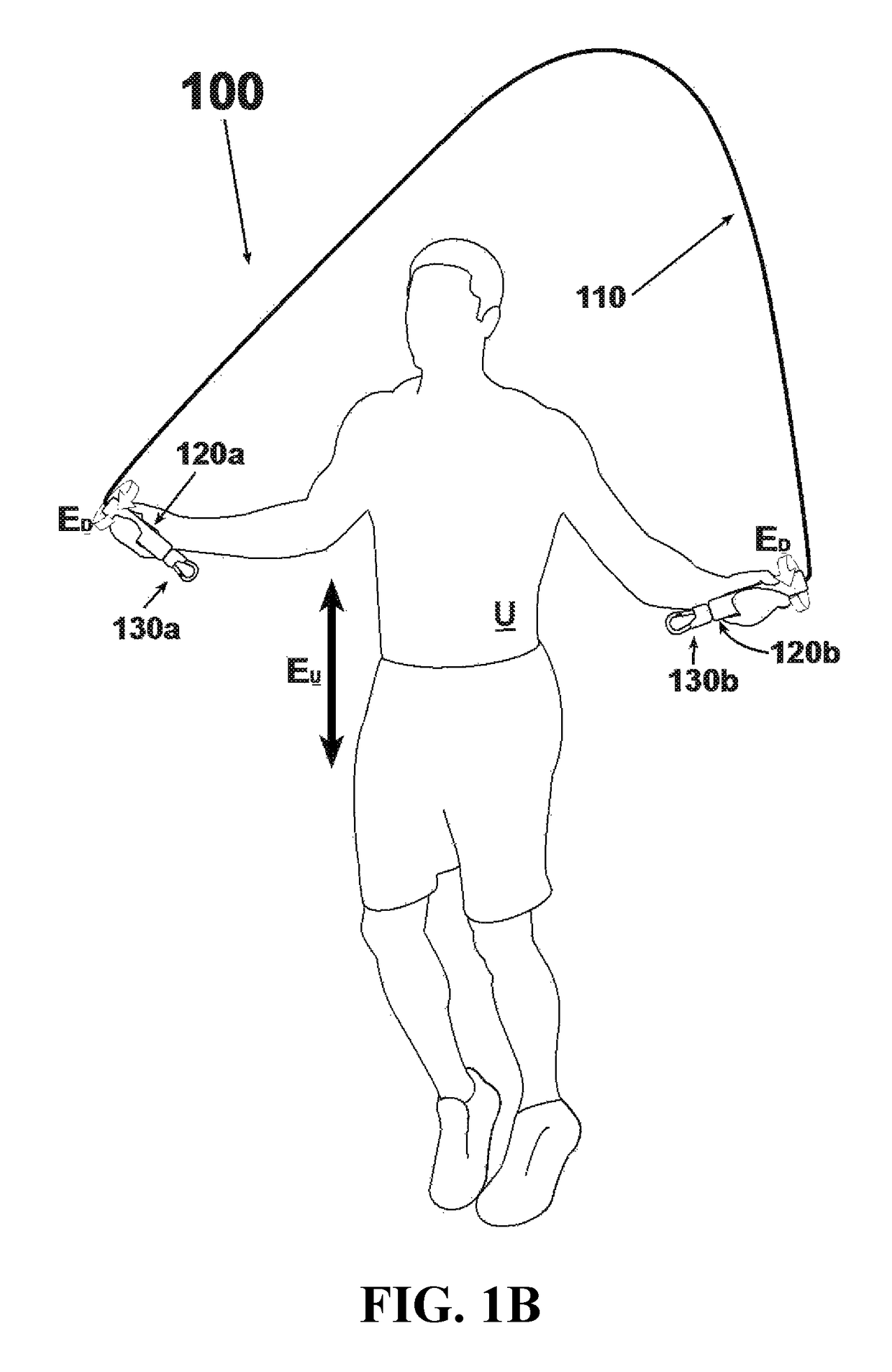
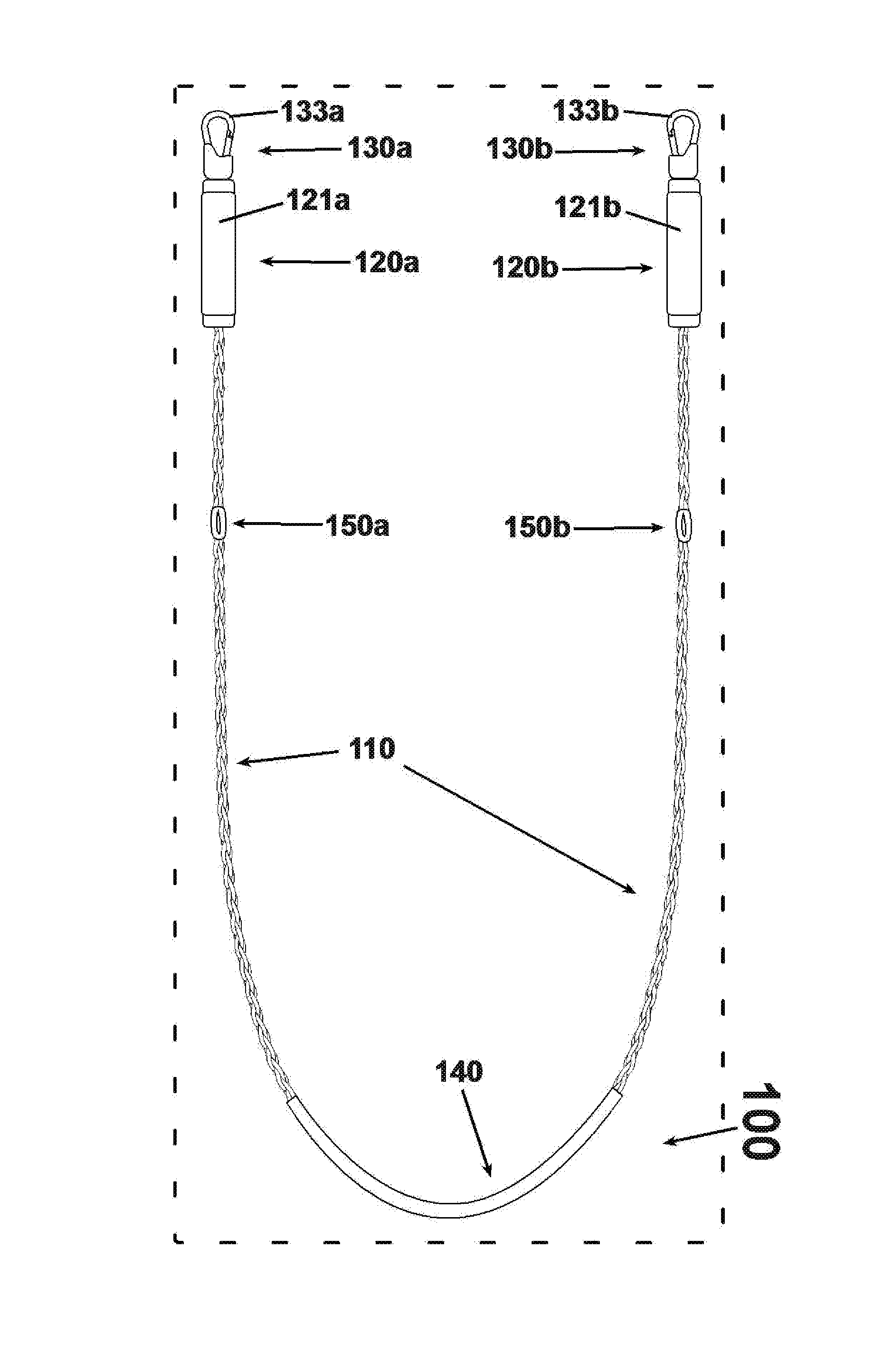
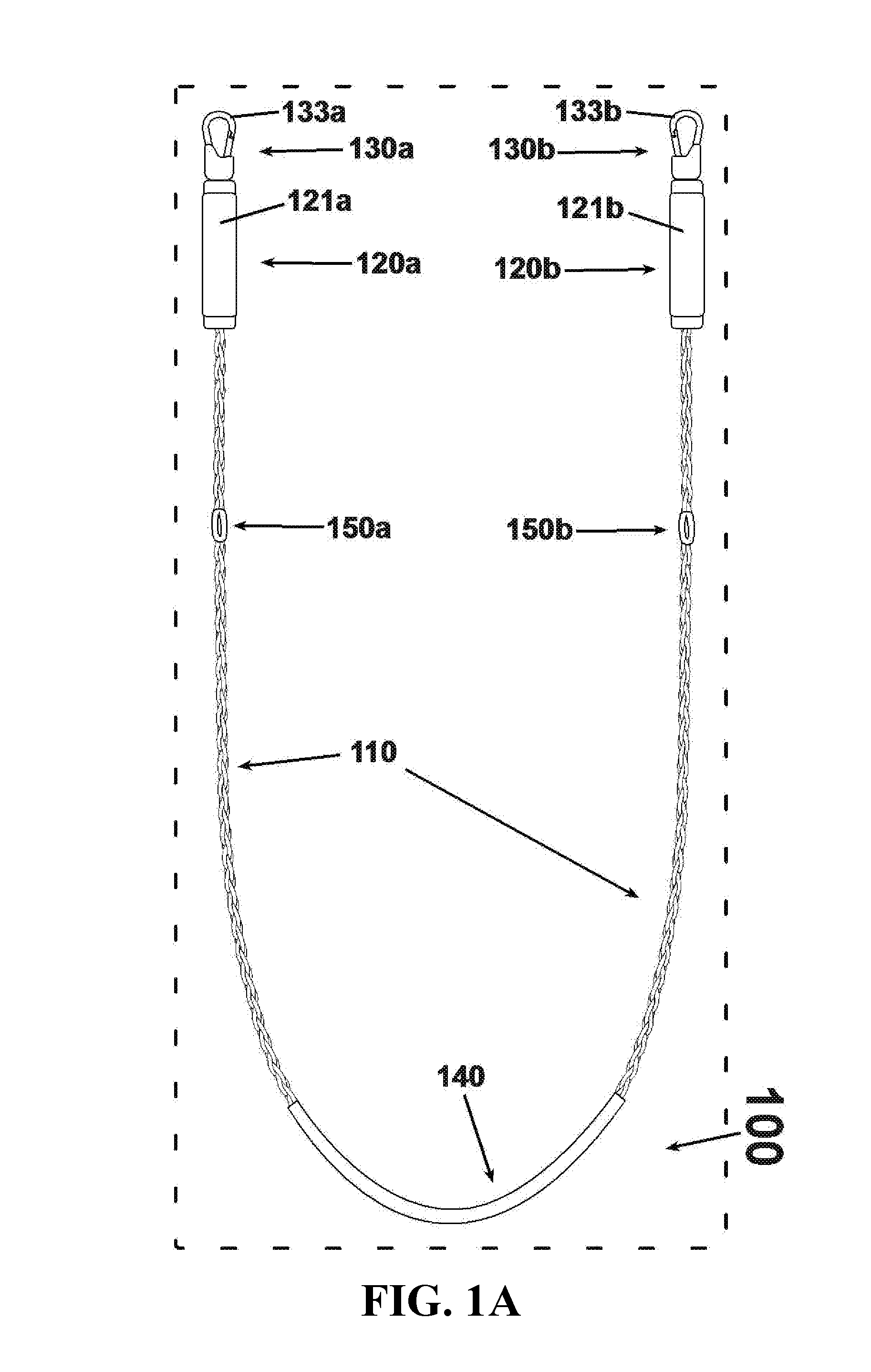
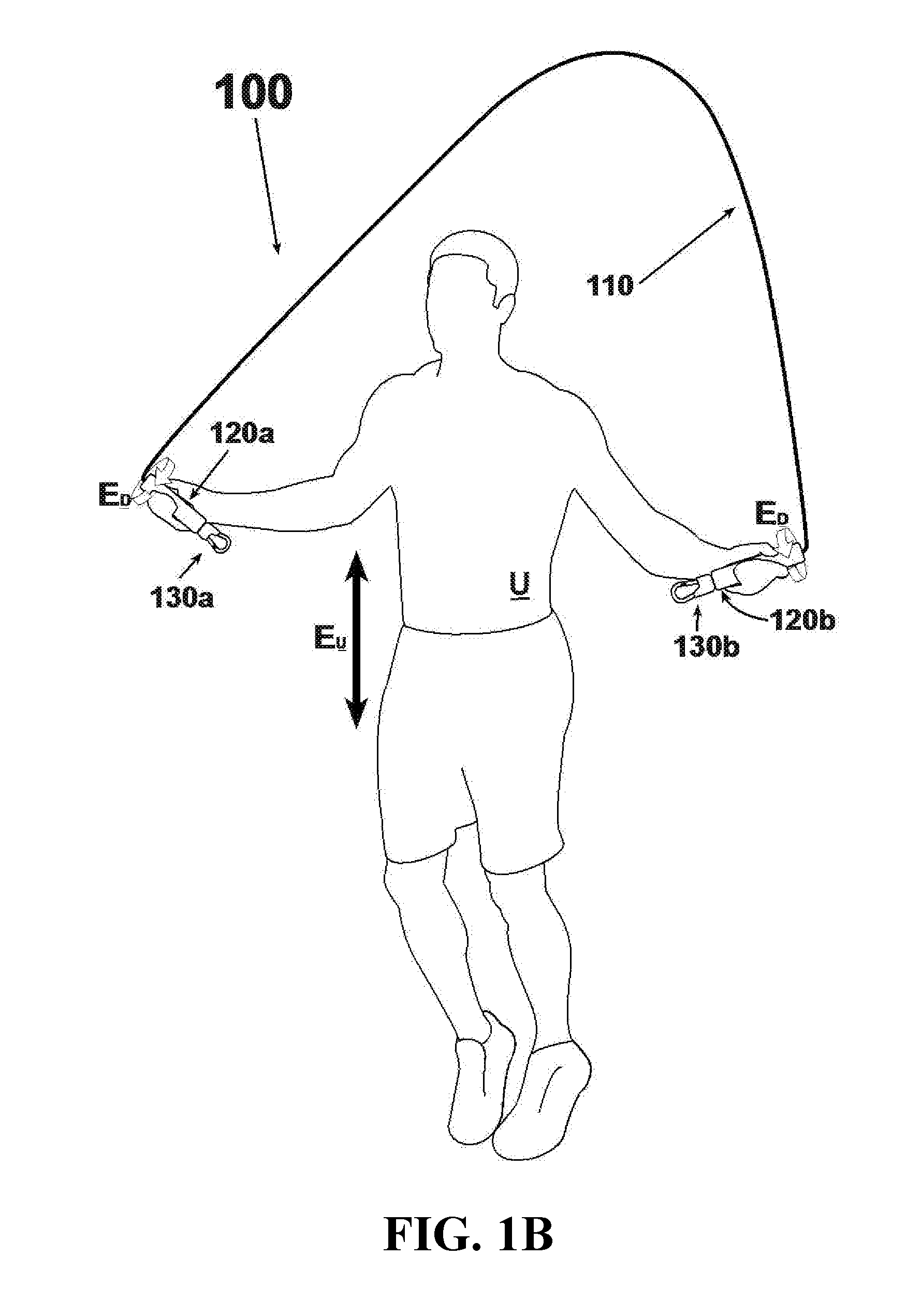
An exercise device having many advantageous features is described, including the ability to provide users with multiple modes of training by transforming the device into different configurations. Users can exercise their cardiovascular endurance by using the device in its jump rope configuration. Users can also engage in muscular resistance training using the same device configured to one of multiple possible resistance training configurations. Resistance training configurations provide a method of suspending users' bodyweight such that they can train their muscular strength capacity by exerting themselves against the force of gravity. Resistance can be selected from nearly zero resistance to a user's full body weight, with the ability to easily adjust between exercises and between users, and the ability to easily transform the device between configurations to provide for versatility and ease-of-use. The device includes an inelastic length member with two clip assemblies and two handles and / or grips at both ends.
Owner:CARTER MATTHEW RODERICK
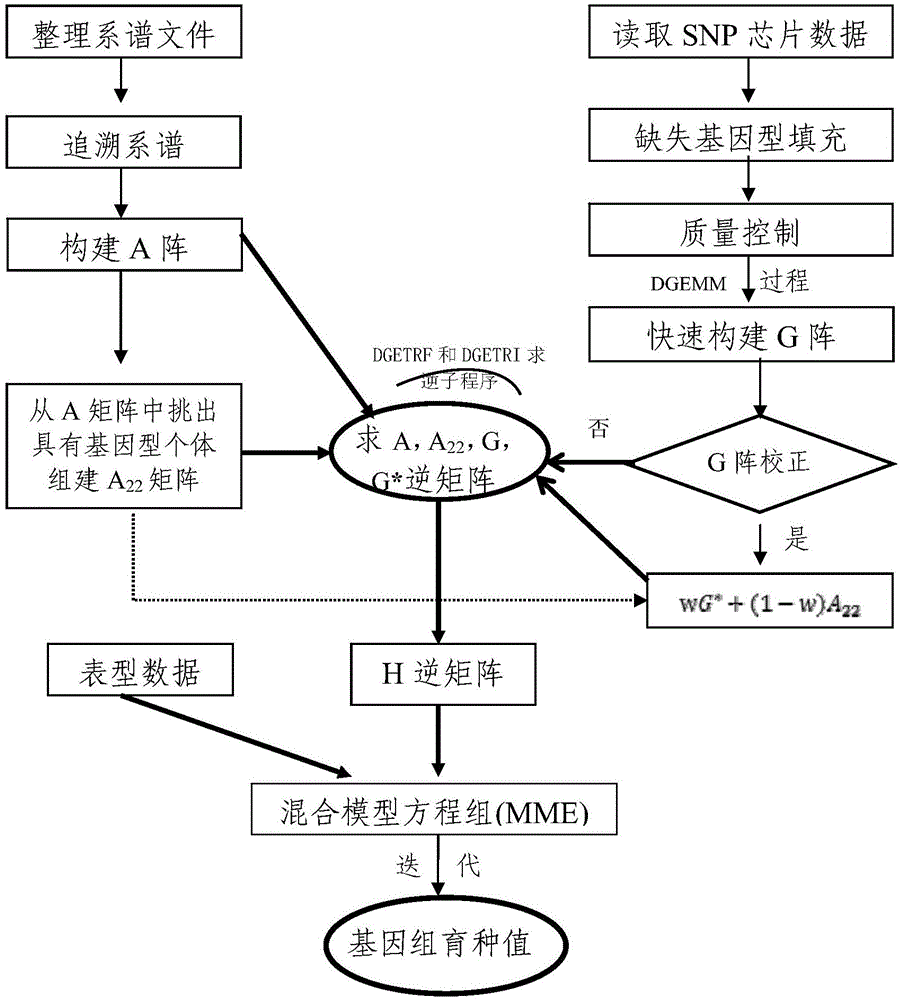
Method for rapidly evaluating genome breeding value and application
InactiveCN103914632ACalculation speedAdvantageSpecial data processing applicationsAnimal breedingHigh density
The invention belongs to the field of bioinformatics and provides a method for rapidly evaluating a genome breeding value. The method includes single nucleotide polymorphism (SNP) chip data edition; arrangement of genealogy files; construction of a matrix A by genealogy information, selection of individuals with SNP genotypes from the matrix A, and construction of a submatrix A22 according to matrix A elements among the individuals; construction of a matrix G by using high-density SNP chip information, and rapid construction of the matrix G by using a DGEMM process in a public open-source LAPACK math library; construction of a matrix H, calling of DGETRF and DGETRI matrixes in the public open-source LAPACK math library to obtain an inverse subprogram, rapid obtaining of inverse matrixes of the matrix A, the matrix A22 and the matrix G, and further solving of an inverse matrix G according to the partitioning inverse matrixes; solving of the genome breeding value of the individuals through a mixed model equation set. According to the method, calculation speeds for evaluating the genome breeding value by means of the comprehensive genealogy information and genome information can be accelerated, and applications of the genome selection to the field of animal breeding can be boosted.
Owner:CHINA AGRI UNIV
Vehicle seat-mounted bow holder
InactiveUS20080047992A1Easy to installEasy to removeMetal working apparatusStowing appliancesHead restraintWrap around
Owner:FABIAN BENJAMIN WADE
Method to split data operational function among system layers
InactiveUS10178073B2AdvantagePlatform integrity maintainanceSoftware simulation/interpretation/emulationData operationsOperating system
A method for accelerating data transfer and improving data storage in a computing environment is provided that includes splitting a function into two layers of an operating system to generate two separate sets of outcomes from the two layers. A set of outcomes from the two layers are combined so as to be imperceptible to a user save for faster operation. The splitting of the function is evaluated and optimized based on machine performance feedback. A computing system for communicating with a network is provided that performs this method. A bifurcated operating system affords additional advantages in performing function splitting.
Owner:TOUCANH LLC
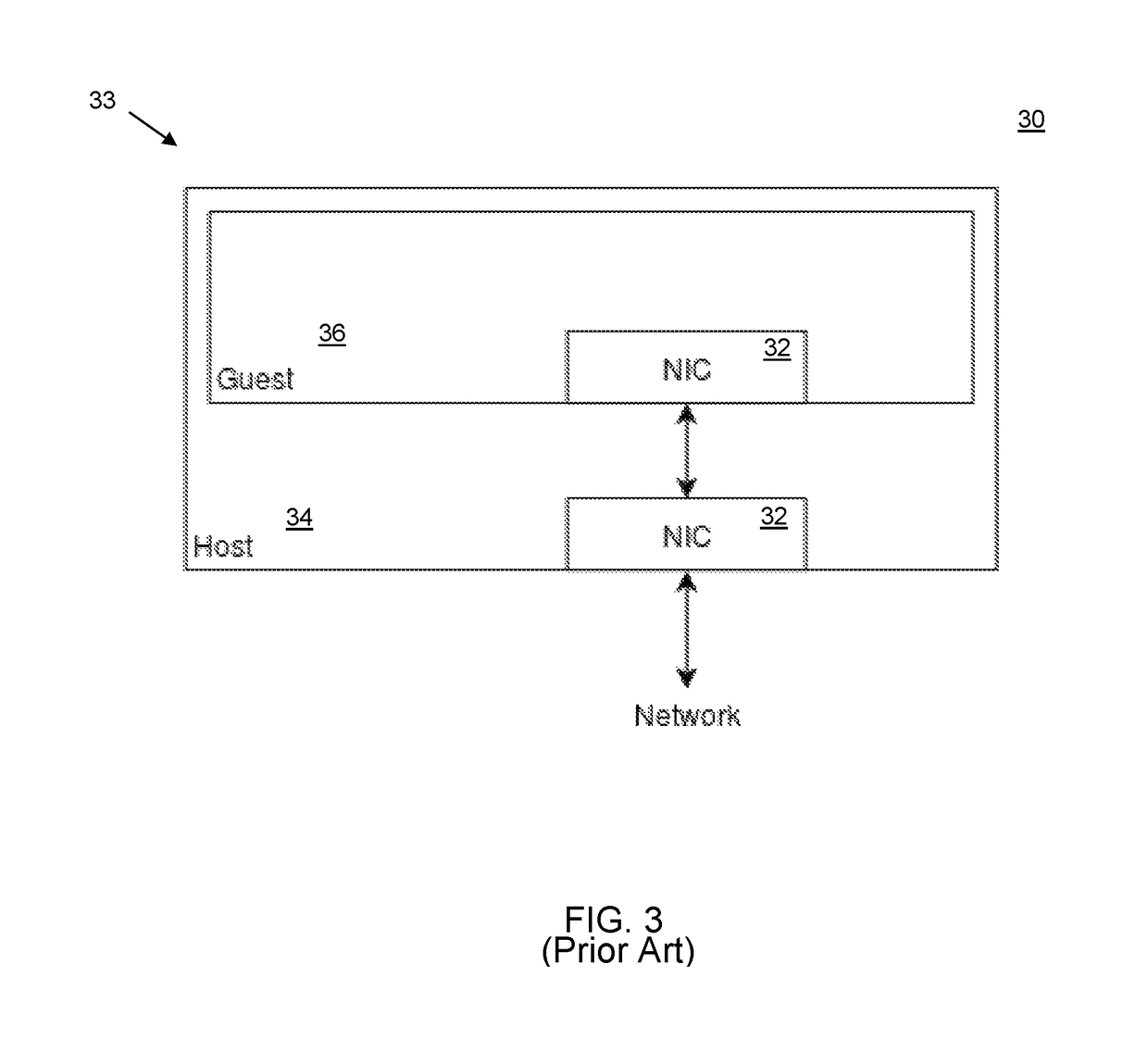
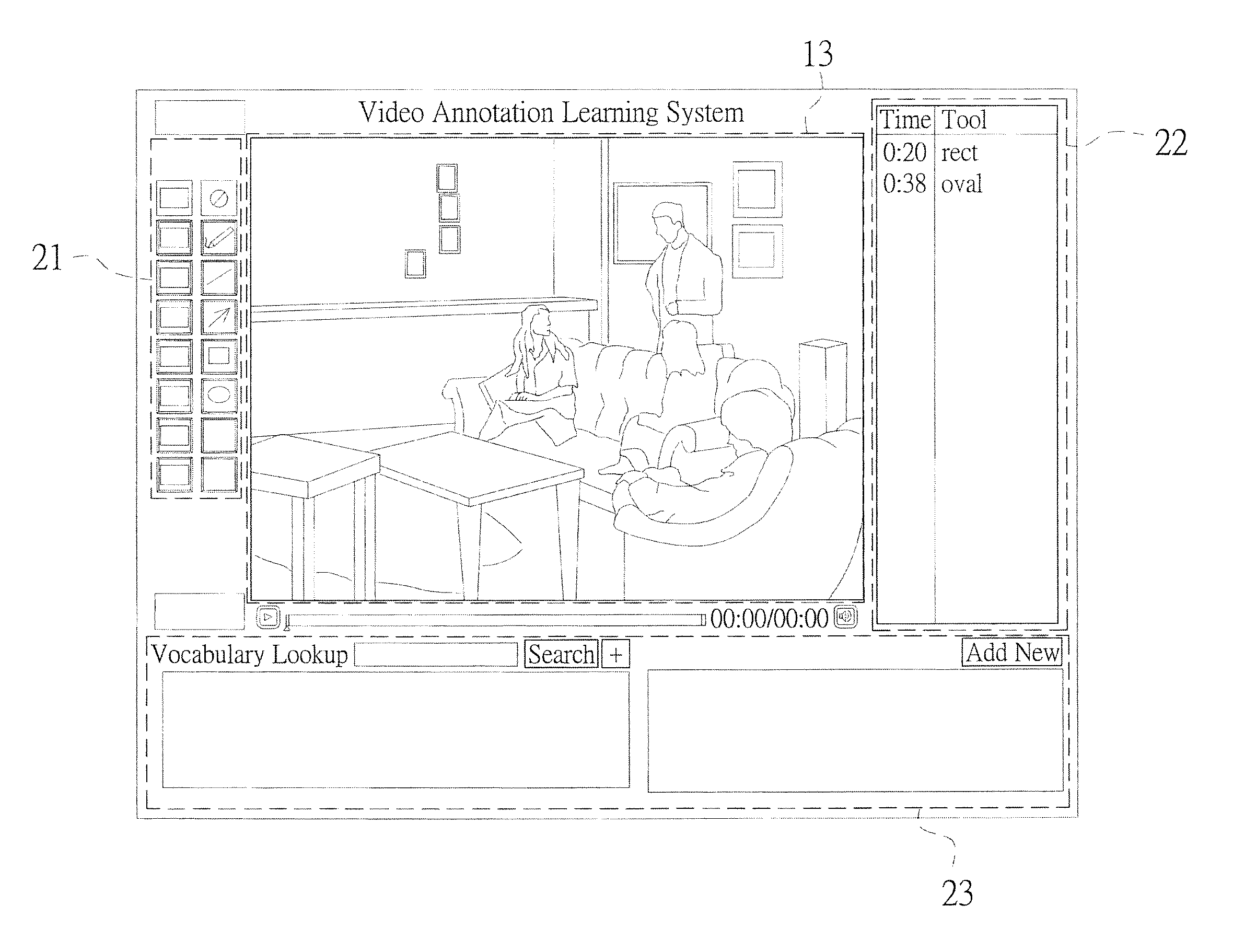
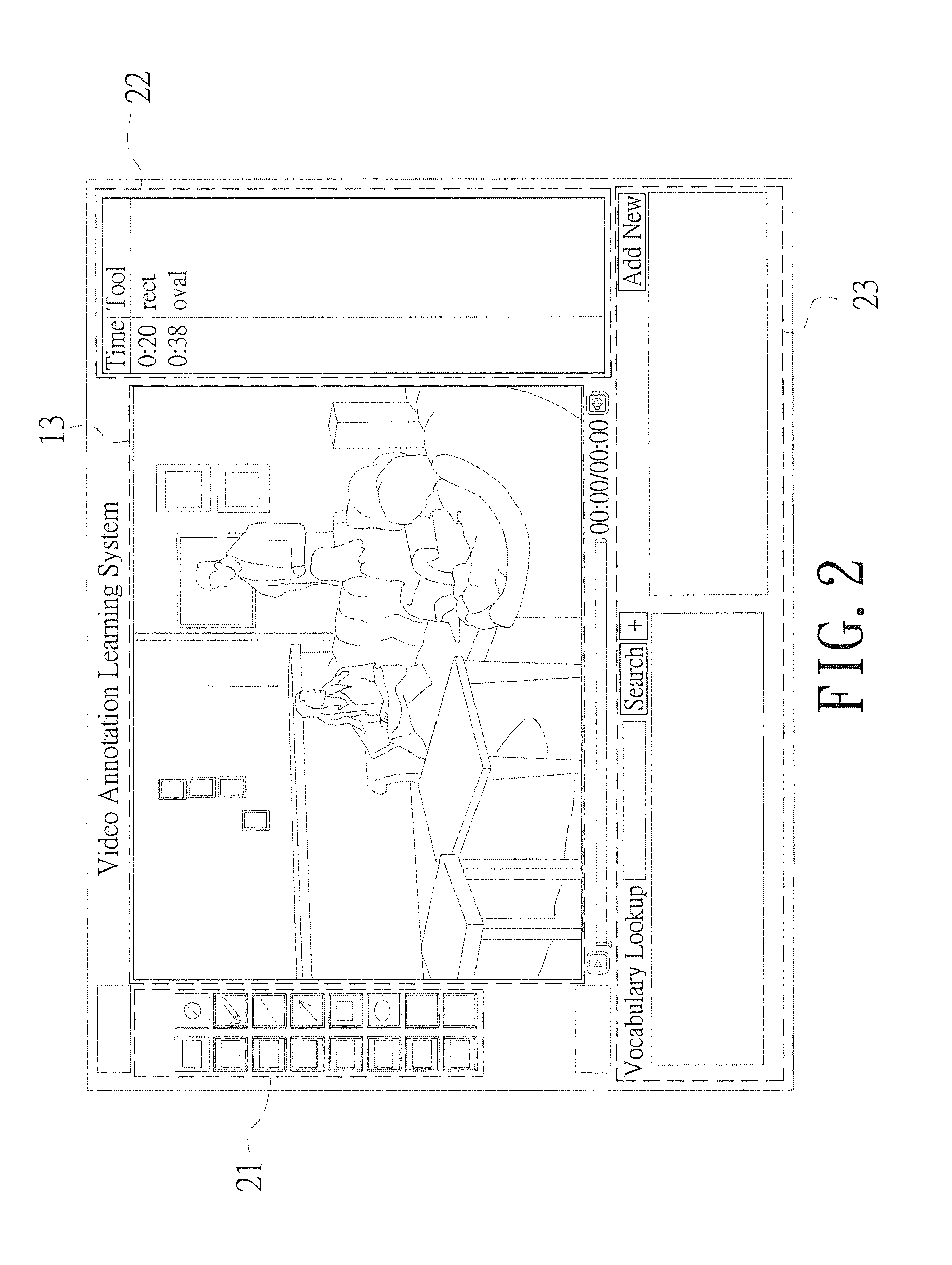
Real-time video annotation learning system and method for the same
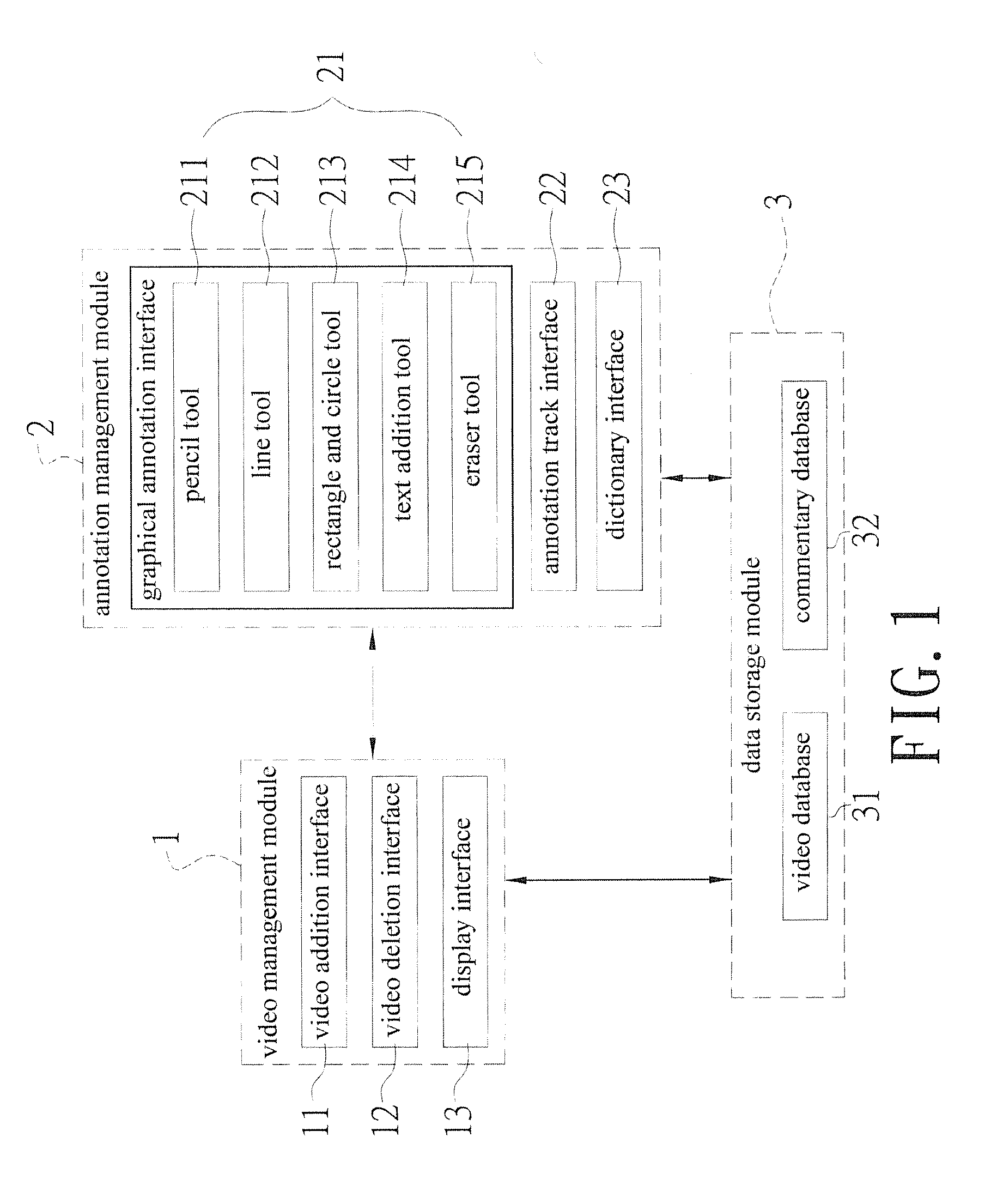
InactiveUS20140335502A1Solve the real problemAdvantageMaps/plans/chartsElectrical appliancesGraphicsVerbal learning
A real-time video annotation learning system and a method for the same are revealed. The real-time video annotation learning system includes a video management module, an annotation management module and a data storage module. A language learner can select videos stored in the data storage module by the video management module and add comments to a location of an image in the video at a specific time while the comments are in the form of text or graphs built in a commentary management module. Thus the comments are shown on the image in the form of text or graphs at a specific time. Users can grasp key points of learning content the video provided exactly. Together with system functions of real-time word look-up and word note, user's learning effect is significantly improved and the video has an advantage in language learning.
Owner:NAT CHENG KUNG UNIV
Polyisocyanate modified with sulphamic acid, preparation method thereof and use thereof
ActiveUS20160280836A1Good water dispersibilityLong application periodPolyurea/polyurethane coatingsCross-linkAdhesive
A polyisocyanate modified with sulphamic acid and a mixture thereof, the preparation method thereof, and the use thereof in the production of polyurethane, especially as a cross-linking ingredient in the field of aqueous coatings and adhesives containing groups that are capable of reacting with isocyanate groups.
Owner:WANHUA CHEM GRP CO LTD +2
High formability super strength cold-roll steel sheet or steel strip, and manufacturing method therefor
InactiveUS20170298466A1High elongationHole expansion rateFurnace typesHeat treatment furnacesUltimate tensile strengthBalance performance
A high formability super strength cold-roll steel sheet or steel strip, and a method for manufacturing same. The steel sheet or steel strip comprises the following ingredients by weight percent: C 0.15% to 0.35%, Si 1.0% to 2.0%, Mn 1.6% to 2.6%, Mo 0.1% to 0.4%, P≦0.02%, S≦0.004%, N≦0.005%, Nb 0.015% to 0.04%, Ti 0.02% to 0.06%, Al 0.015% to 0.045%, B 0.0003% to 0.001% and B≧P % / 30, and the balance being Fe and inevitable impurities. The steel sheet or steel strip has a tensile strength greater than or equal to 980 MPa, an elongation rate greater than or equal to 15% and a hole expansion rate greater than or equal to 40%, and has balanced performance.
Owner:CHINA BAOWU STEEL GRP CORP LTD
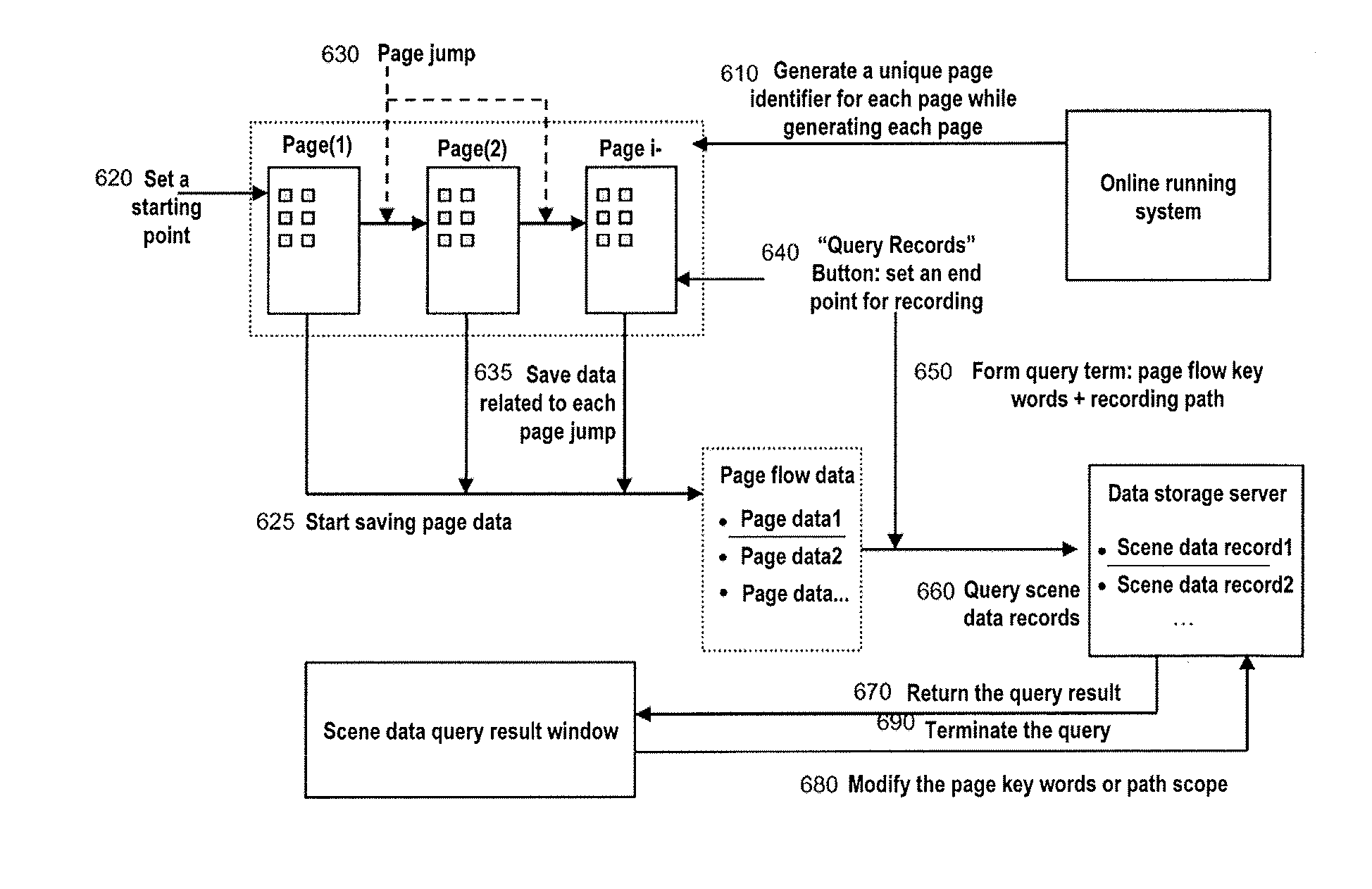
Method and system of saving and querying context data for online applications
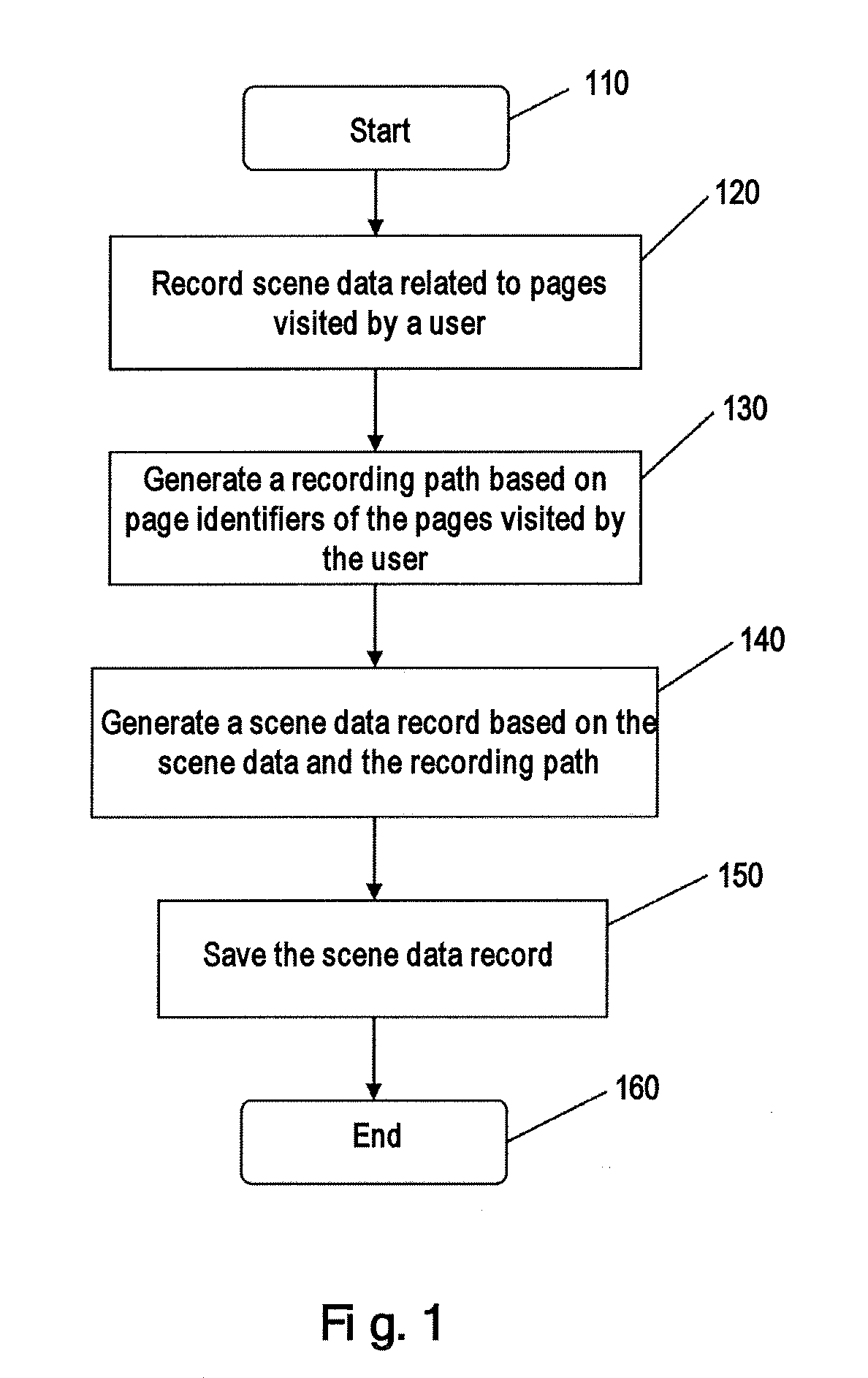
InactiveUS20110106841A1The implementation process is simpleAdvantageDatabase queryingDigital data processing detailsPath generationContext data
The present invention provides methods and systems for saving and querying context data for an online application. The context data of an online application related to pages visited by a user are collected, where the context data is associated with page identifiers of the visited pages. A step-by-step path is generated based on the page identifiers of the pages visited by the user, and a context data record is generated and saved based on the collected context data and the step-by-step path. According to the methods and systems, a query term is further generated using on the collected context data and the step-by-step path, for performing query to the context data. By applying the methods and systems of the present invention to different contexts, the user is able to easily save and later reference the previous actual running data of some functional contexts.
Owner:IBM CORP
Multipurpose Exercise Training Device
InactiveUS20170050065A1Easily transformed in configurationEasy to transformDumb-bellsSkipping-ropesJumping ropeMuscle strength
An exercise device having many advantageous features is described, including the ability to provide users with multiple modes of training by transforming the device into different configurations. Users can exercise their cardiovascular endurance by using the device in its jump rope configuration. Users can also engage in muscular resistance training using the same device configured to one of multiple possible resistance training configurations. Resistance training configurations provide a method of suspending users' bodyweight such that they can train their muscular strength capacity by exerting themselves against the force of gravity. Resistance can be selected from nearly zero resistance to a user's full body weight, with the ability to easily adjust between exercises and between users, and the ability to easily transform the device between configurations to provide for versatility and ease-of-use. The device includes an inelastic length member with two clip assemblies and two handles and / or grips at both ends.
Owner:CARTER MATTHEW RODERICK
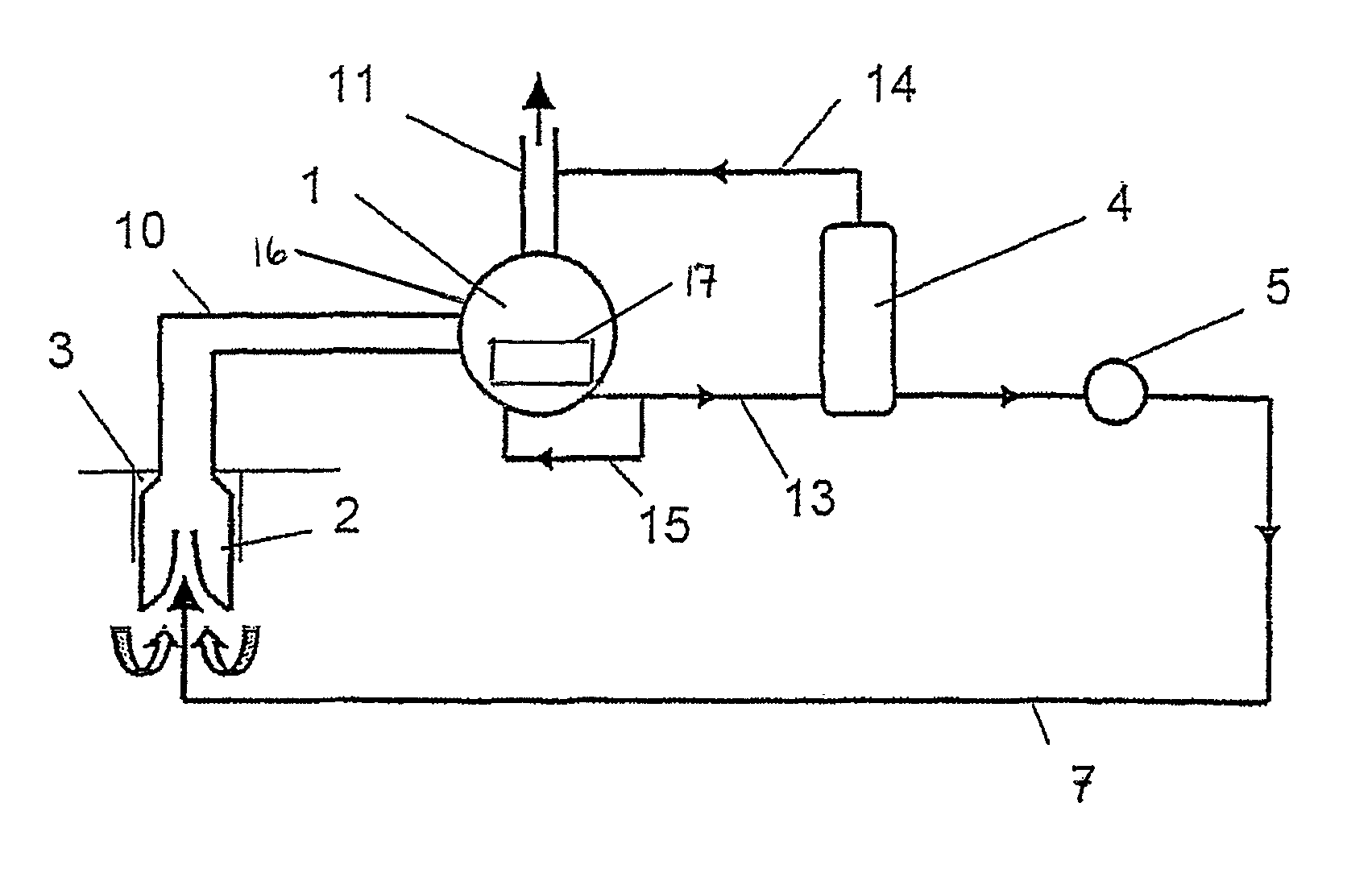
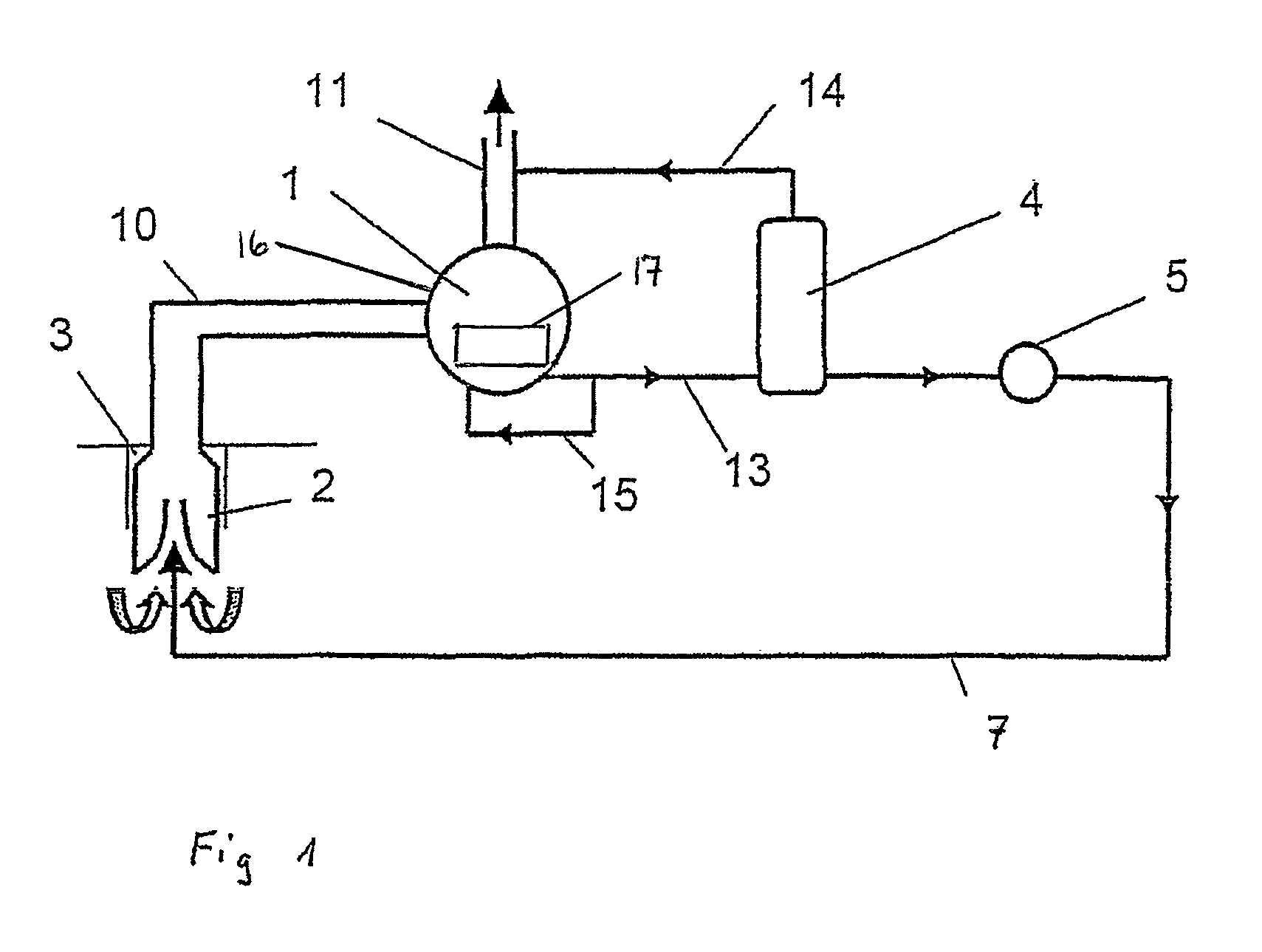
Method for delivering a multi phase mixture and pump installation
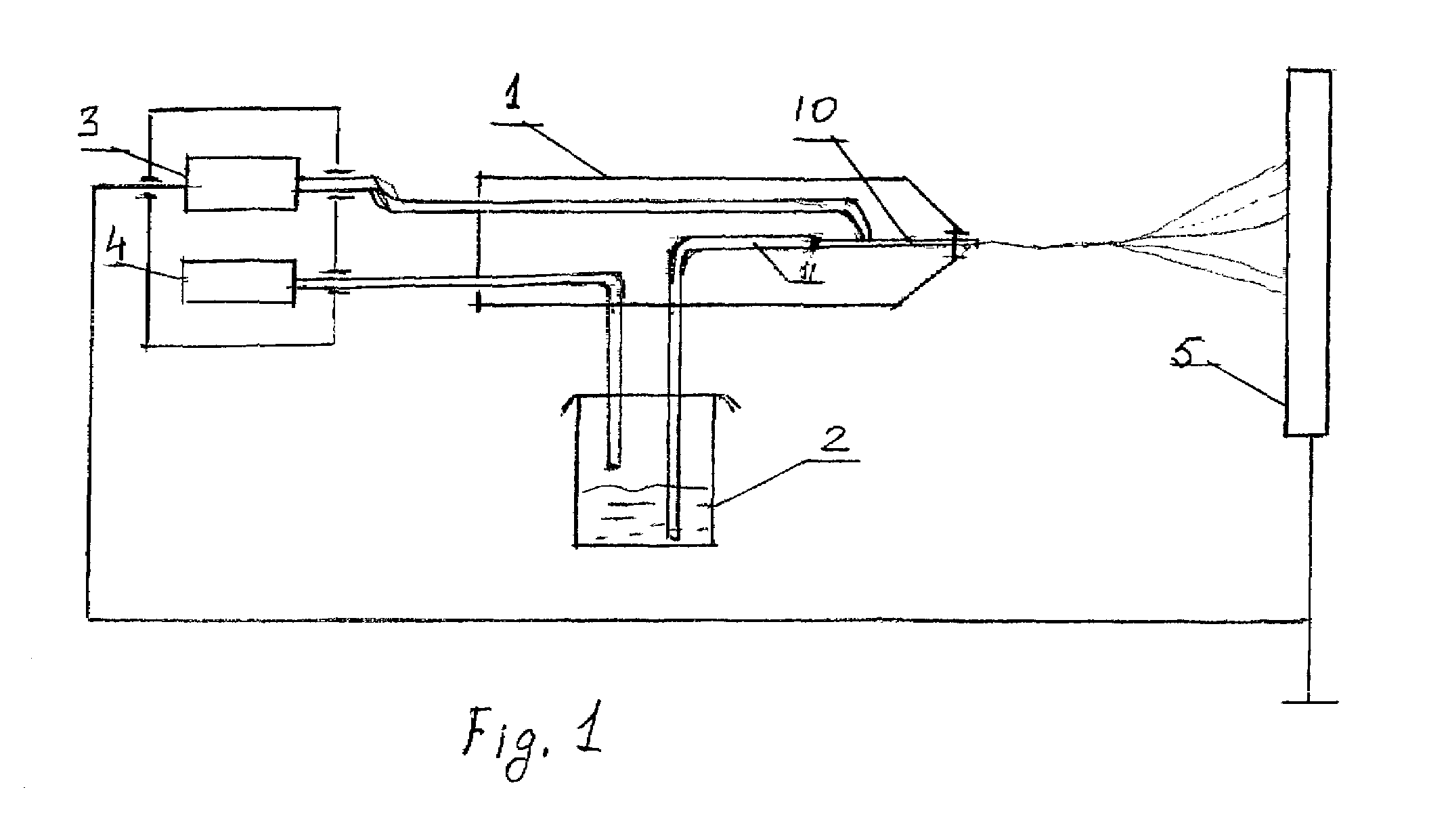
The object of the invention is to improve delivery of the multi-phase mixture, in particular hydrocarbons from a well, and to limit the free gas volume. According to the invention, this object is attained in that a partial liquid flow (13) is split off on the pressure side from the main delivery flow and guided to the high-pressure side of at least one ejector pump (2) arranged on the suction side as an auxiliary delivery device. The pump installation provides a feed line (7) connecting the pressure chamber of the displacement pump (1) with the high-pressure side of at least one ejector pump (2), whereby the ejector pump (2) is arranged on the suction side in the delivery direction of the displacement pump (1).
Owner:JOH HEINRICH BORNEMANN
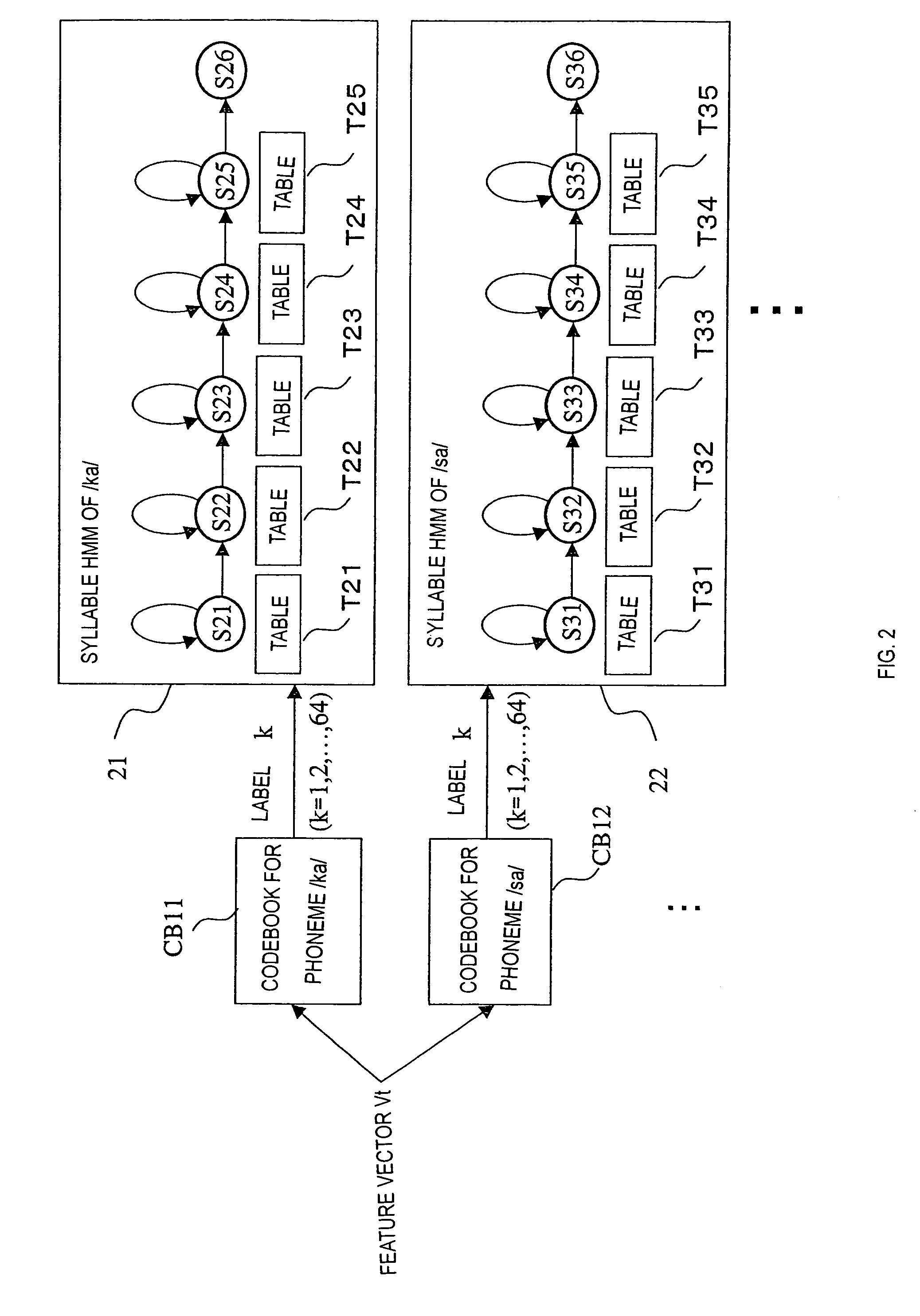
Method of calculating HMM output probability and speech recognition apparatus
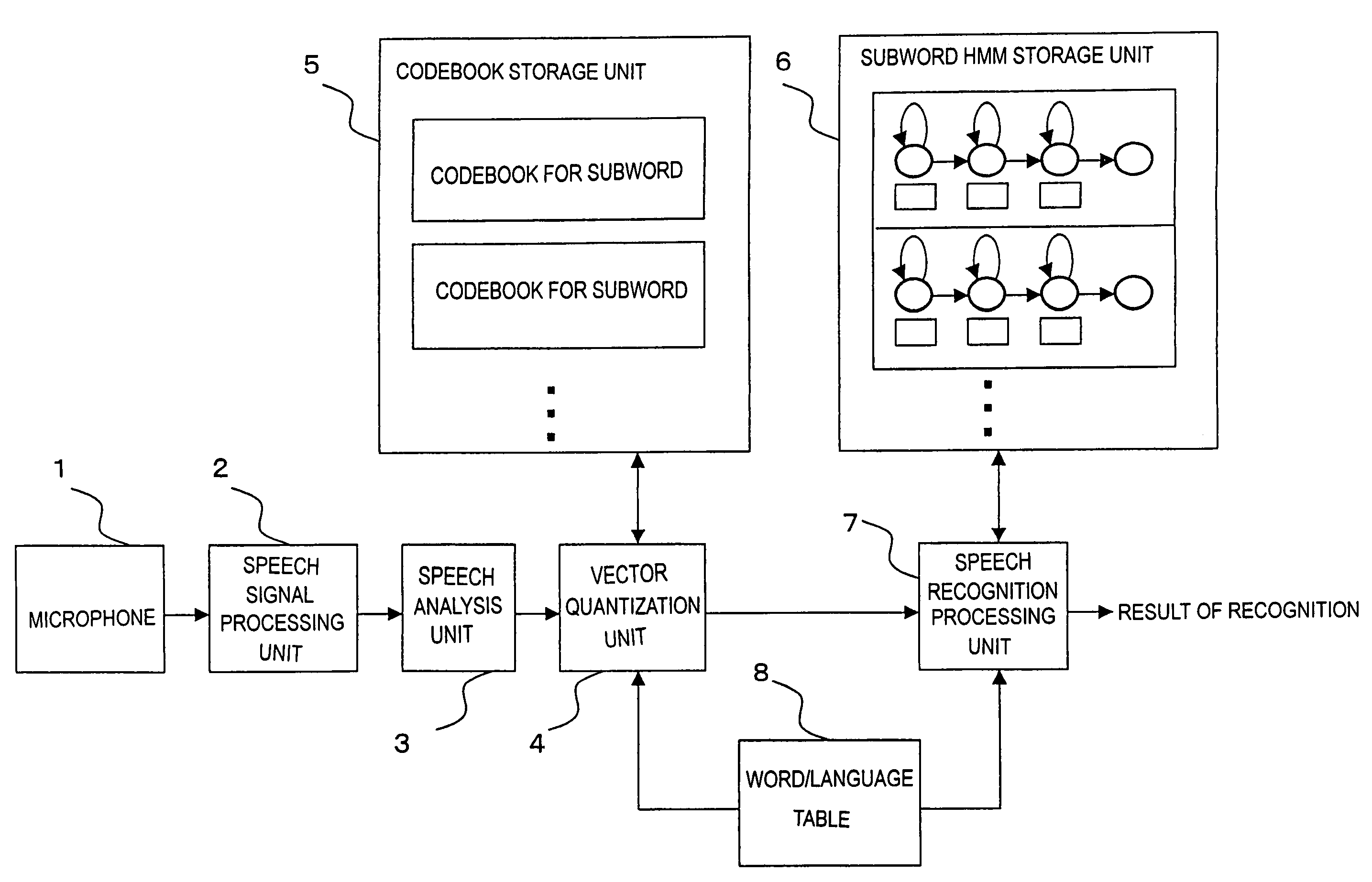
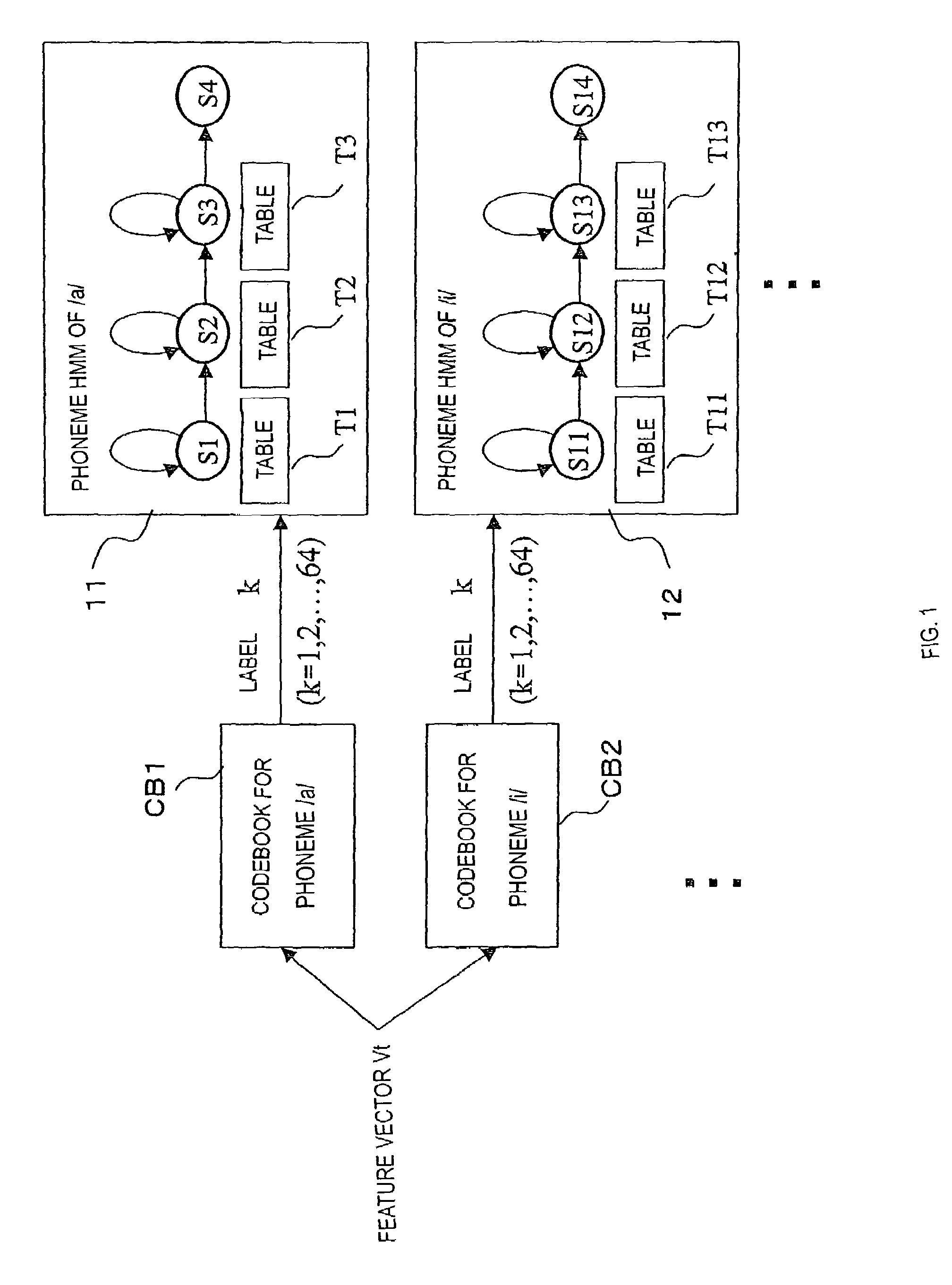
InactiveUS7058576B2Reduce in quantityReduce data sizeSpeech recognitionSpeech identificationSpeech analytics
The invention relates to speech recognition based on HMM, in which speech recognition is performed by performing vector quantization and obtaining an output probability by table reference, and the amount of computation and use of memory area are minimized while achieving a high ability of recognition. Exemplary codebooks used for vector quantization can be provided as follows: if phonemes are used as subwords, codebooks for respective phonemes, such that a codebook CB1 is a codebook for a phoneme / a / and a codebook CB2 is a codebook for a phoneme / i / , and these codebooks are associated with respective phoneme HMMs. When a feature vector obtained by speech analysis is vector quantized based on, for example, the codebook CB1 and a code (label) is output, tables for respective states of the phoneme HMM associated with the codebook CB1 are each referred to in order to obtain state output probabilities corresponding to the label, and speech recognition is performed using the state output probabilities as a parameter.
Owner:SEIKO EPSON CORP
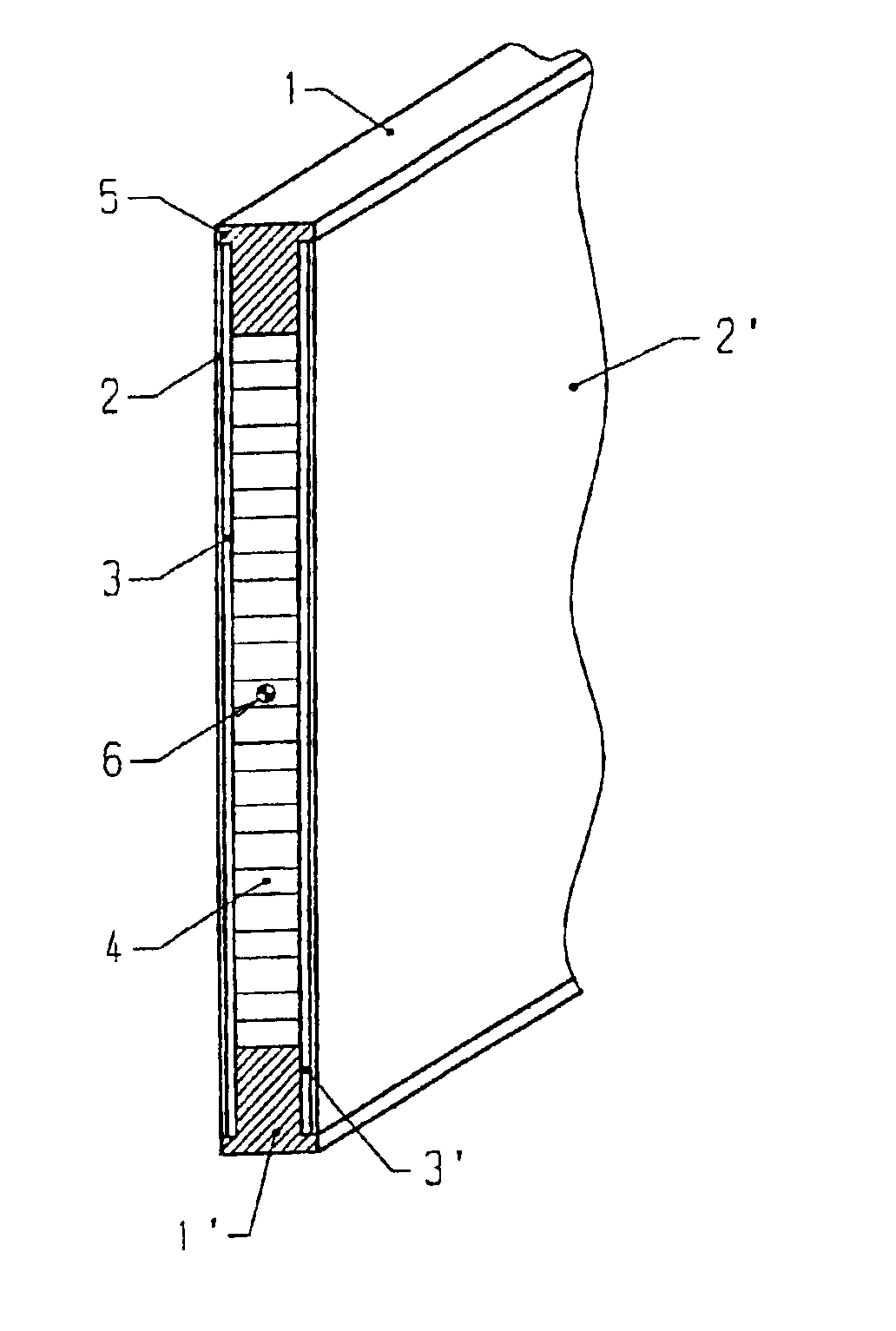
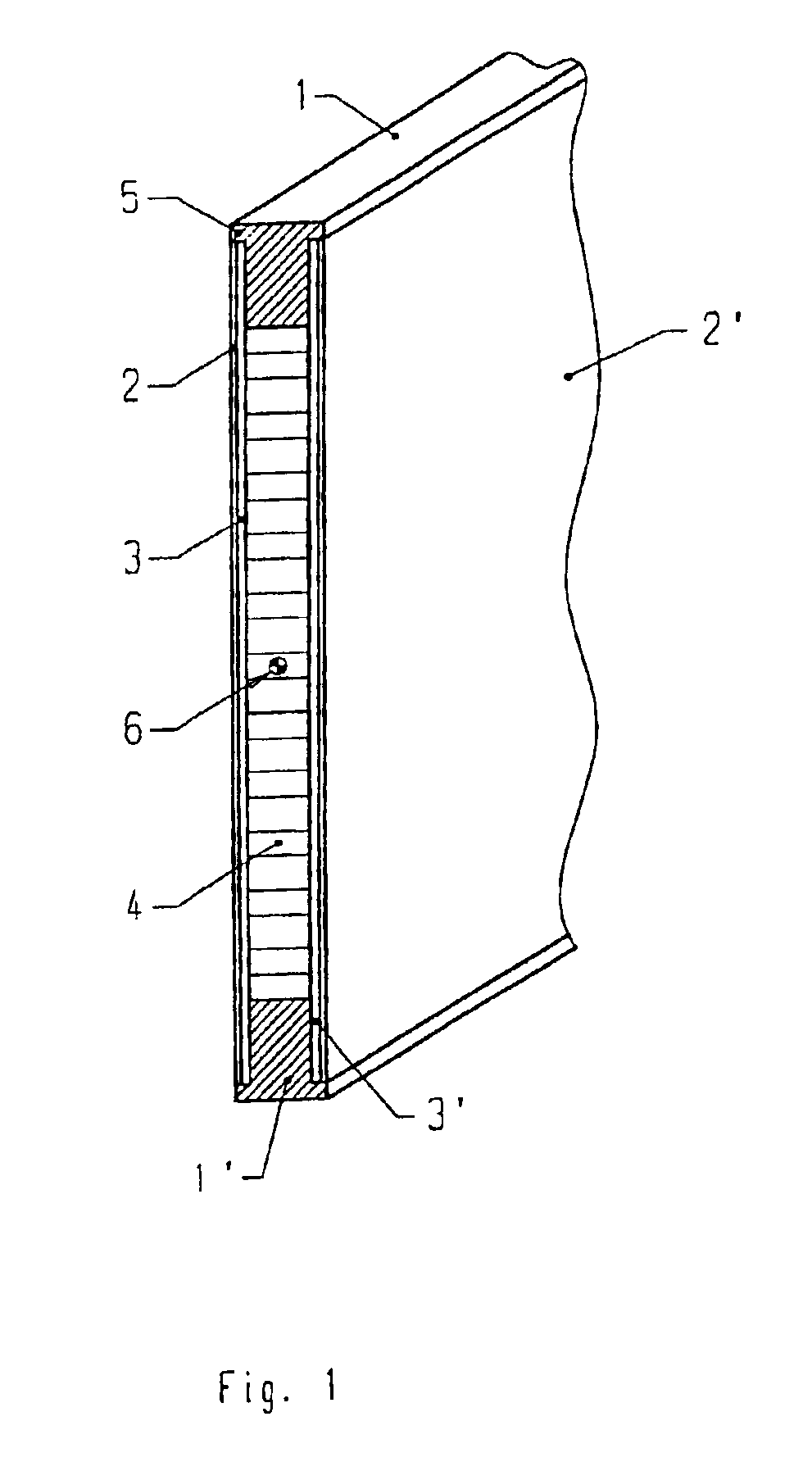
Support element for a heddle frame
InactiveUS6994123B2Material requirementHigh elastic modulusHealdsOther shedding mechanismMetallic materialsEngineering
Owner:GROZ BECKERT KG
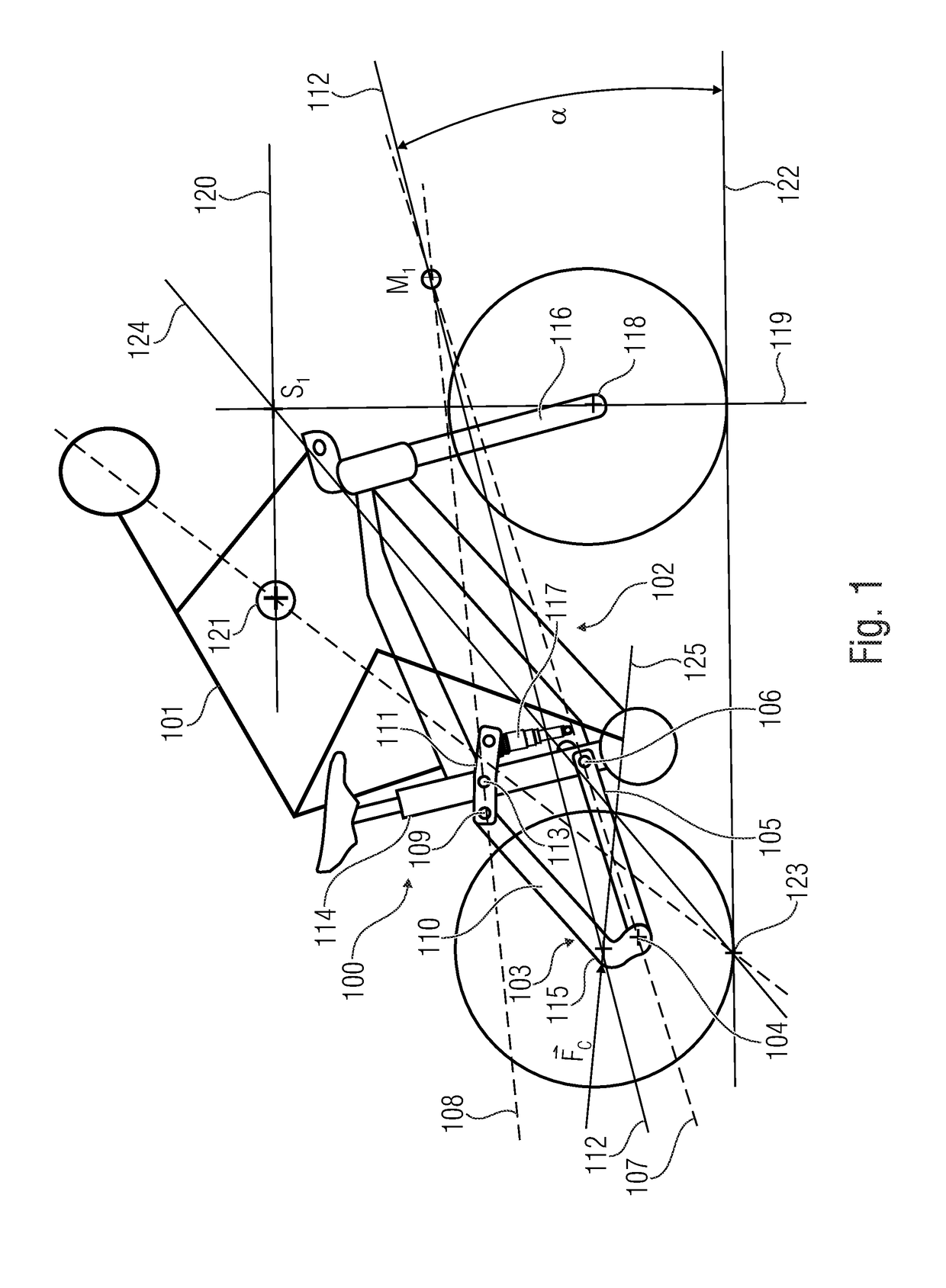
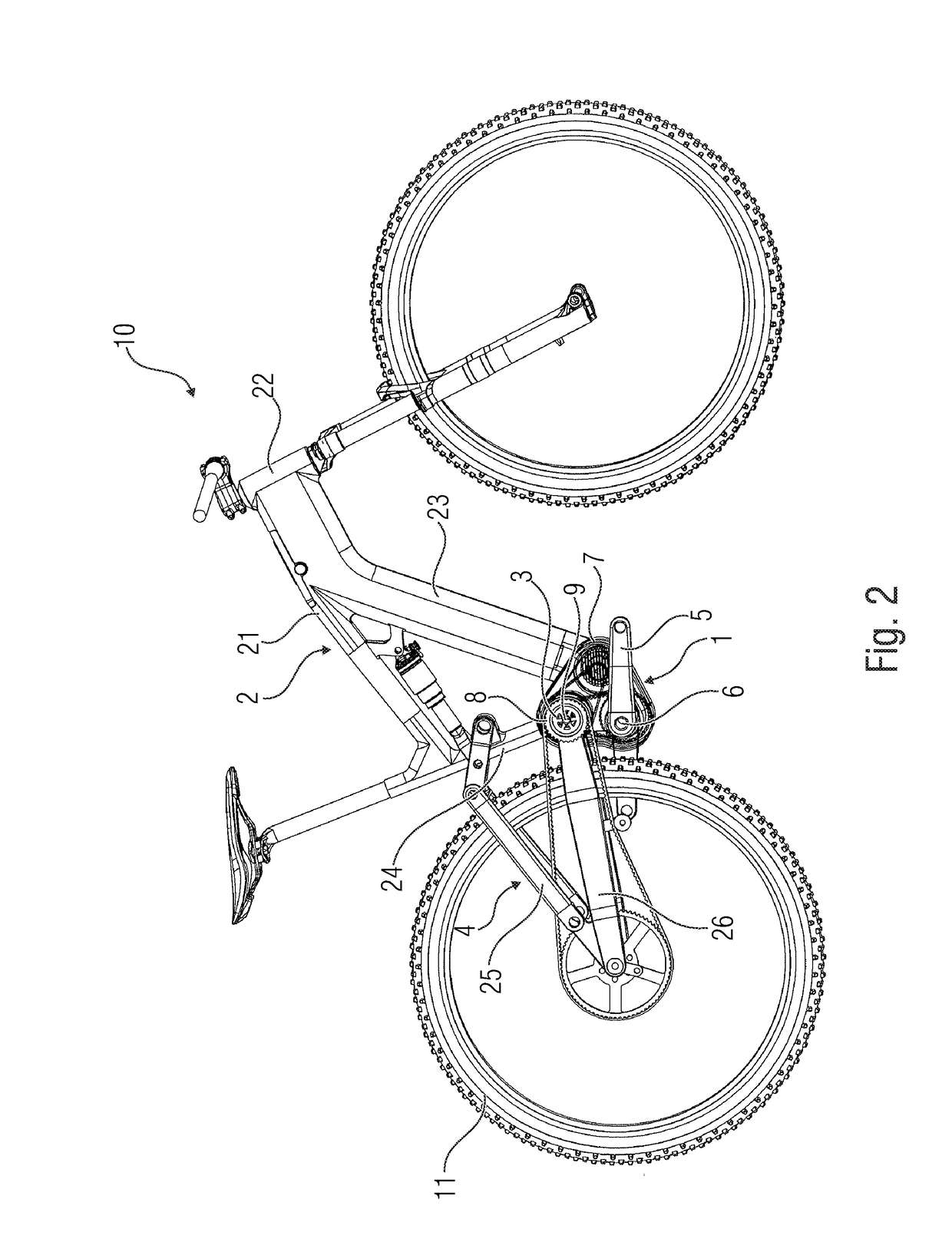
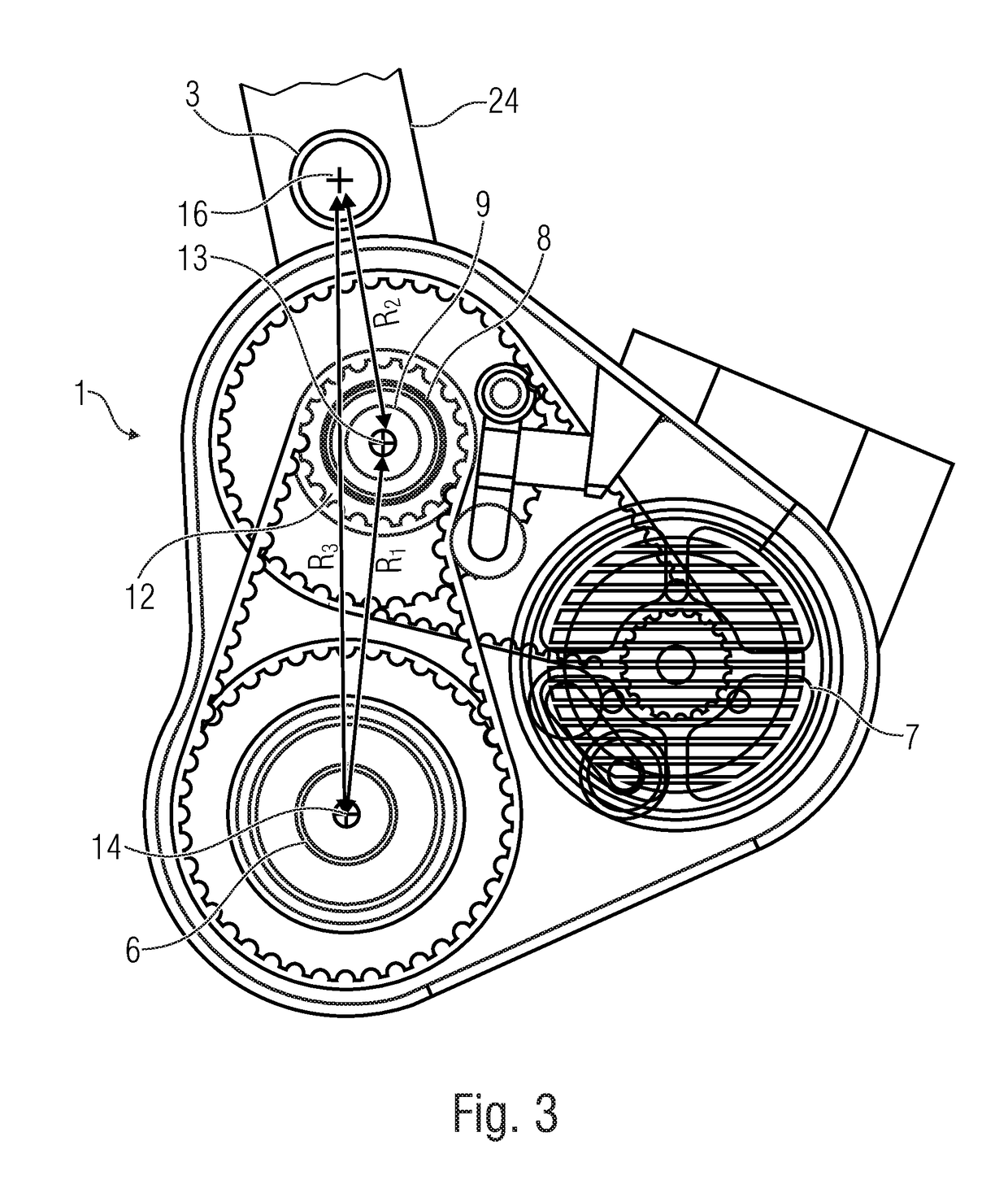
Drive device for a bicycle driven by an electric motor
ActiveUS20190031278A1Strong decompressionClosely arrangedChain/belt transmissionAxle suspensionsDrive shaftEngineering
What is disclosed is a drive device for a bicycle driven by an electric motor, having a main frame having a swing arm bearing, and a rear linkage element arranged at the swing arm bearing. The drive device has a pedal crank as a first drive for providing a first drive force, having a first drive shaft, a center electric motor as a second drive for providing a second drive force, and an output element having an output shaft, configured to receive the first and / or second drive forces and transfer same to the bicycle wheel to be driven. The center axis of the output shaft is radially distanced from the center axis of the first drive shaft and the output shaft is arranged relative to the swing arm bearing such that a radial distance between the center axis of the swing arm bearing and the center axis of the output shaft is smaller than a radial distance between the center axis of the swing arm bearing and the center axis of the first drive shaft.
Owner:QCS QUALITY CONSULT SERVICE GMBH
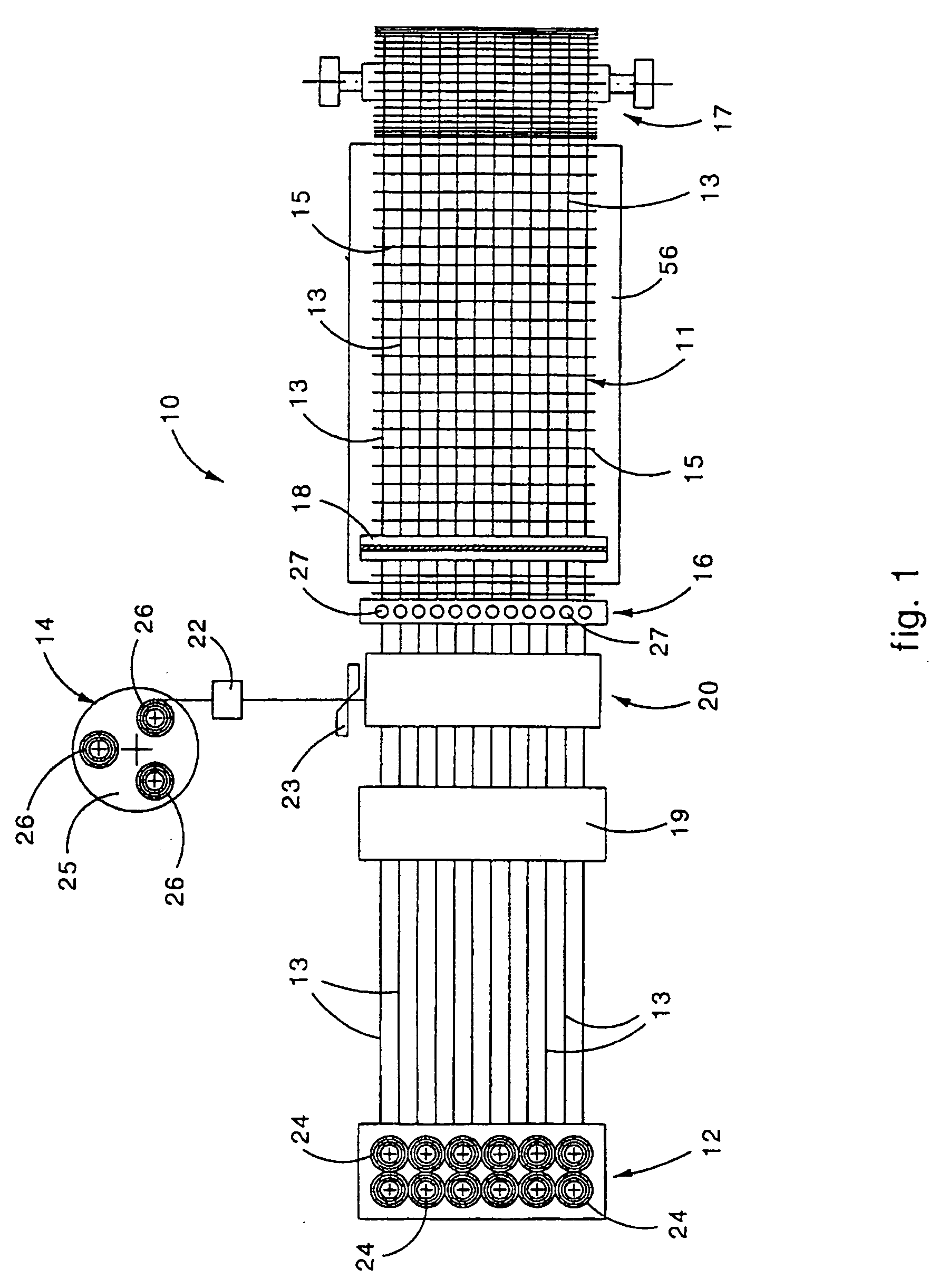
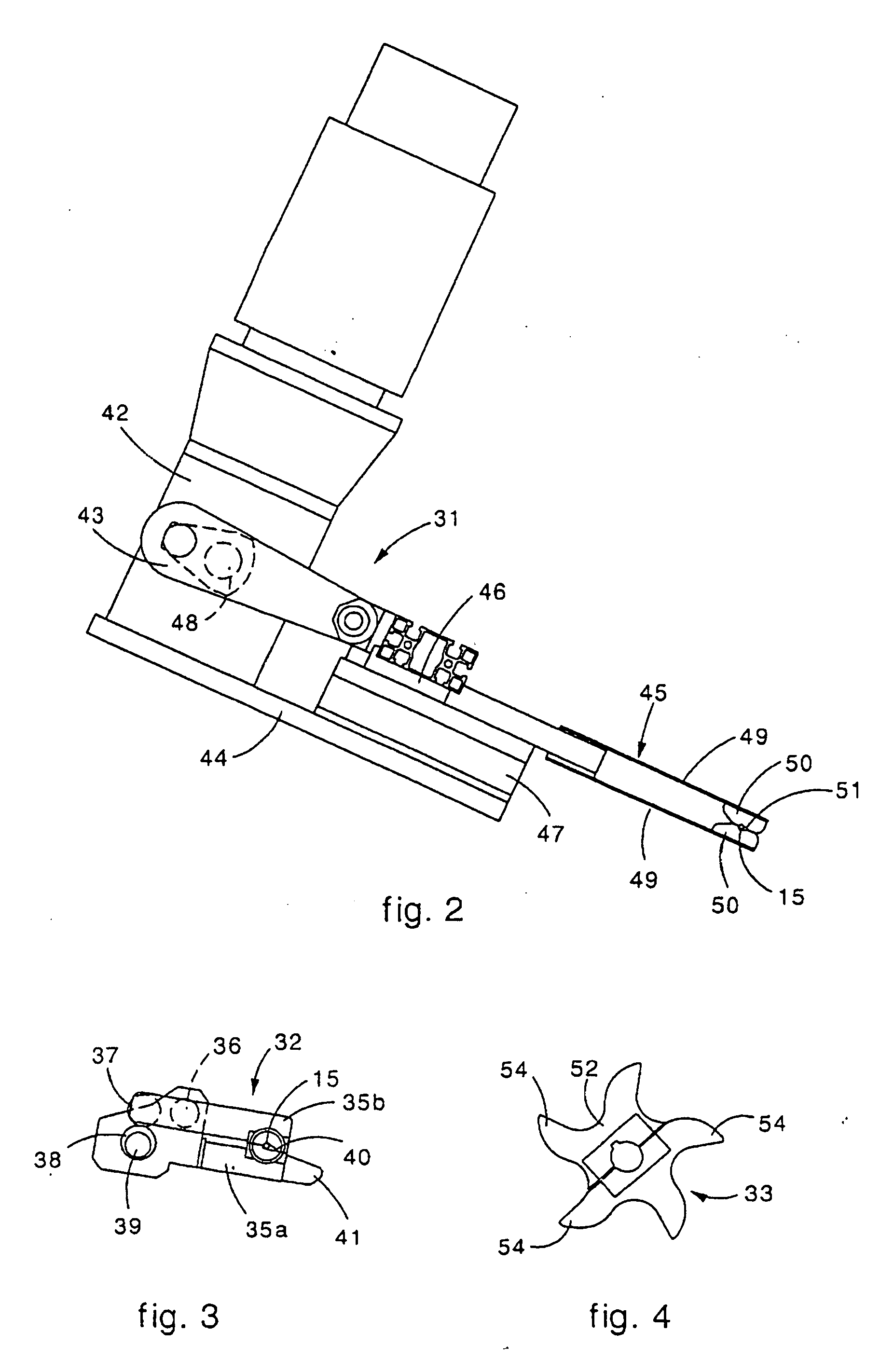
Machine for the formation of metal mesh and relative method
Machine for the formation of a metal mesh obtained by attaching together a plurality of longitudinal metal wires to a plurality of transverse metal wires. The machine includes a first feed assembly to make the longitudinal wires advance step-wise, a second feed assembly to arrange one transverse wire at a time in a first preparation position, a positioning apparatus to arrange the transverse wire in a second attachment position, and a welding assembly to attach the transverse wires to the longitudinal wires. The positioning apparatus includes a loading assembly, provided with a gripping and transfer device, to locate the transverse wire in the second attachment position. The machine also includes a thrusting device to take the transverse wire from the first preparation position to a third intermediate pick-up position, near the second attachment position, wherein the transverse wire is picked up by the gripping and transfer device. The thrusting device includes a rotary element provided with blade able to thrust the transverse wire from the first preparation position to the third intermediate pick-up position.
Owner:BETA SYSTEMS
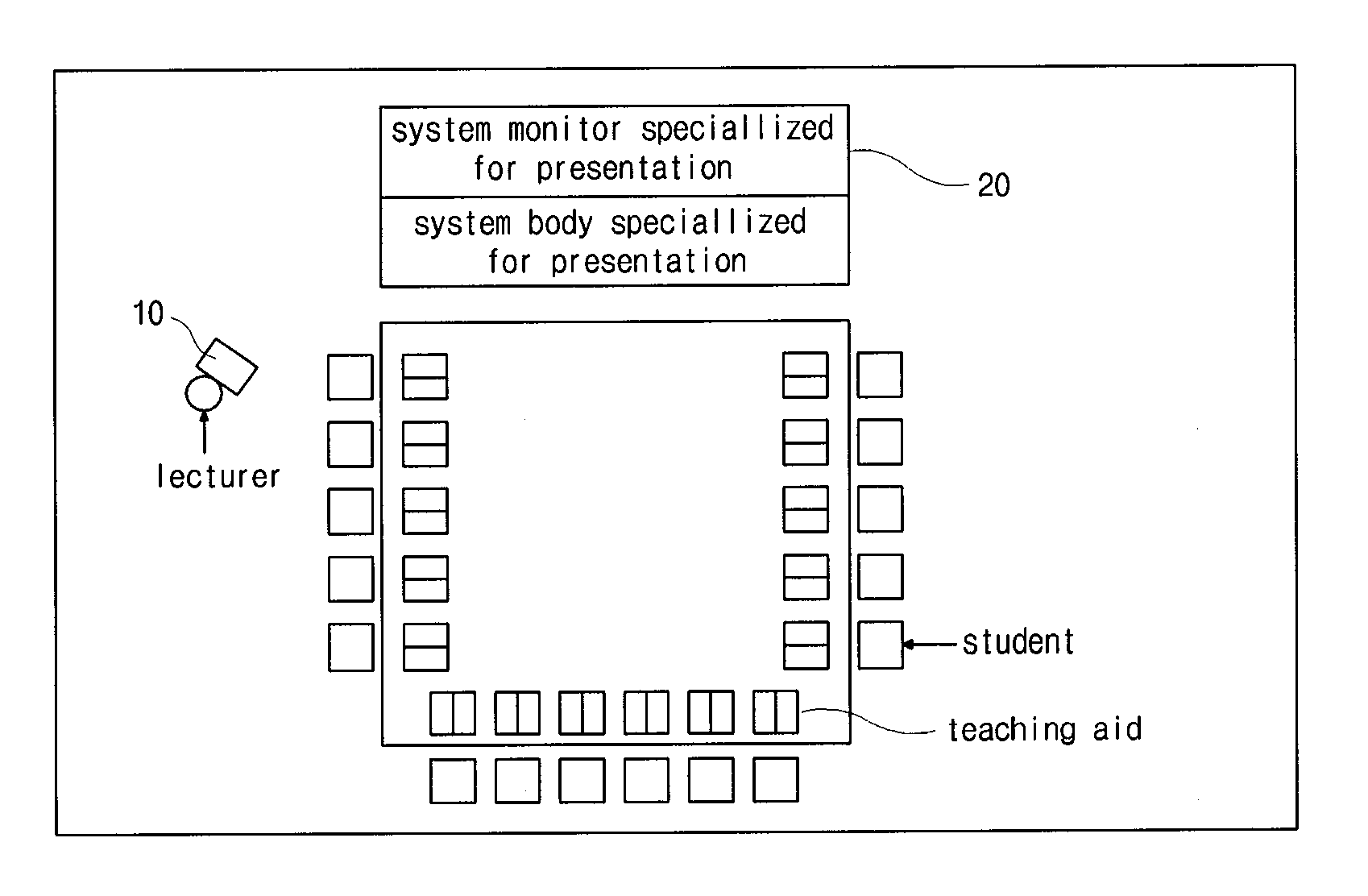
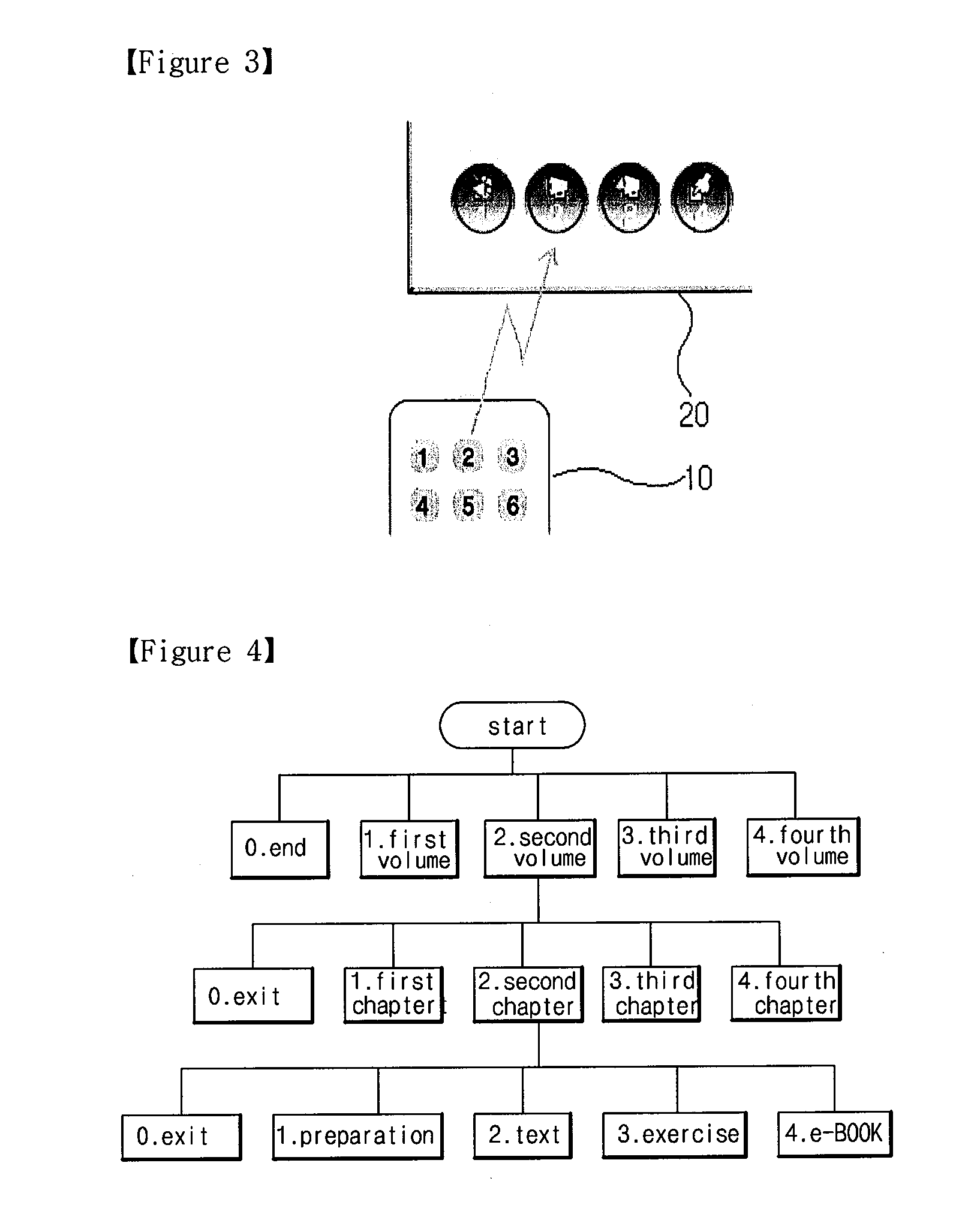
Method of controlling computer program and searching fold and file using object-oriented button, control and presentation system using the same and remote controller using the same
InactiveUS20110020782A1AdvantageQuickly and correctly searchedInput/output for user-computer interactionTelevision system detailsRemote controlControl signal
The present invention is to provide a method of controlling a program executed on a computer using a remote control apparatus and a method of searching a folder or a file using the method of controlling the program. The method of controlling the program according to the present invention includes defining a plurality of object linking button groups in the remote control, each having an identification mark; arranging a plurality of objects each having the same identification mark as one of the plurality of object linking button groups, wherein each of the objects has a text or an image each including an execution information or a data content; assigning and interlocking the execution information with objects, respectively; transmitting a control signal to a computer by selecting one object linking button of the remote control apparatus; and selecting the execution information interlocked with corresponding object to the one object linking button after the program receives the control signal. In the present invention, since data or a file is quickly selected using an object linking button, there is advantages in processing a dynamic presentation. Moreover, a desired presentation method is easily selected and executed.
Owner:KO YUN YONG
Light emitting diode chip
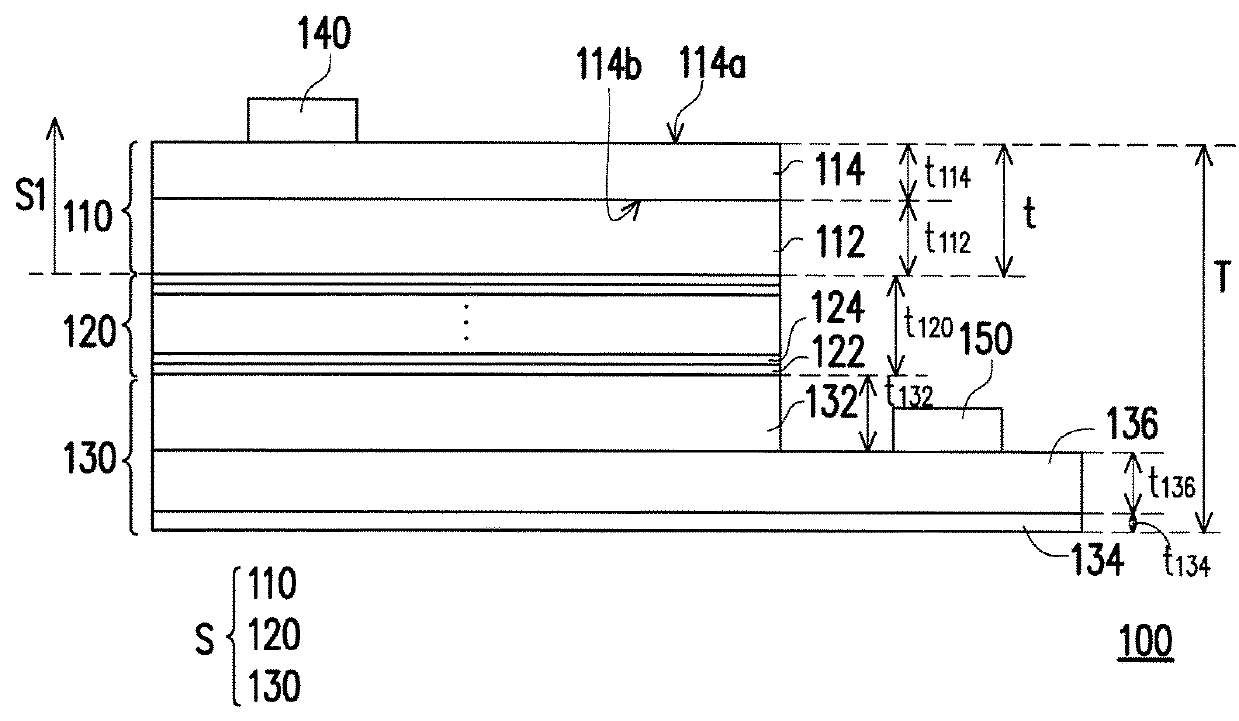
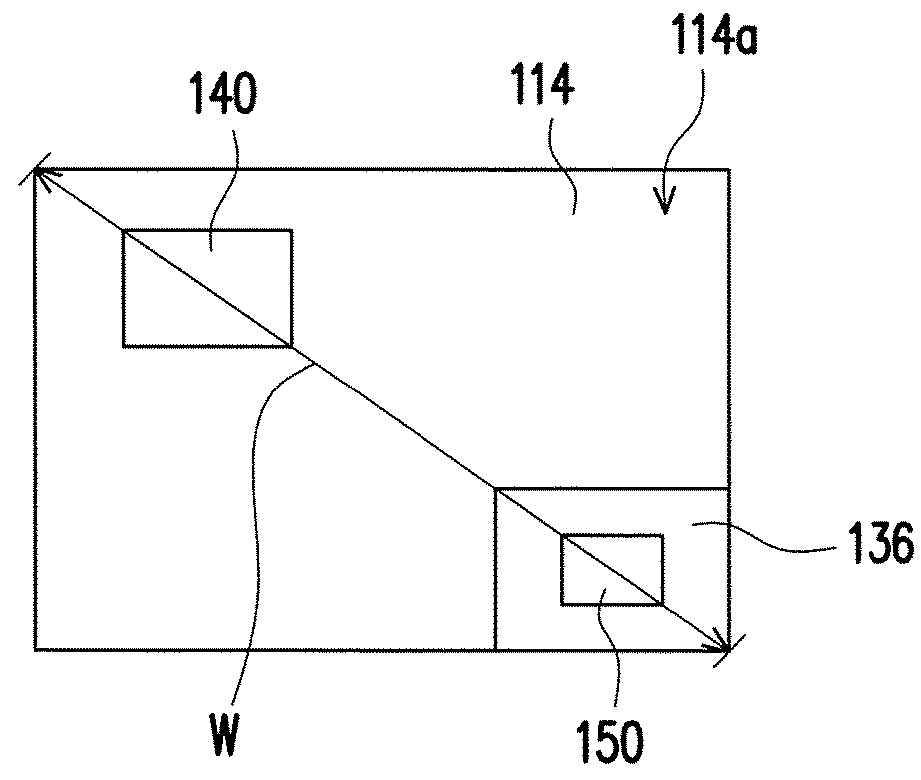
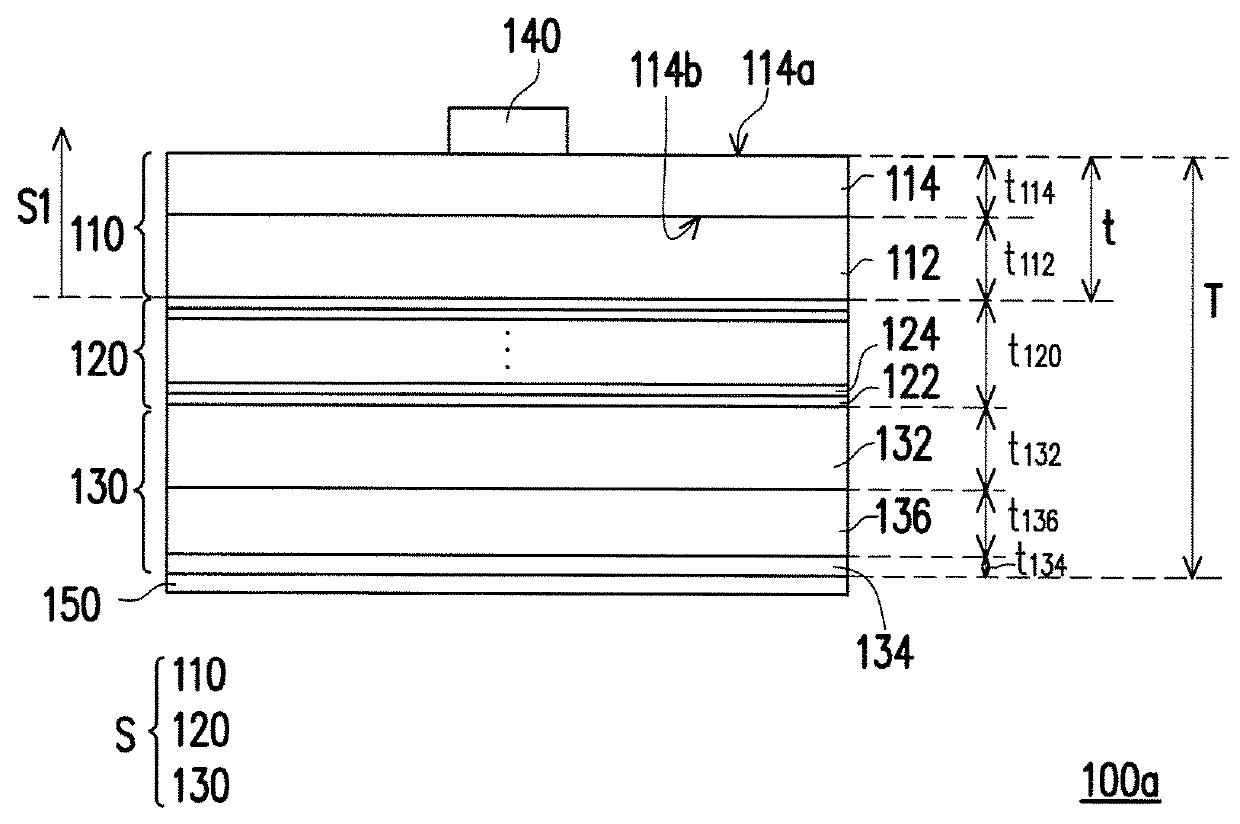
ActiveUS20180166607A1Improve performanceReduce areaSemiconductor devicesLight-emitting diodeSemiconductor
A light-emitting diode chip including a p-type semiconductor layer, a light-emitting layer and an n-type semiconductor layer is provided. The light-emitting layer is disposed between the p-type semiconductor layer and the n-type semiconductor layer. A ratio of a sum of thicknesses of all semiconductor layers of the light-emitting diode chip over a maximum width of the light-emitting diode chip ranges from 0.02 to 1.5. A ratio of a sum of thicknesses of all semiconductor layers located in a side of the light-emitting layer toward the p-type semiconductor layer over the sum of thicknesses of all semiconductor layers of the light-emitting diode chip ranges from 0.05 to 0.2.
Owner:PLAYNITRIDE
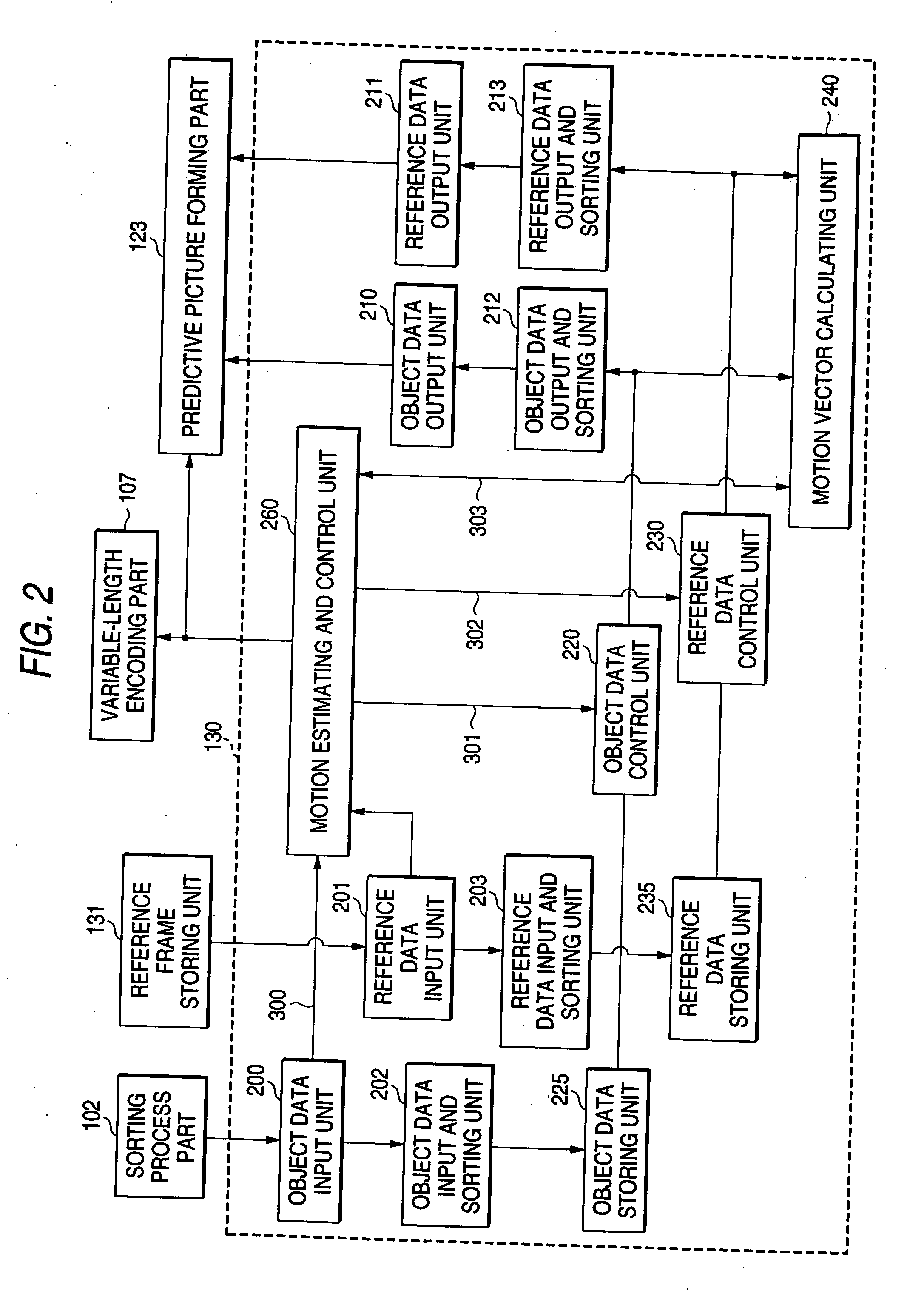
Motion vector estimating method and motion picture processor
ActiveUS20060039474A1AdvantageIncrease speedTelevision system detailsImage analysisPattern recognitionComputer graphics (images)
A motion vector estimator includes an object data input and sorting unit and a reference data input and sorting unit for sorting packed input picture data to separate first picture data used for estimating the motion vector from second picture data that is not used for estimating the motion vector and pack the picture data again respectively, an object data storing unit and a reference data storing unit for storing the repacked first picture data and the second picture data, and a motion vector calculating unit for reading the first picture data from the object data storing unit and the reference data storing unit to perform a calculation for estimating the motion vector.
Owner:PANASONIC SEMICON SOLUTIONS CO LTD
Bacteriorhodopsin protein variants and methods of use for long term data storage
ActiveUS8883719B2Enhanced protein performanceGeneration densityPeptide/protein ingredientsNanoinformaticsLong term dataWild type
Bacteriorhodopsin protein variants and methods using the bacteriorhodopsin variants for performance in holographic and three-dimensional (3D) memory storage devices are described. The amino acid and chemical modifications of bacteriorhodopsin provided herein achieve greatly enhanced protein performance. The memory storage devices write, read and erase data proficiently. The bacteriorhodopsin protein variants are useful in optical memory storage and associative processor systems. Irradiation of the light-sensitive protein with light of known wavelength causes the protein to switch between different states. The variants enter the branched photocycle via a single or a two photon process and form the permanent ‘Q’ state more efficiently than the wild-type bacteriorhodopsin protein. This branching photocycle of the variants is exploited in the fabrication of 3D memory storage devices.
Owner:UNIV OF CONNECTICUT
Features
- R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
Why Patsnap Eureka
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Social media
Patsnap Eureka Blog
Learn More Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com