Patents
Literature
Hiro is an intelligent assistant for R&D personnel, combined with Patent DNA, to facilitate innovative research.
100results about How to "Mechanically robust" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
High density electrical connector with variable insertion and retention force
ActiveUS8182289B2Lower insertion forceImprove retentionTwo-part coupling devicesCoupling protective earth/shielding arrangementsElectricityHigh density
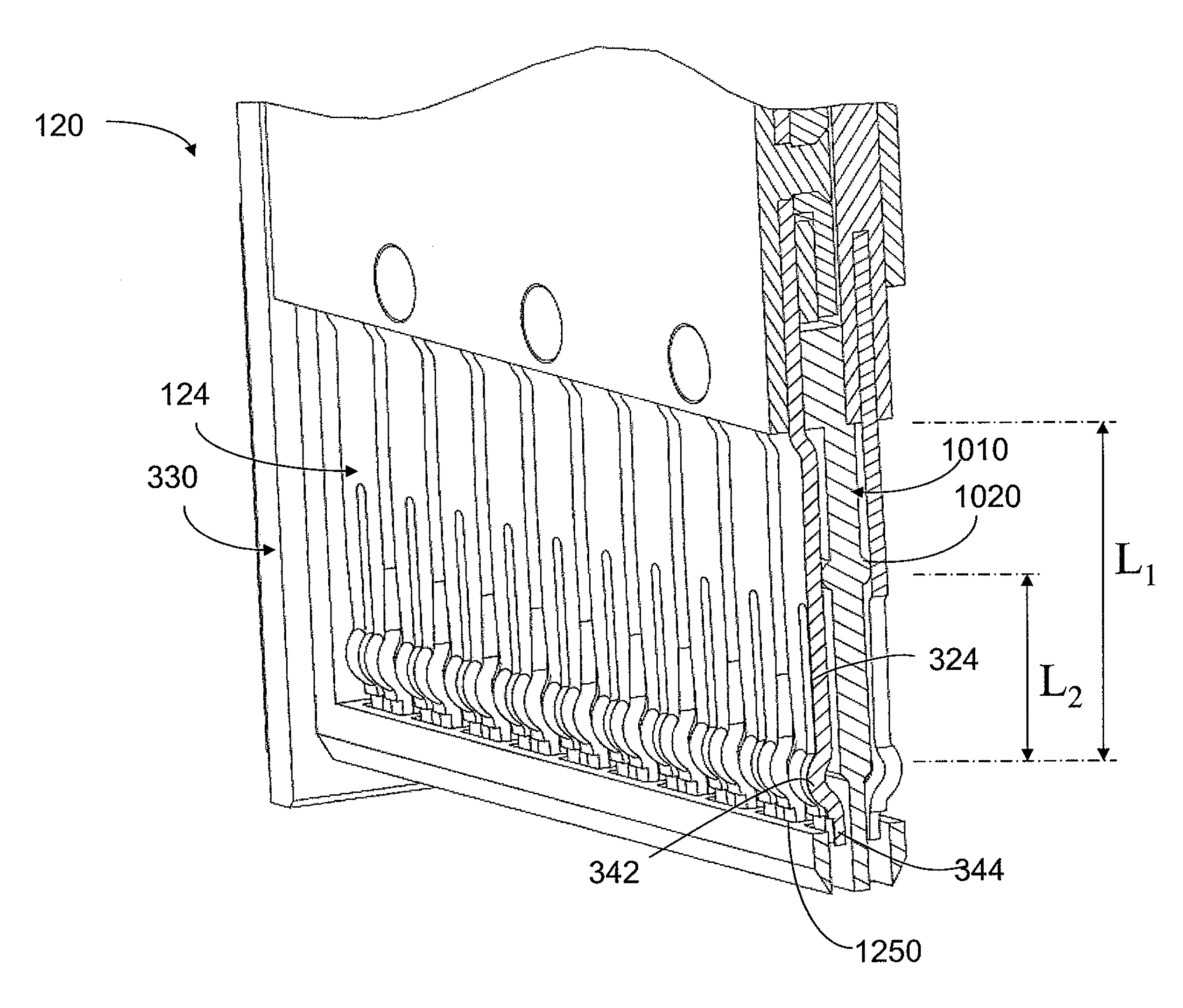
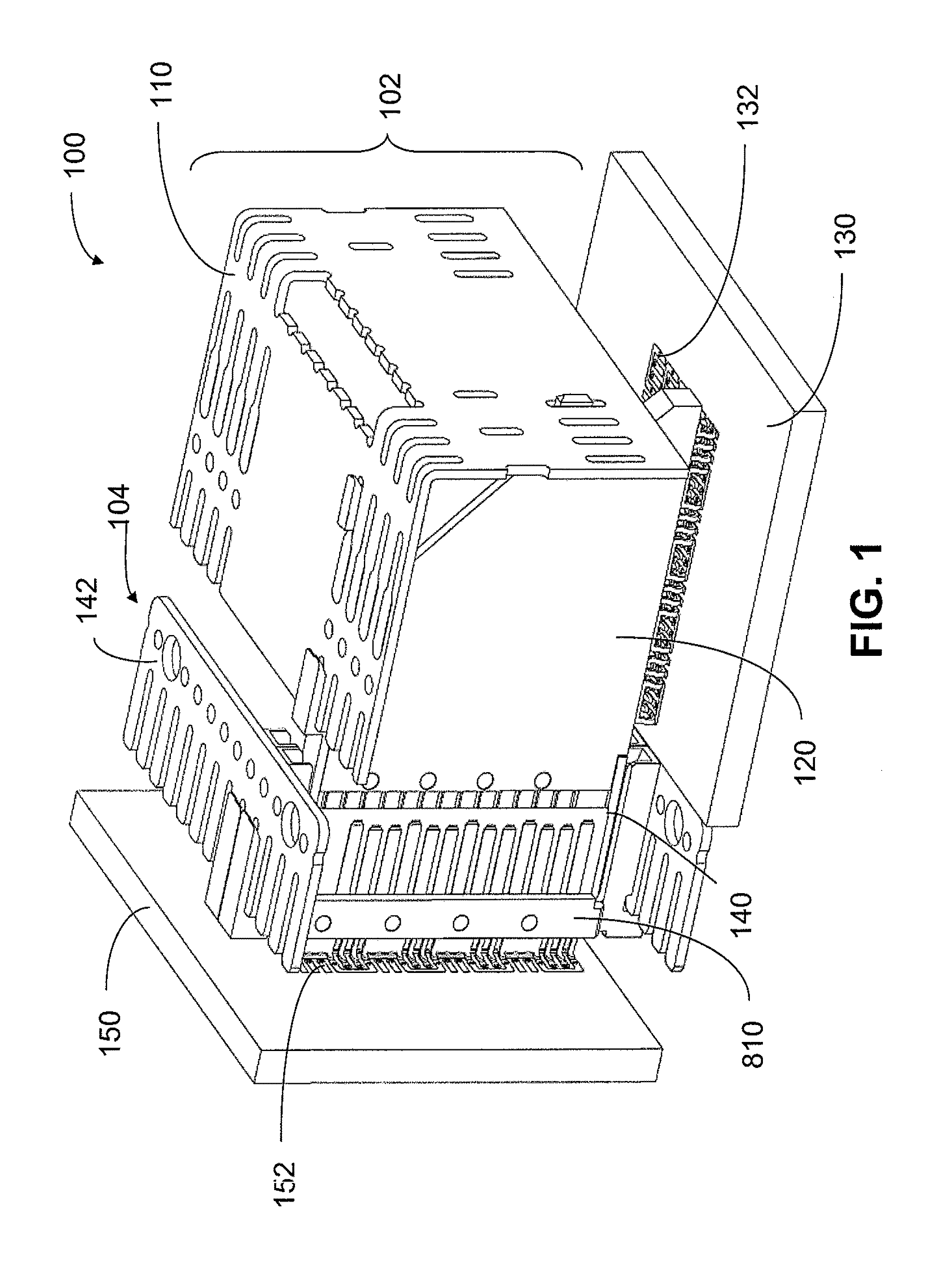
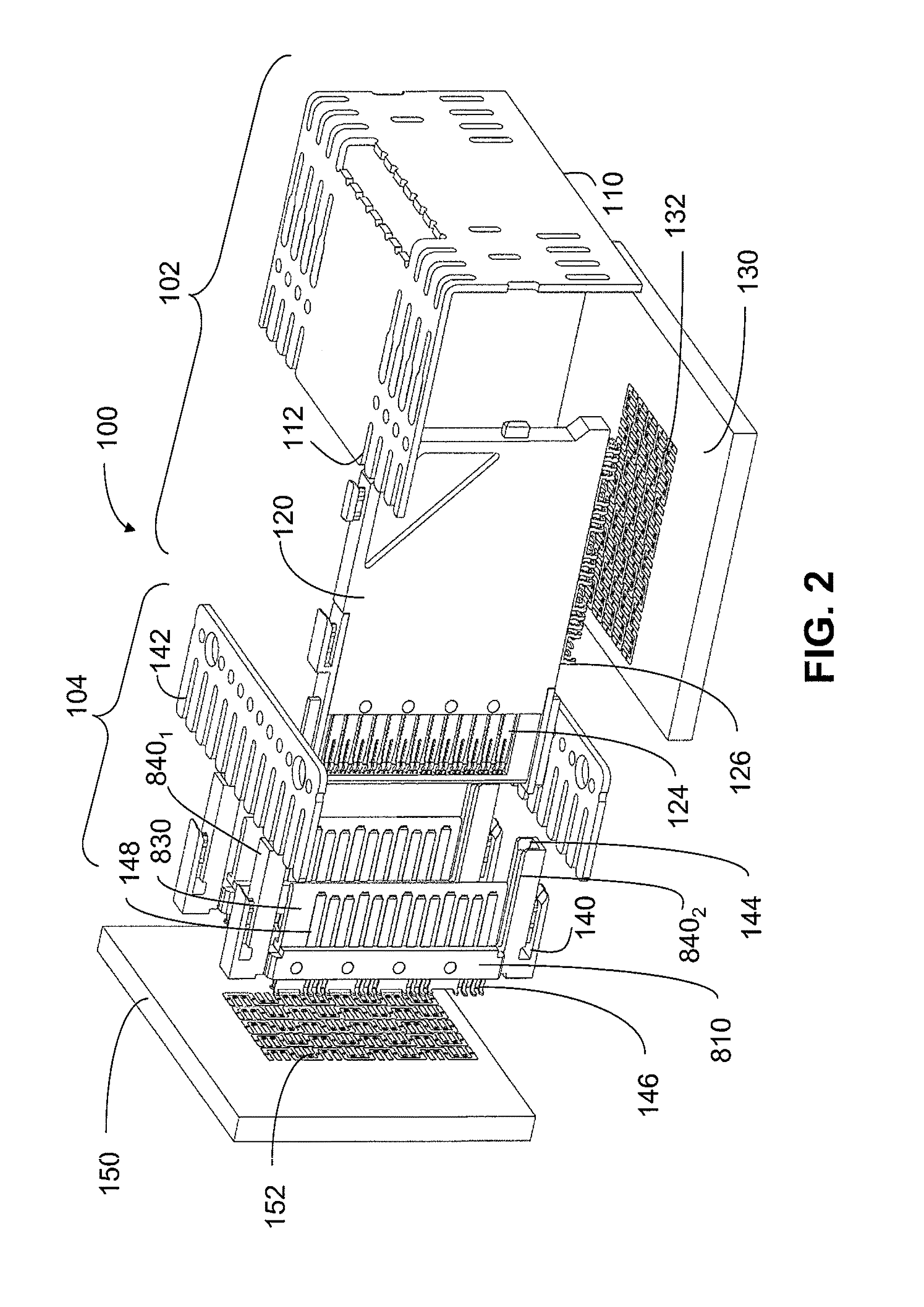
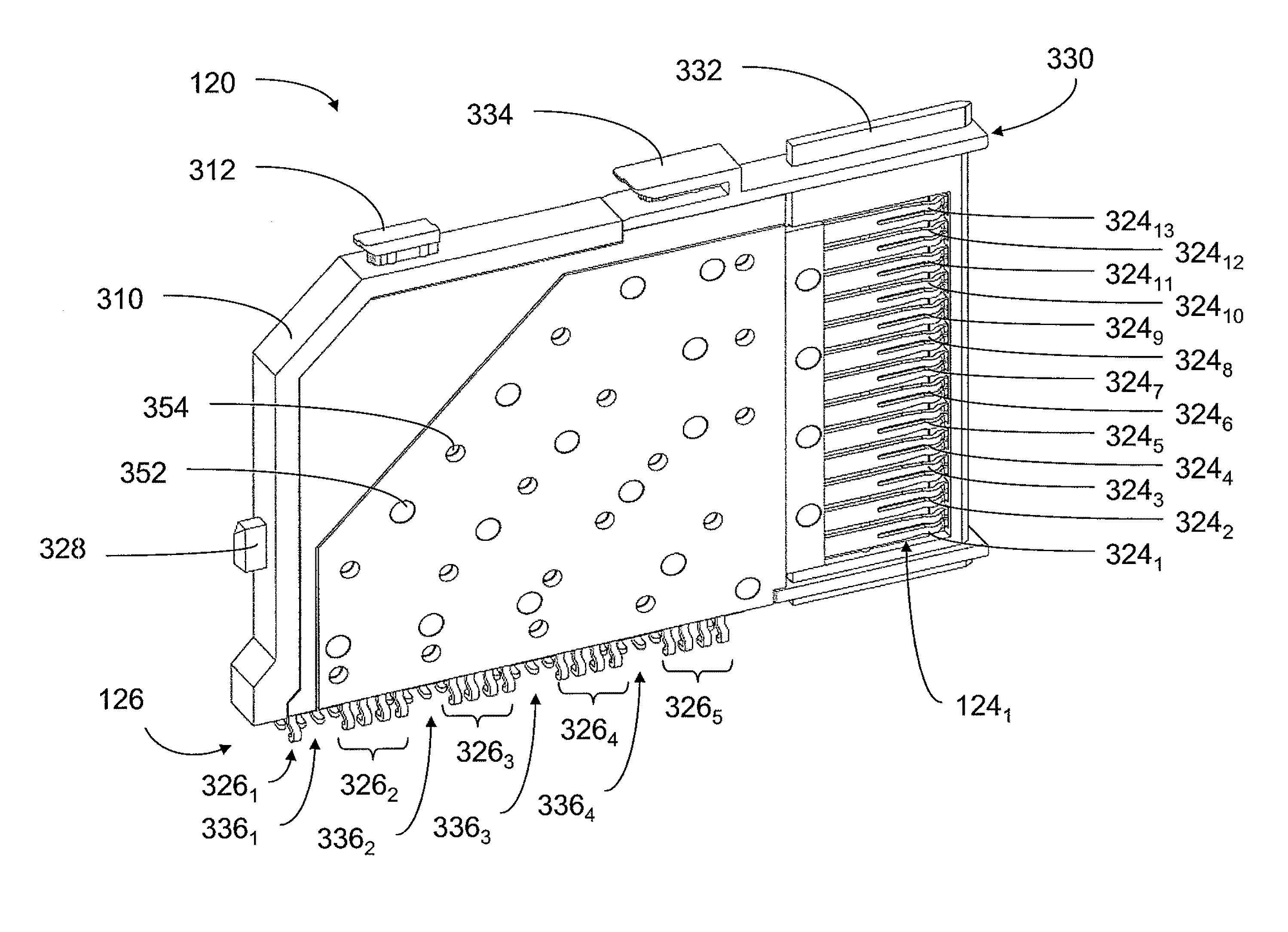
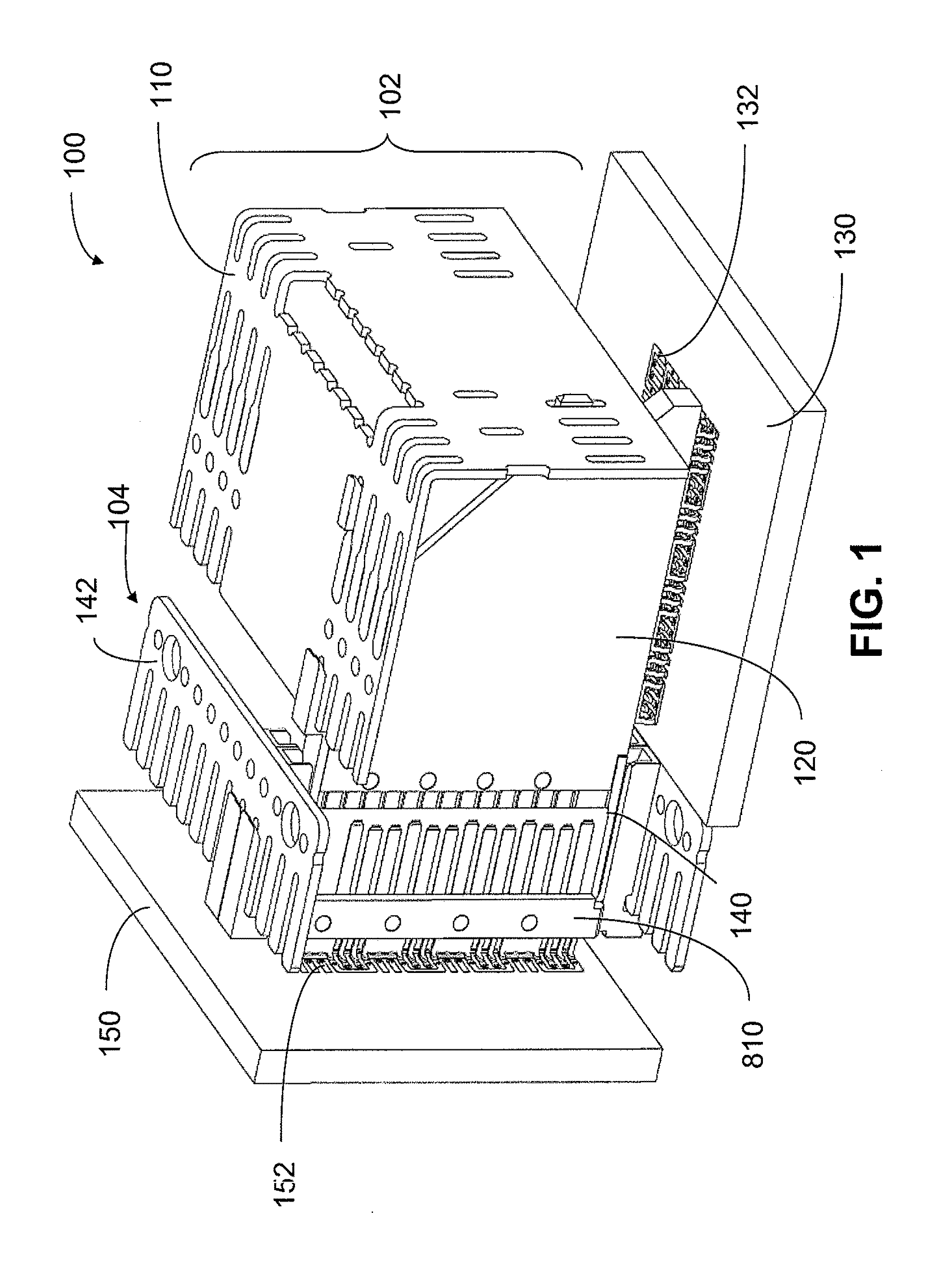
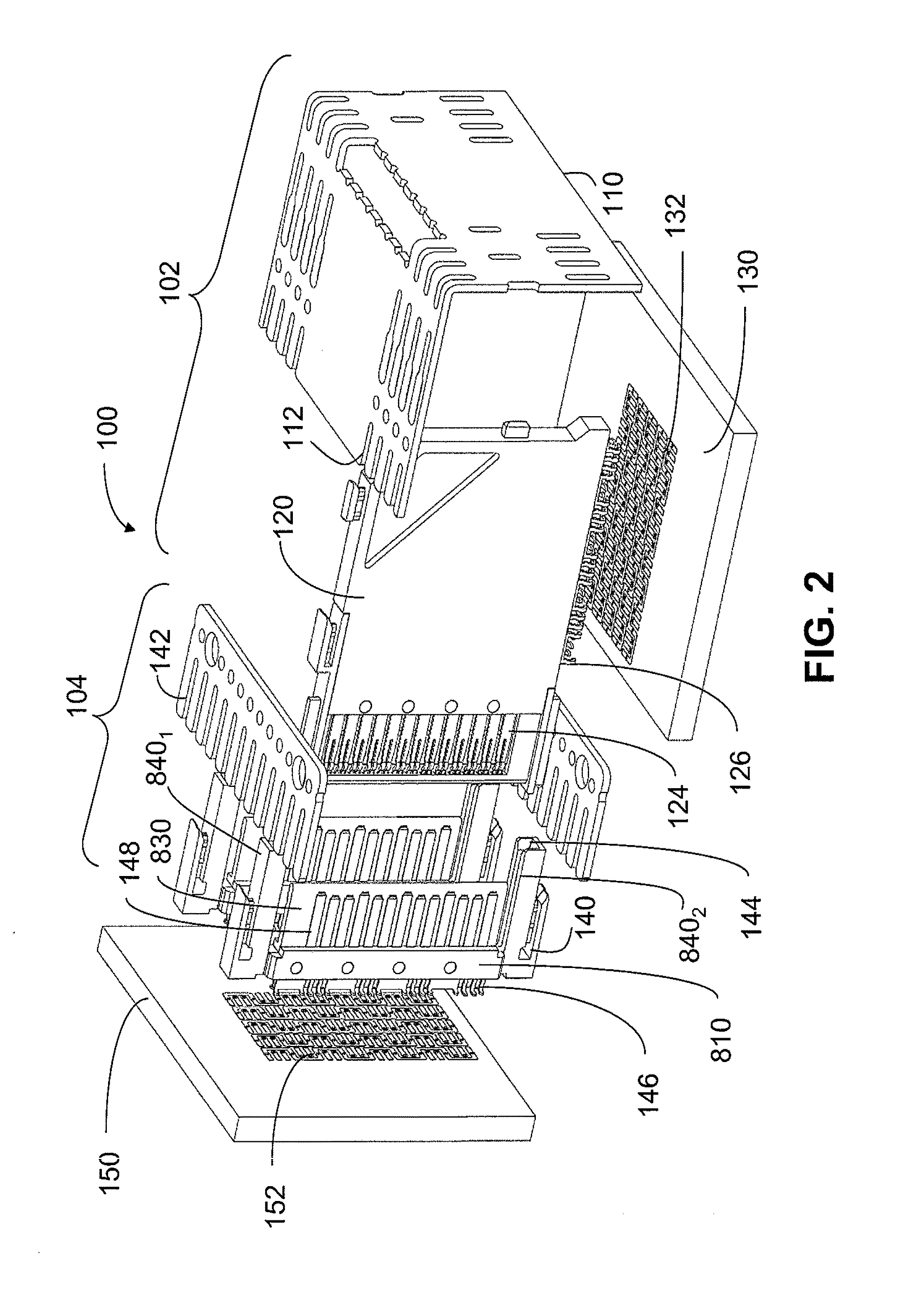
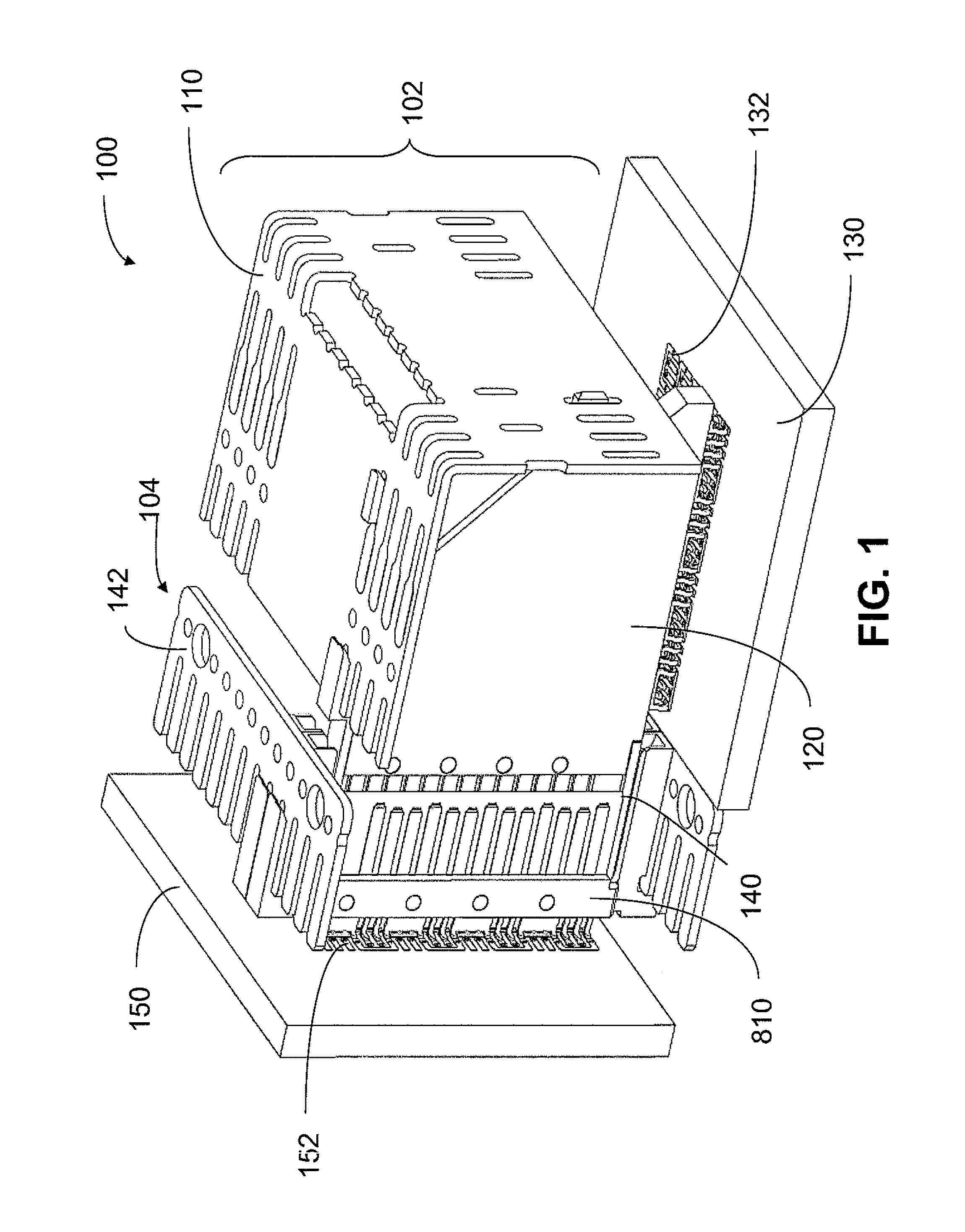
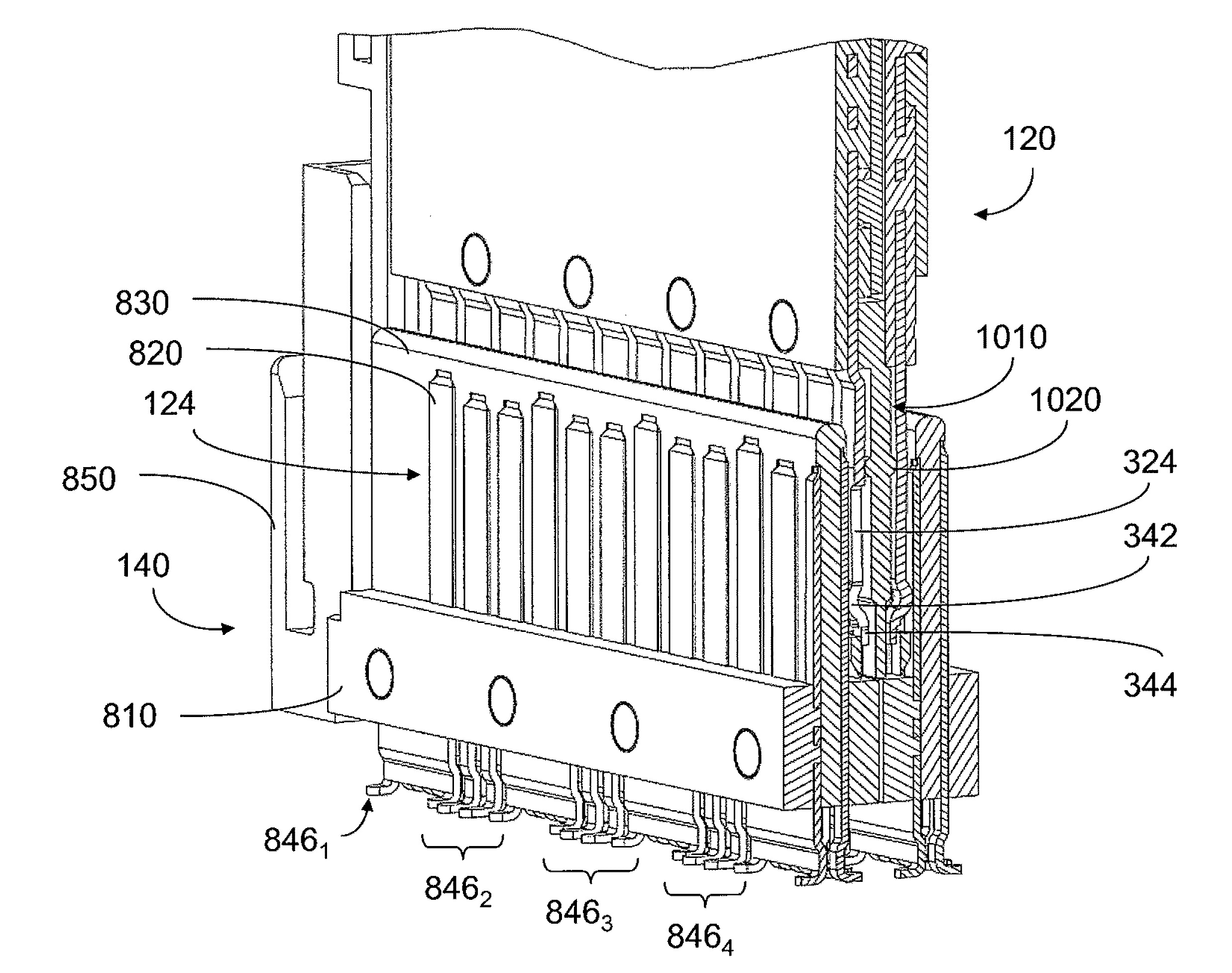
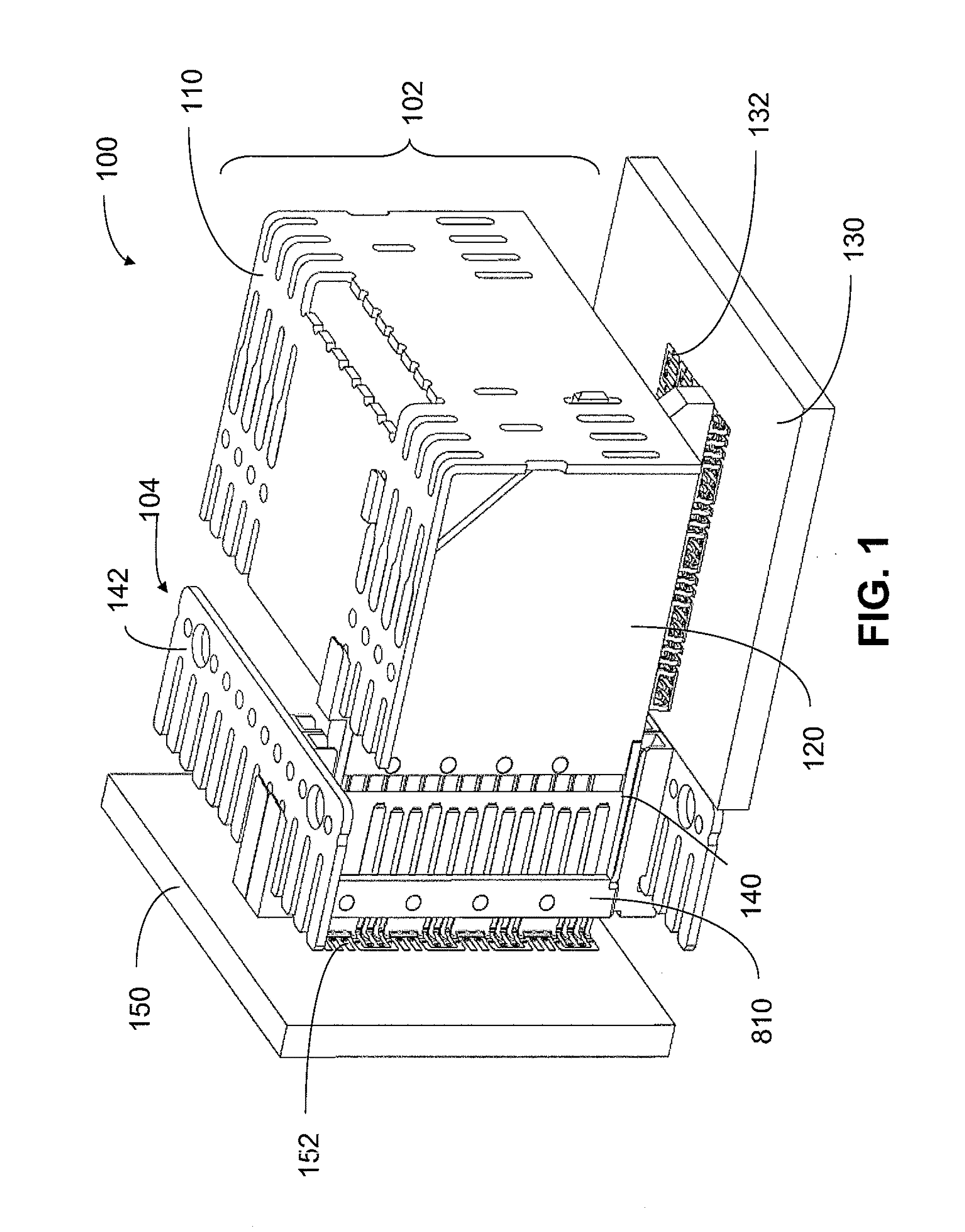
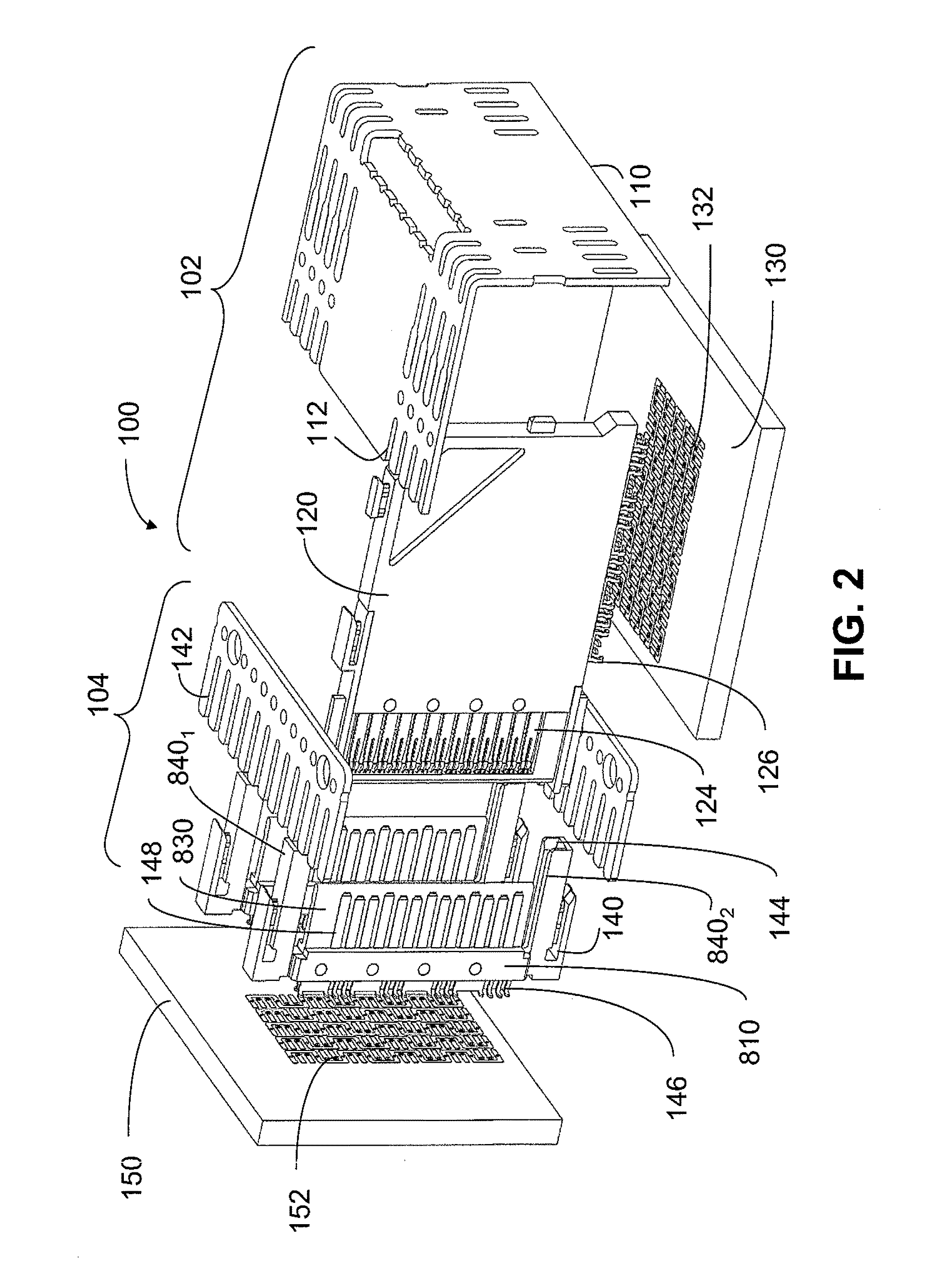
An interconnection system that includes a daughter card and backplane electrical connectors mounted to printed circuit boards at connector footprints. The spring rate of beam-shaped contacts in the daughter card connector increases while mating with the backplane connector so that the retention force may be greater than the insertion force. Such a change in spring rate may be achieved by positioning the beam-shaped contacts adjacent a surface of a connector housing. That surface may include a projection that aligns with the beam-shaped contact. When the connectors are unmated, the beam-shaped contact may be spaced from the projection. As the connectors begin to mate, a central portion of the beam-shaped contact may be pressed against the projection, which has the effect of shortening the beam length and increasing its stiffness.
Owner:AMPHENOL CORP
High density electrical connector with variable insertion and retention force
ActiveUS20110212649A1High retention forceHigh spring rateTwo-part coupling devicesCoupling protective earth/shielding arrangementsElectricityBackplane
An interconnection system that includes a daughter card and backplane electrical connectors mounted to printed circuit boards at connector footprints. The spring rate of beam-shaped contacts in the daughter card connector increases while mating with the backplane connector so that the retention force may be greater than the insertion force. Such a change in spring rate may be achieved by positioning the beam-shaped contacts adjacent a surface of a connector housing. That surface may include a projection that aligns with the beam-shaped contact. When the connectors are unmated, the beam-shaped contact may be spaced from the projection. As the connectors begin to mate, a central portion of the beam-shaped contact may be pressed against the projection, which has the effect of shortening the beam length and increasing its stiffness.
Owner:AMPHENOL CORP
High density electrical connector and PCB footprint
ActiveUS8272877B2Reduces instanceImprove signal integrityTwo-part coupling devicesElectrical connection printed elementsHigh densityElectrical conductor
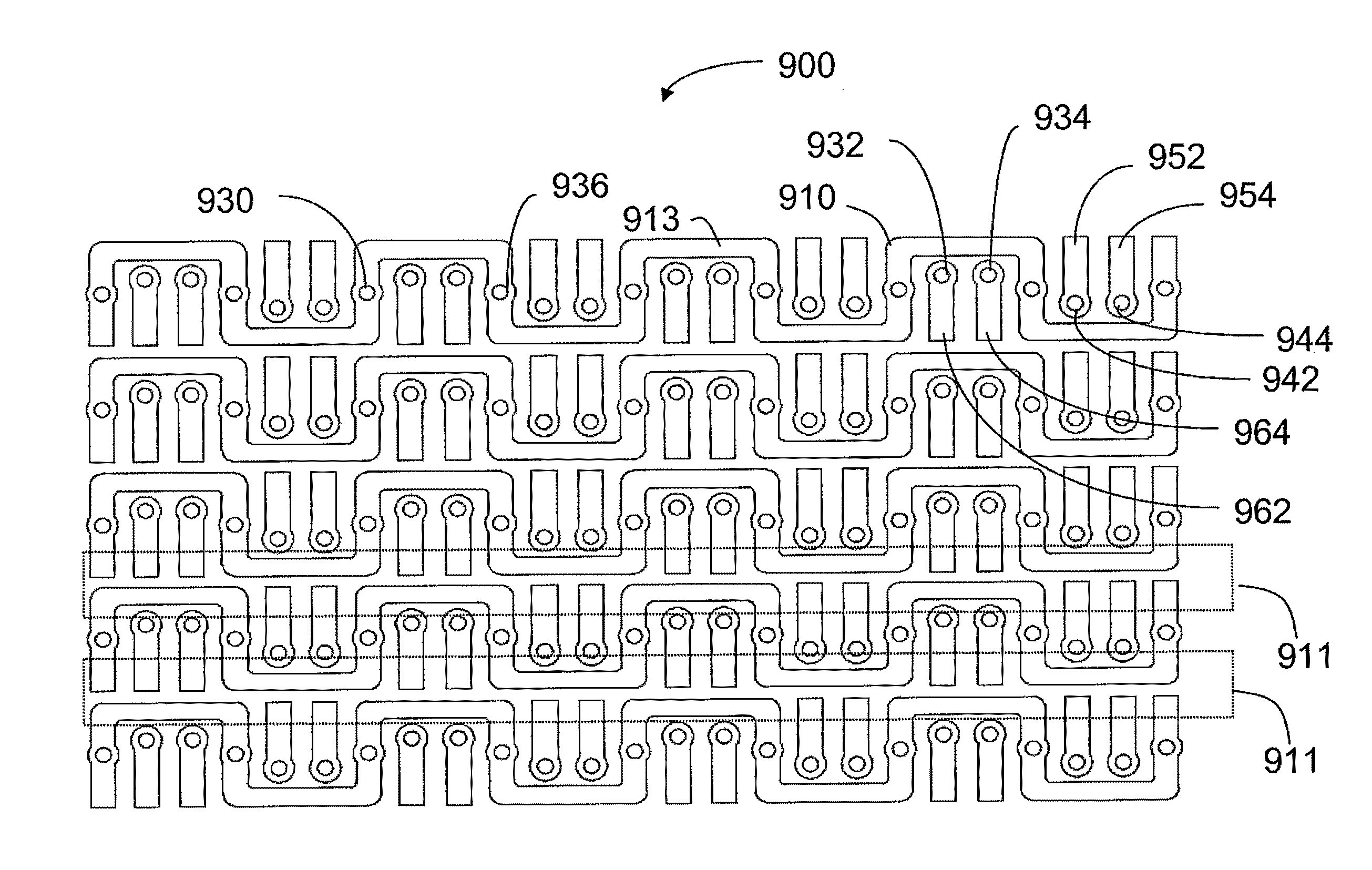
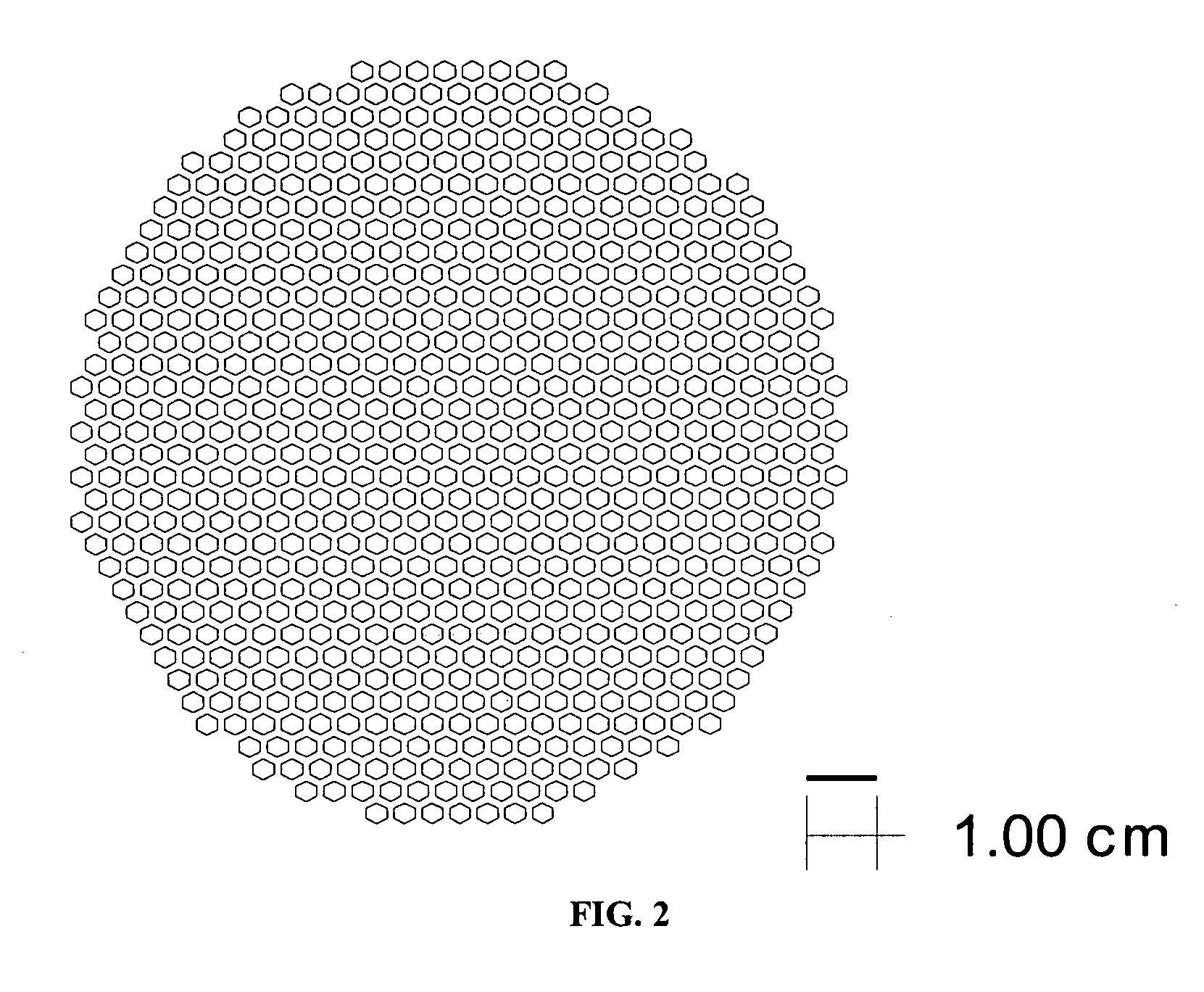
An interconnection system that includes a daughter card and backplane electrical connectors, each mounted to a printed circuit board at a connector footprint. The backplane connector has conductive elements with transition regions that allow the mating contact portions to be positioned on a uniform pitch while contact tail portions can be shaped to improve signal integrity or to provide a more compact and / or mechanically robust footprint. The conductive elements in both connectors are configured such that the contact tails of the ground conductors align from column to column, but the planar portions of the ground conductors in one column align with a pair of signal conductors in the other column, which improves mechanical and signal integrity. Mechanical integrity may be improved by forming the connector footprints with pads for the ground conductors that span multiple columns.
Owner:AMPHENOL CORP
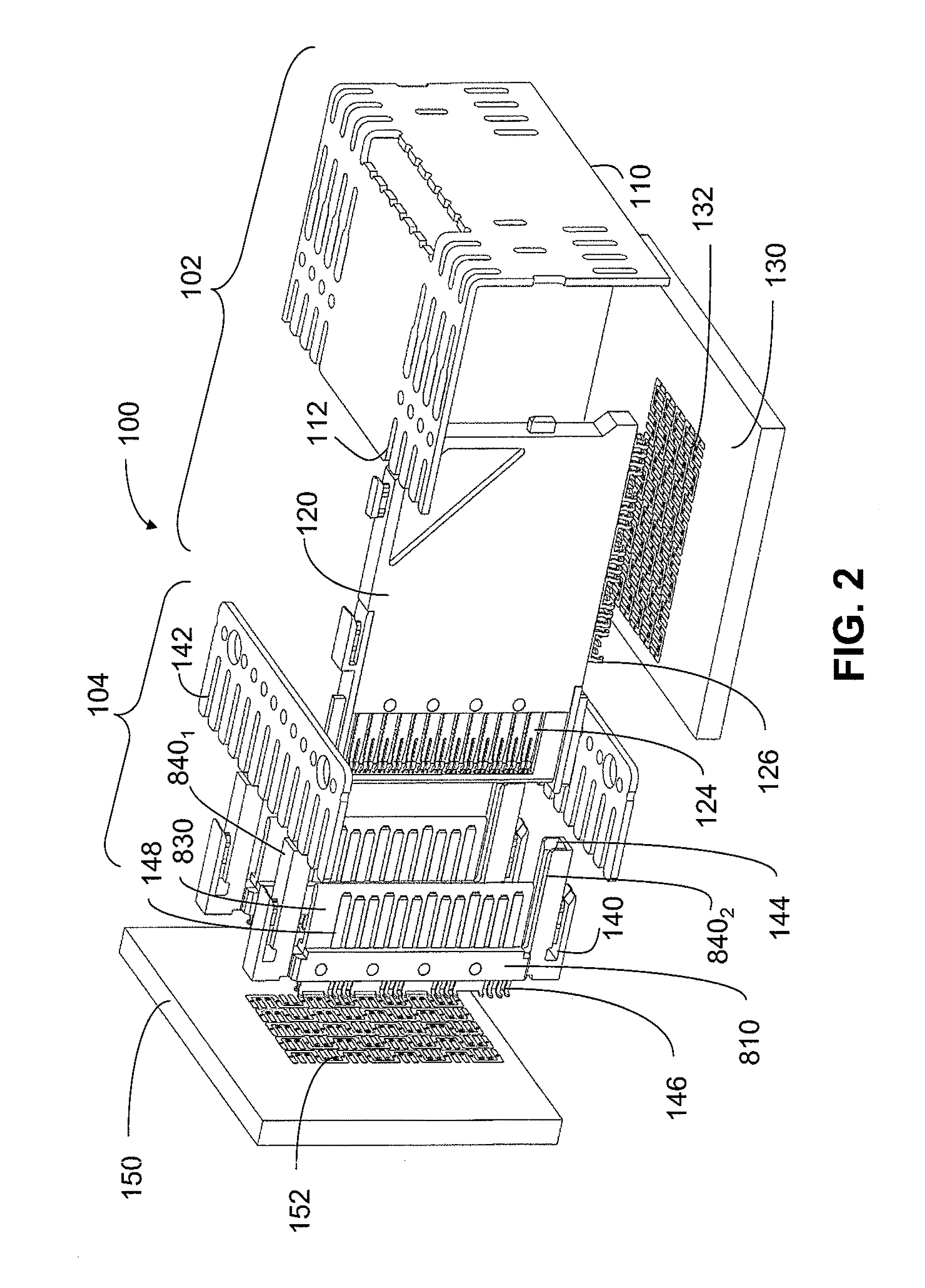
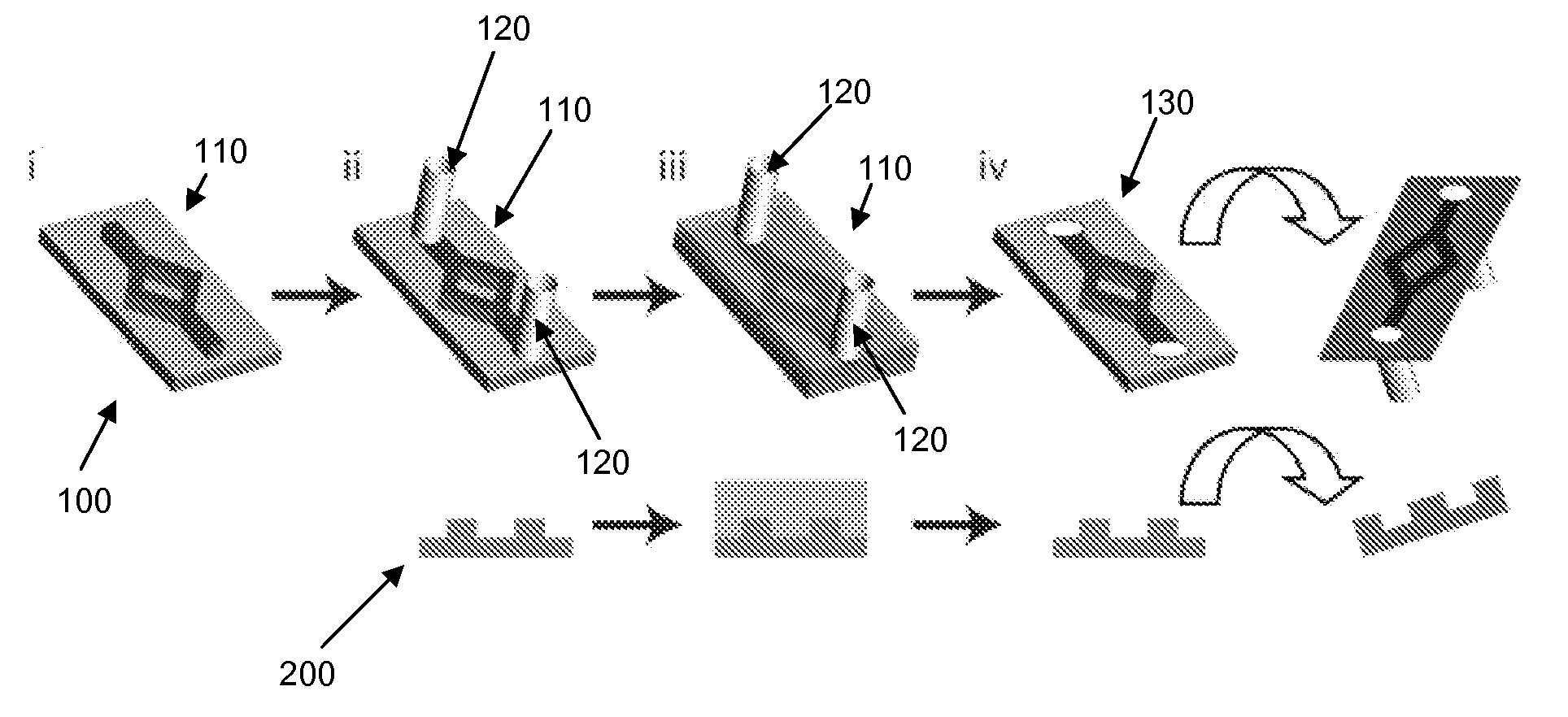
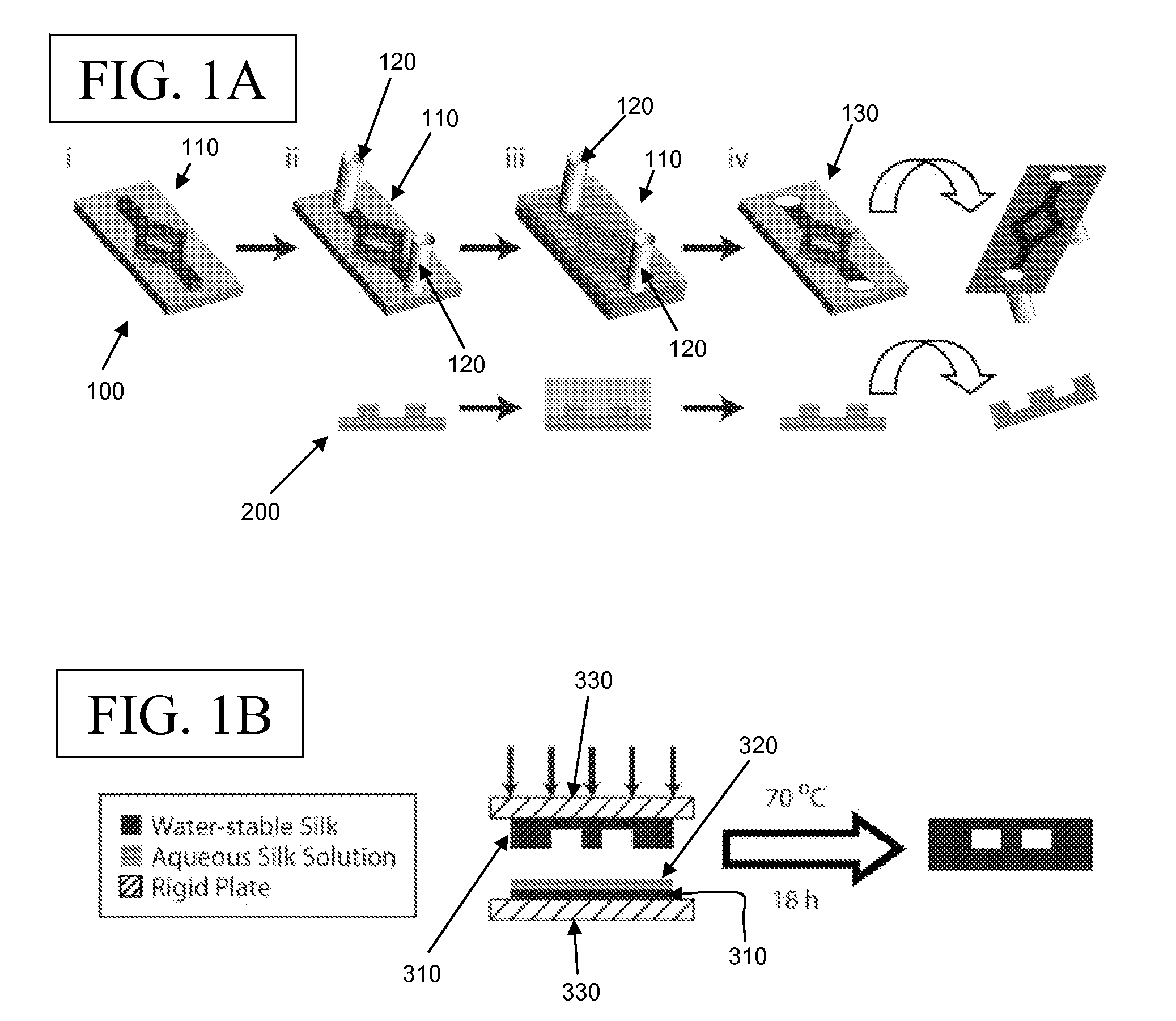
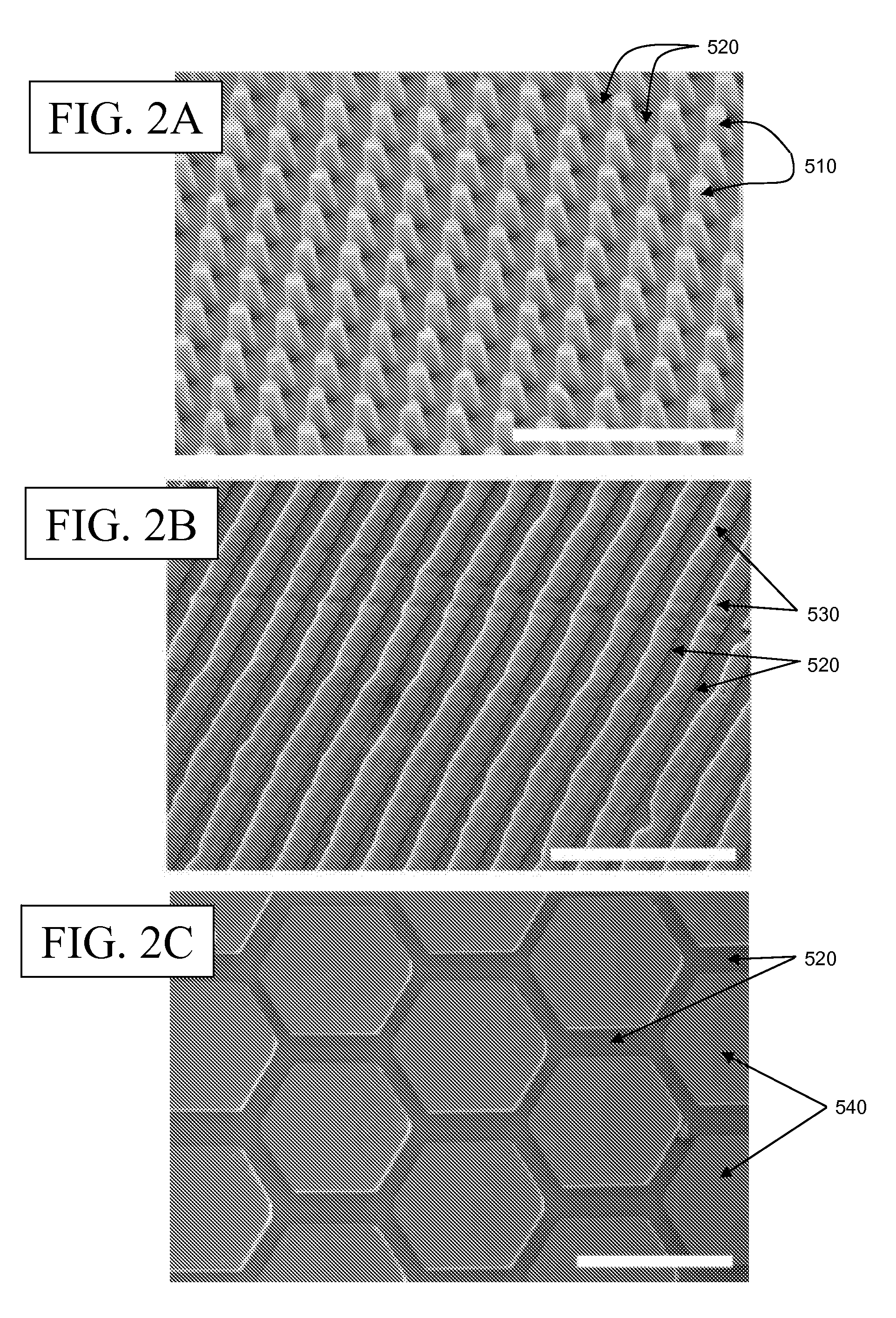
Microfluidic Devices and Methods for Fabricating the Same
ActiveUS20090004737A1Mechanically robustRobust mechanical propertyBioreactor/fermenter combinationsBiological substance pretreatmentsEngineeringElectrical and Electronics engineering
Owner:TRUSTEES OF TUFTS COLLEGE TUFTS UNIV +1
Three-dimensional thin-film semiconductor substrate with through-holes and methods of manufacturing
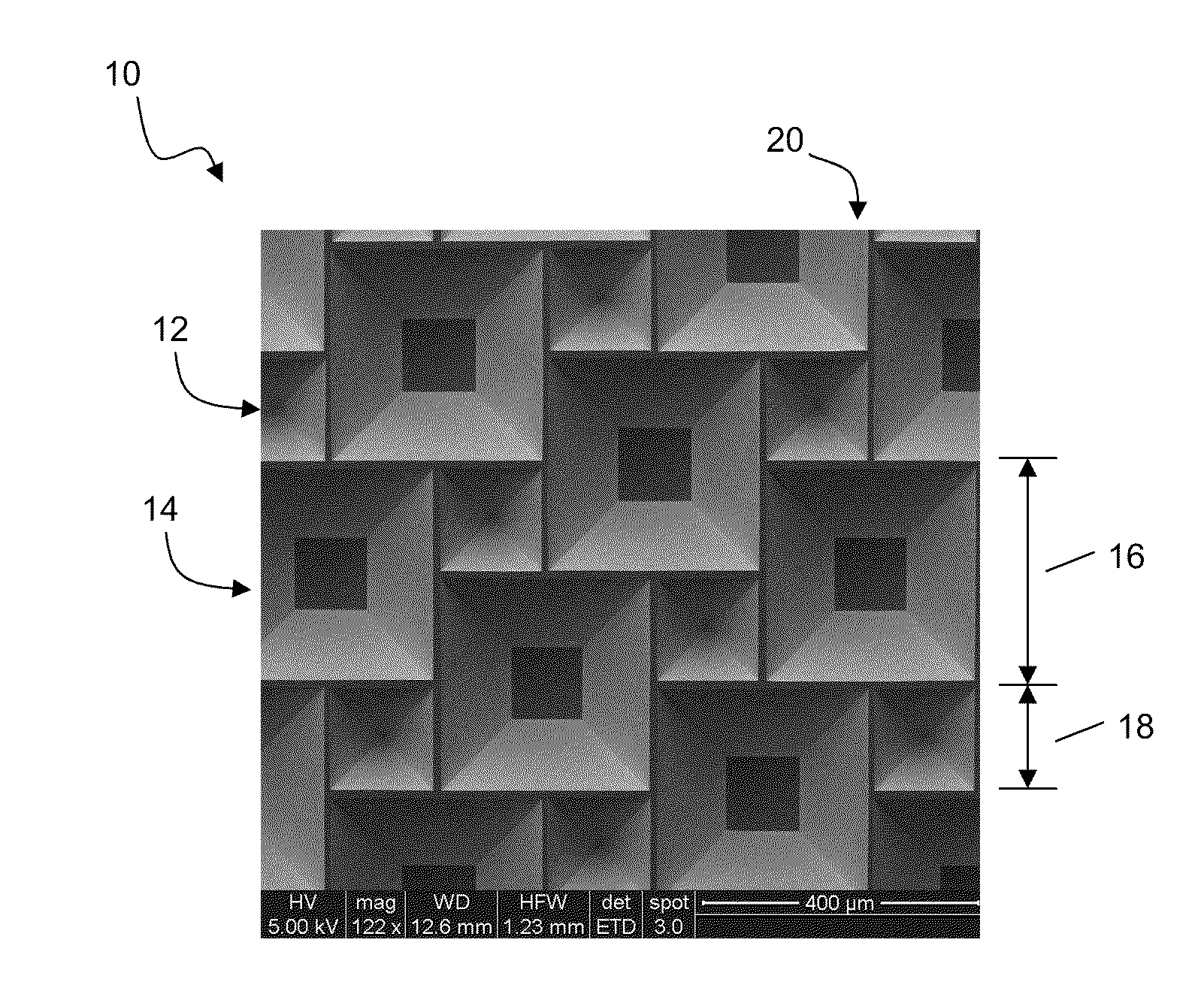
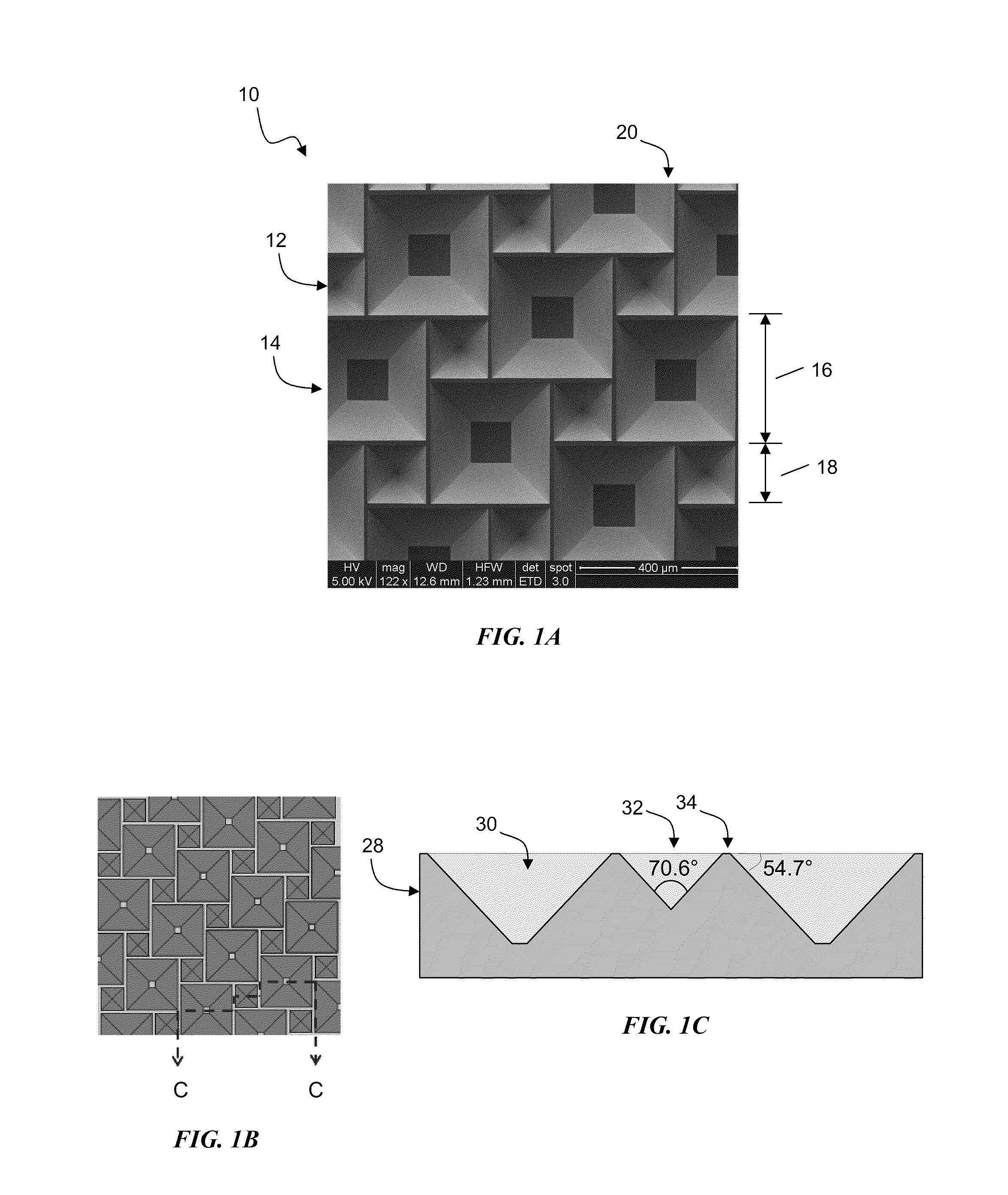
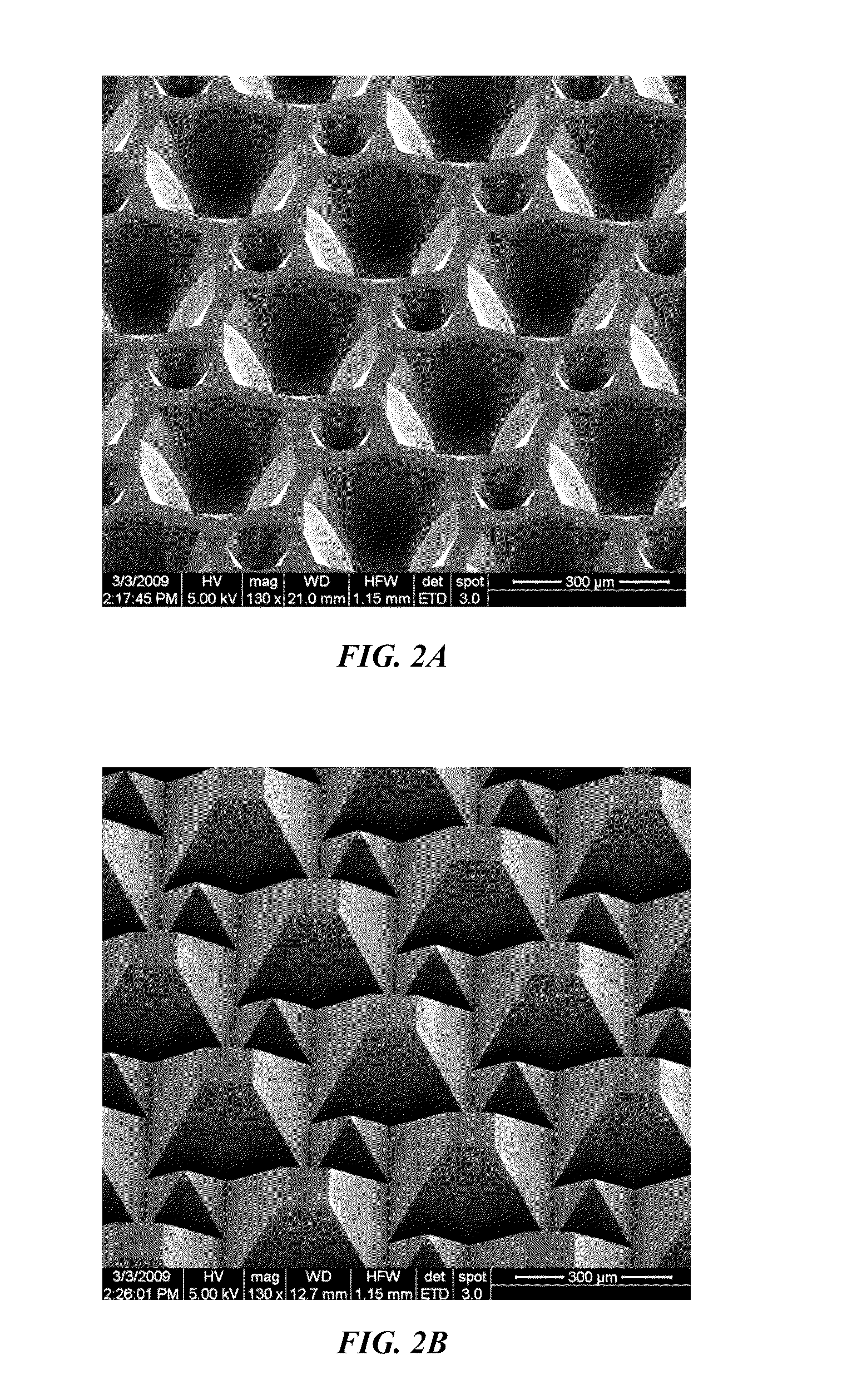
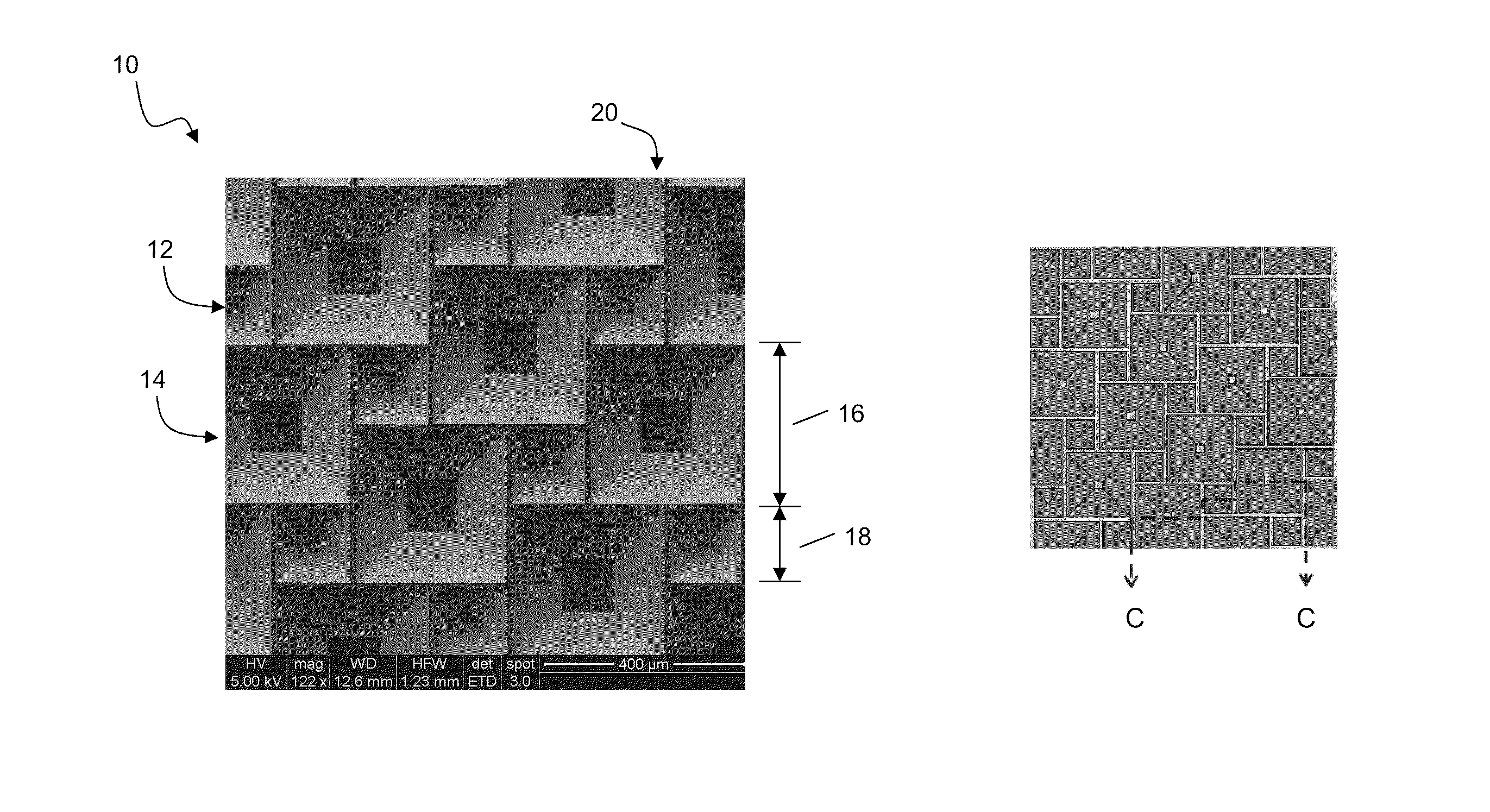
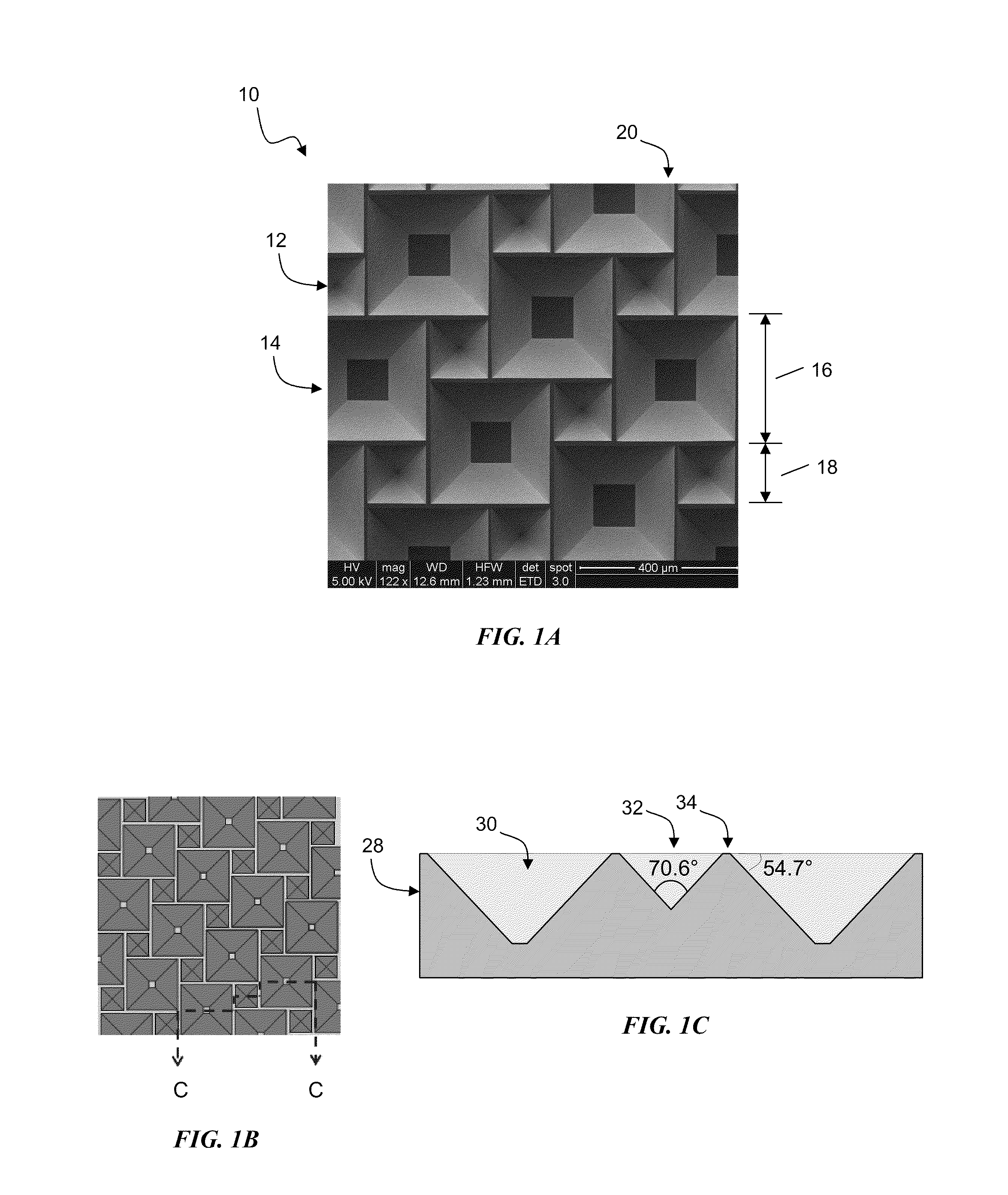
InactiveUS20100300518A1Eliminate and reduce disadvantageEliminate and reduce and problemFinal product manufactureSemiconductor/solid-state device manufacturingOptoelectronicsPorous semiconductors
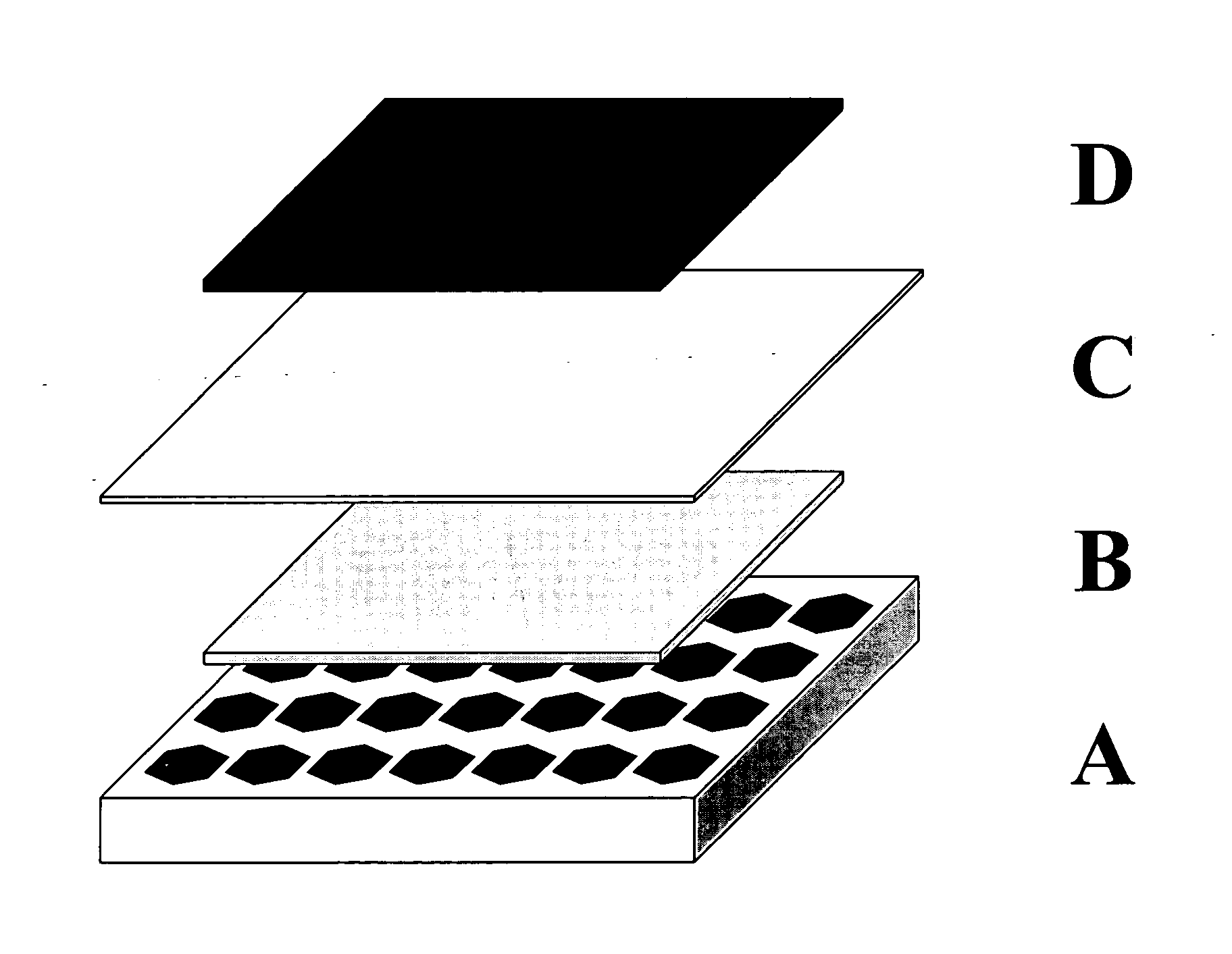
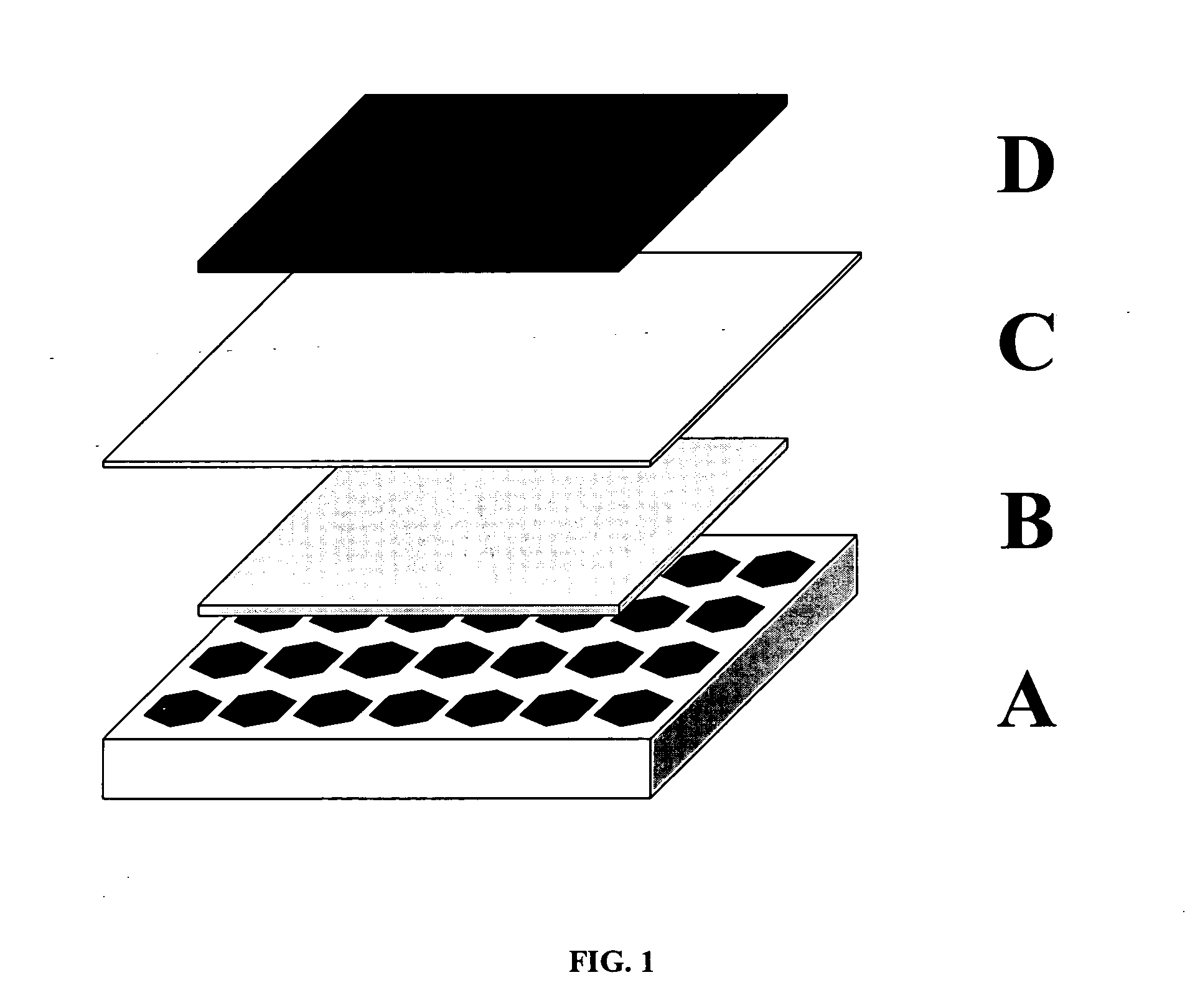
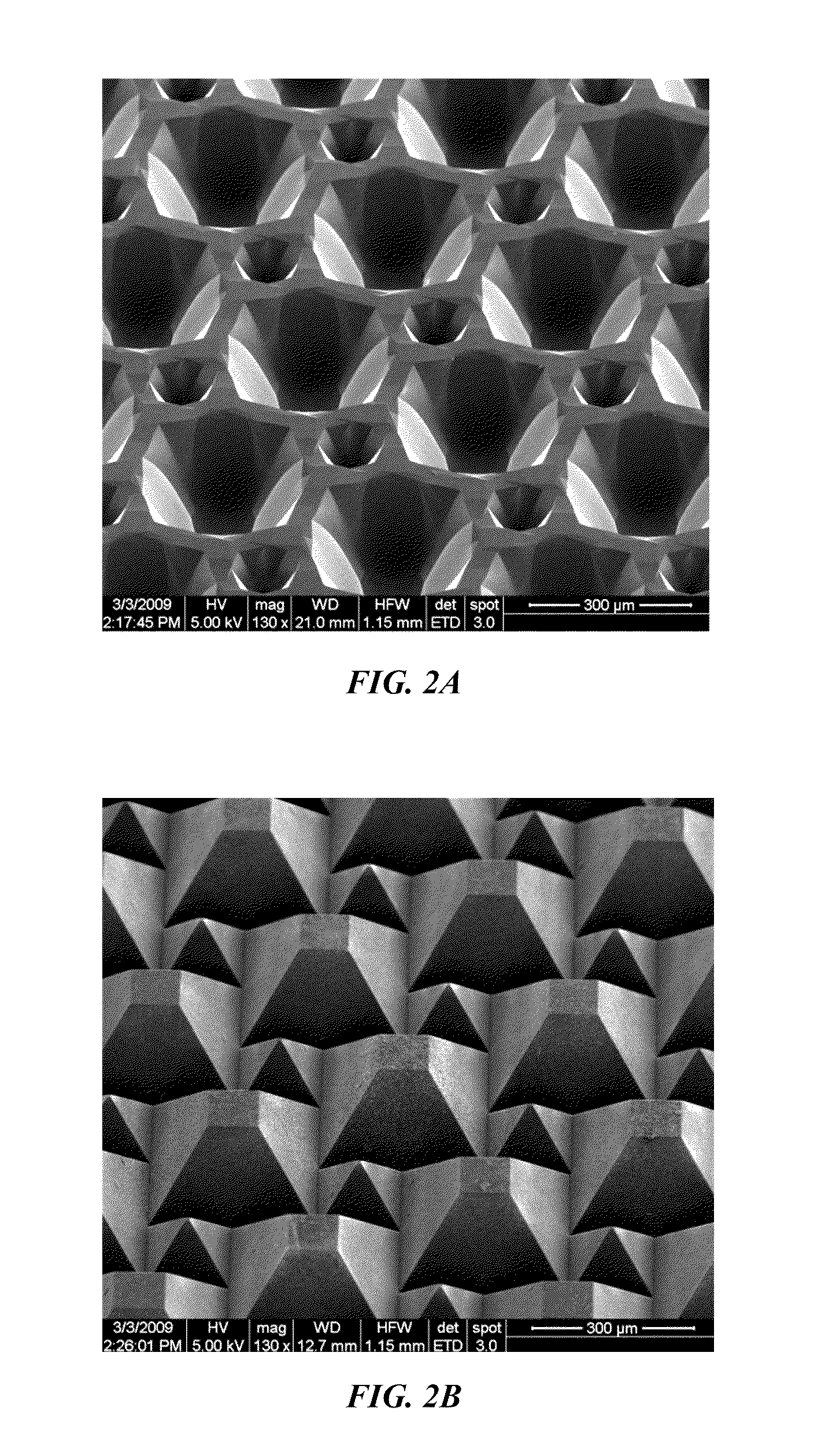
A method for the fabrication of a three-dimensional thin-film semiconductor substrate with selective through-holes is provided. A porous semiconductor layer is conformally formed on a semiconductor template comprising a plurality of three-dimensional inverted pyramidal surface features defined by top surface areas aligned along a (100) crystallographic orientation plane of the semiconductor template and a plurality of inverted pyramidal cavities defined by sidewalls aligned along the (111) crystallographic orientation plane of the semiconductor template. An epitaxial semiconductor layer is conformally formed on the porous semiconductor layer. The epitaxial semiconductor layer is released from the semiconductor template. Through-holes are selectively formed in the epitaxial semiconductor layer with openings between the front and back lateral surface planes of the epitaxial semiconductor layer to form a partially transparent three-dimensional thin-film semiconductor substrate.
Owner:BEAMREACH SOLAR INC
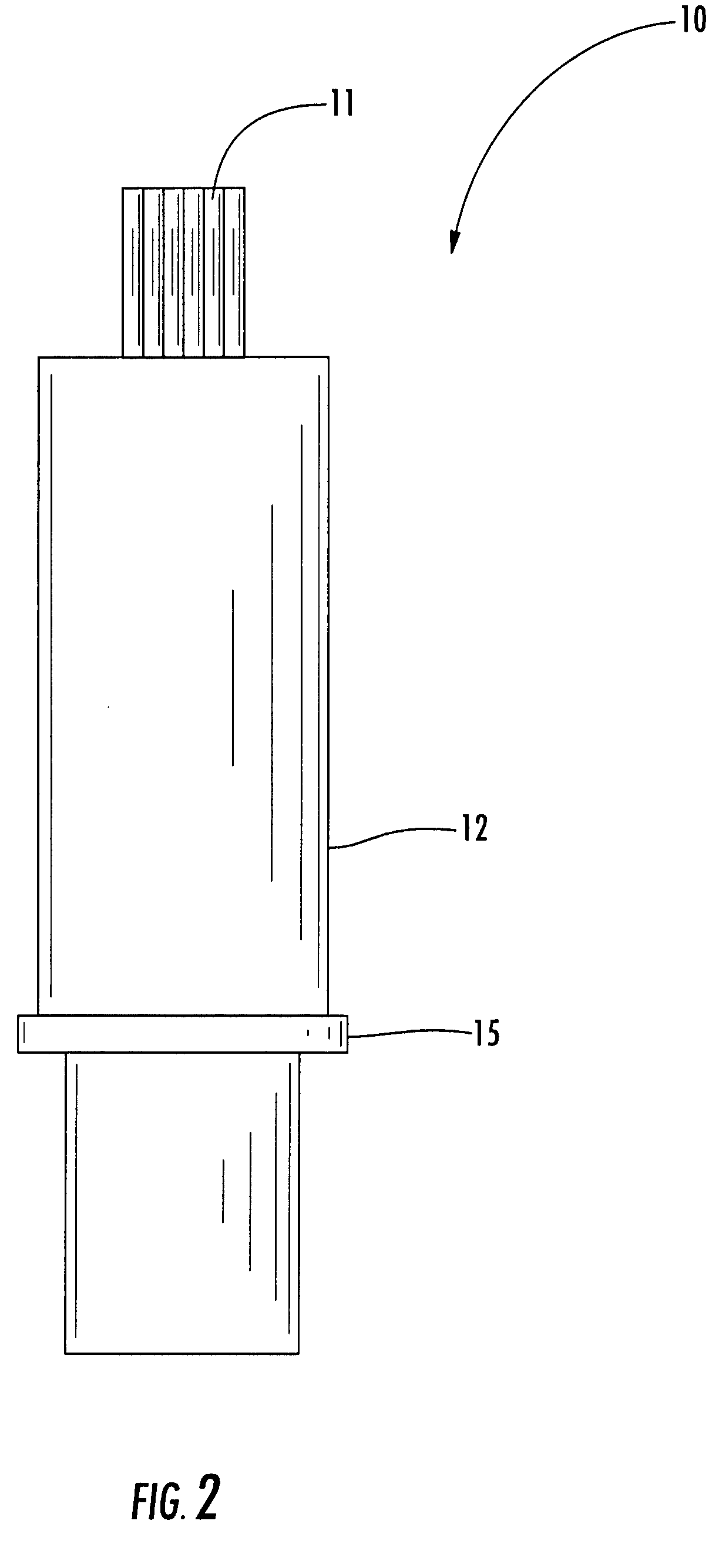
Housing Component for a Drug Delivery Device
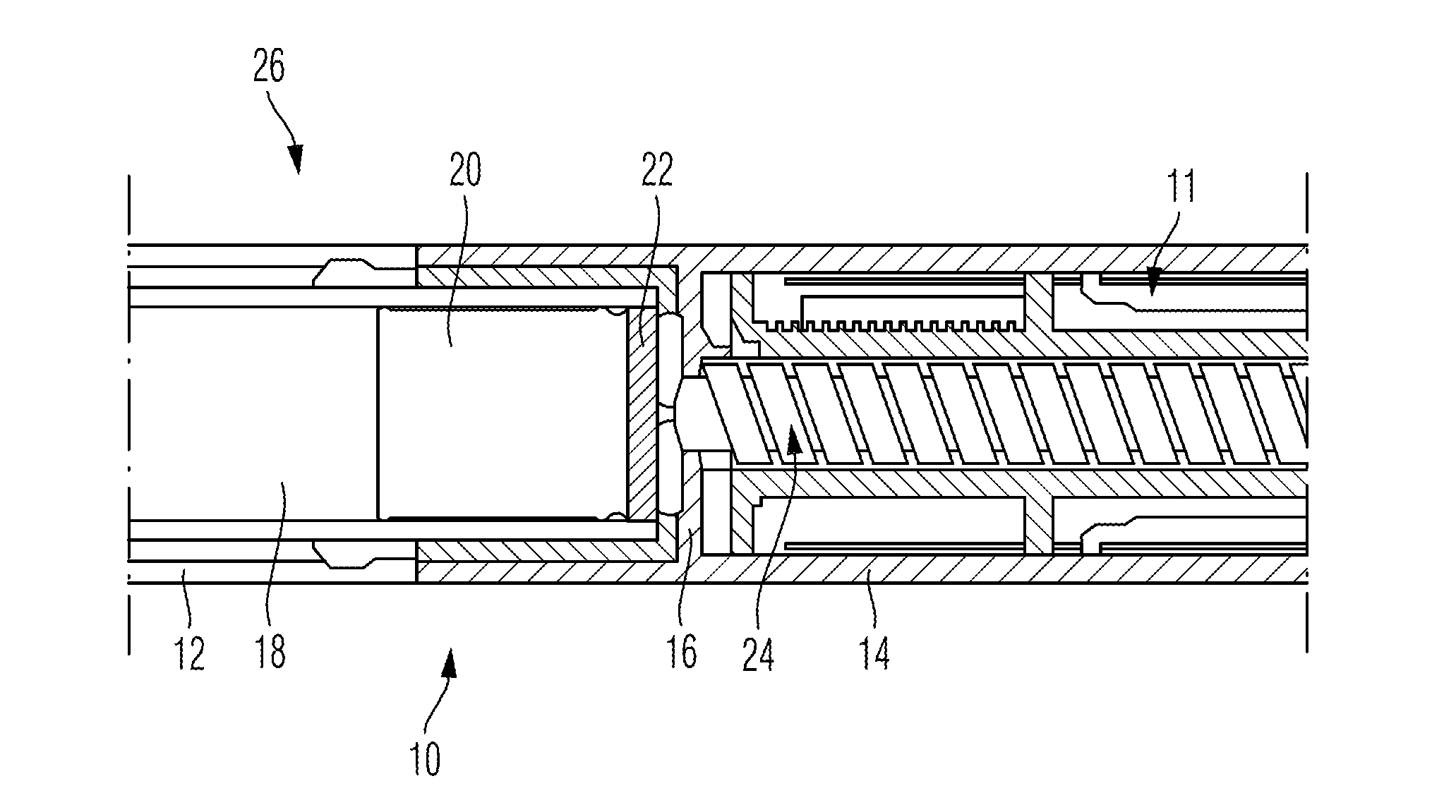
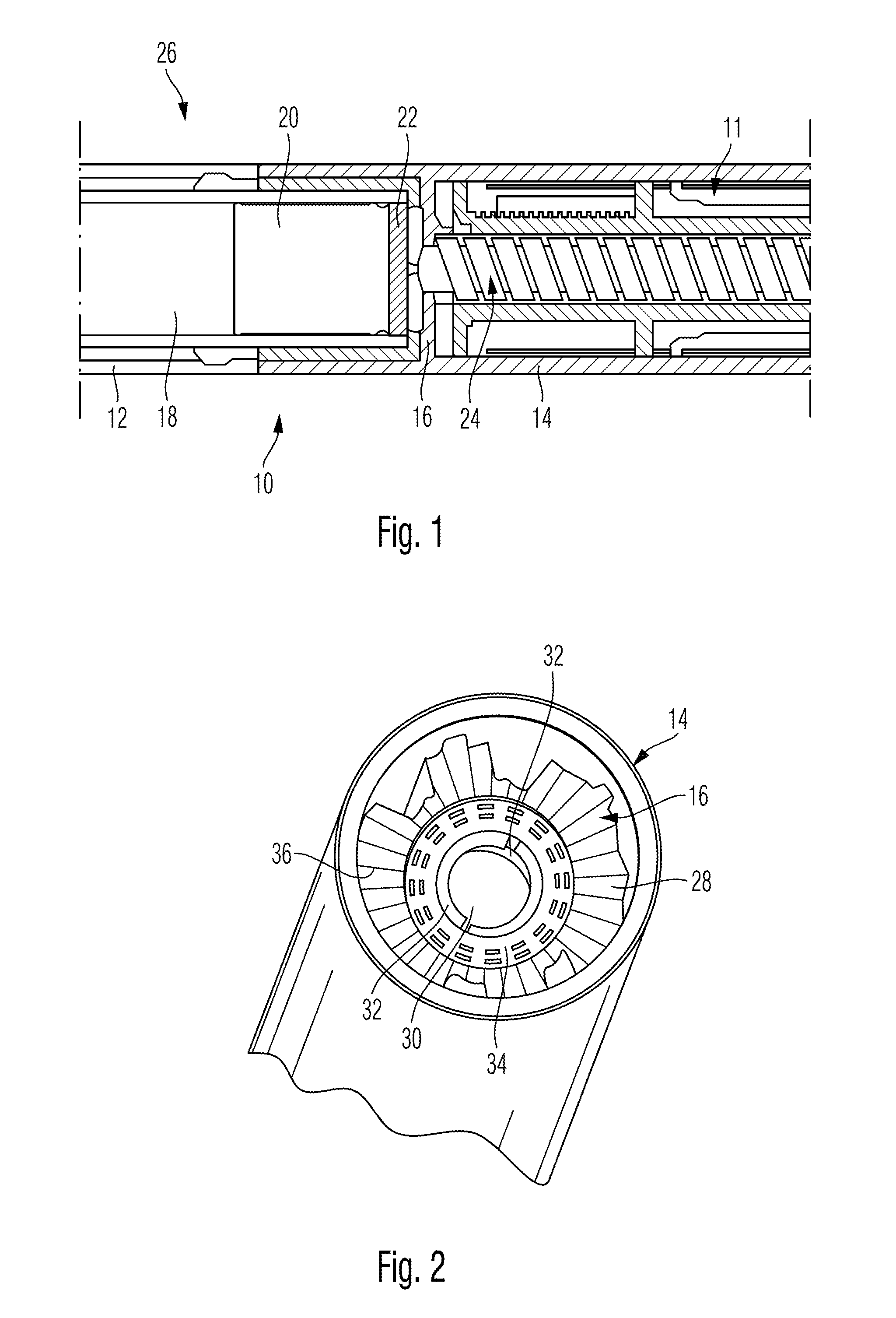
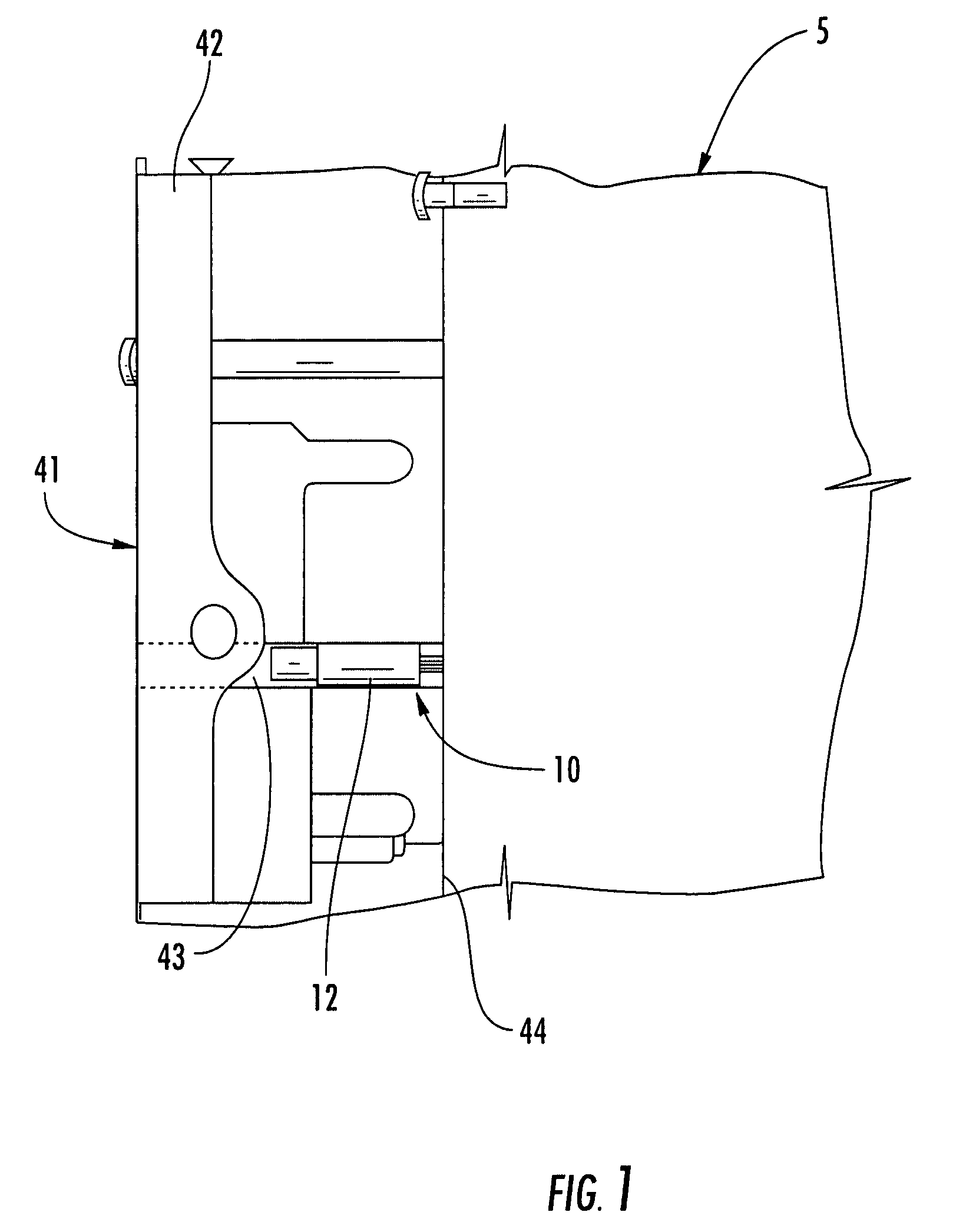
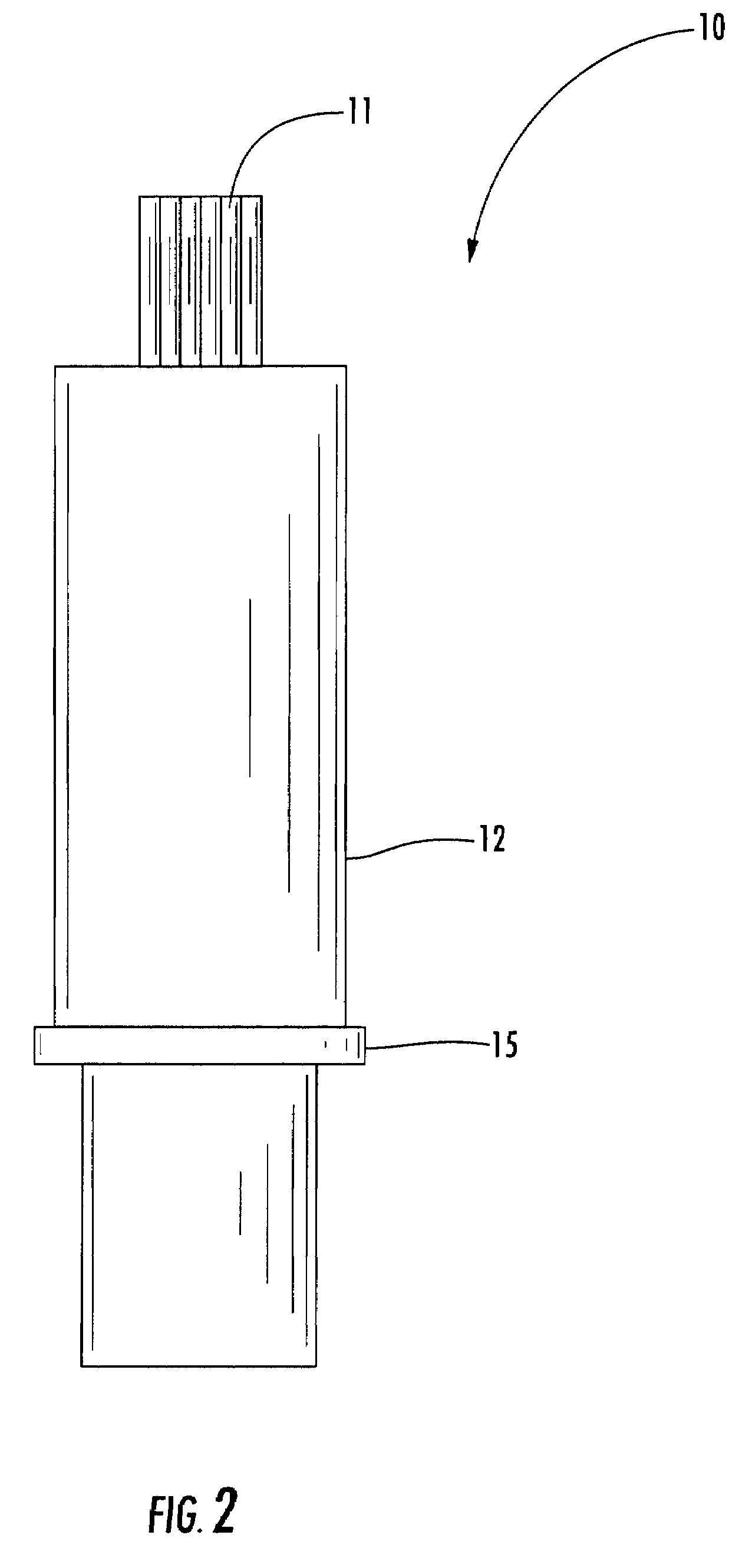
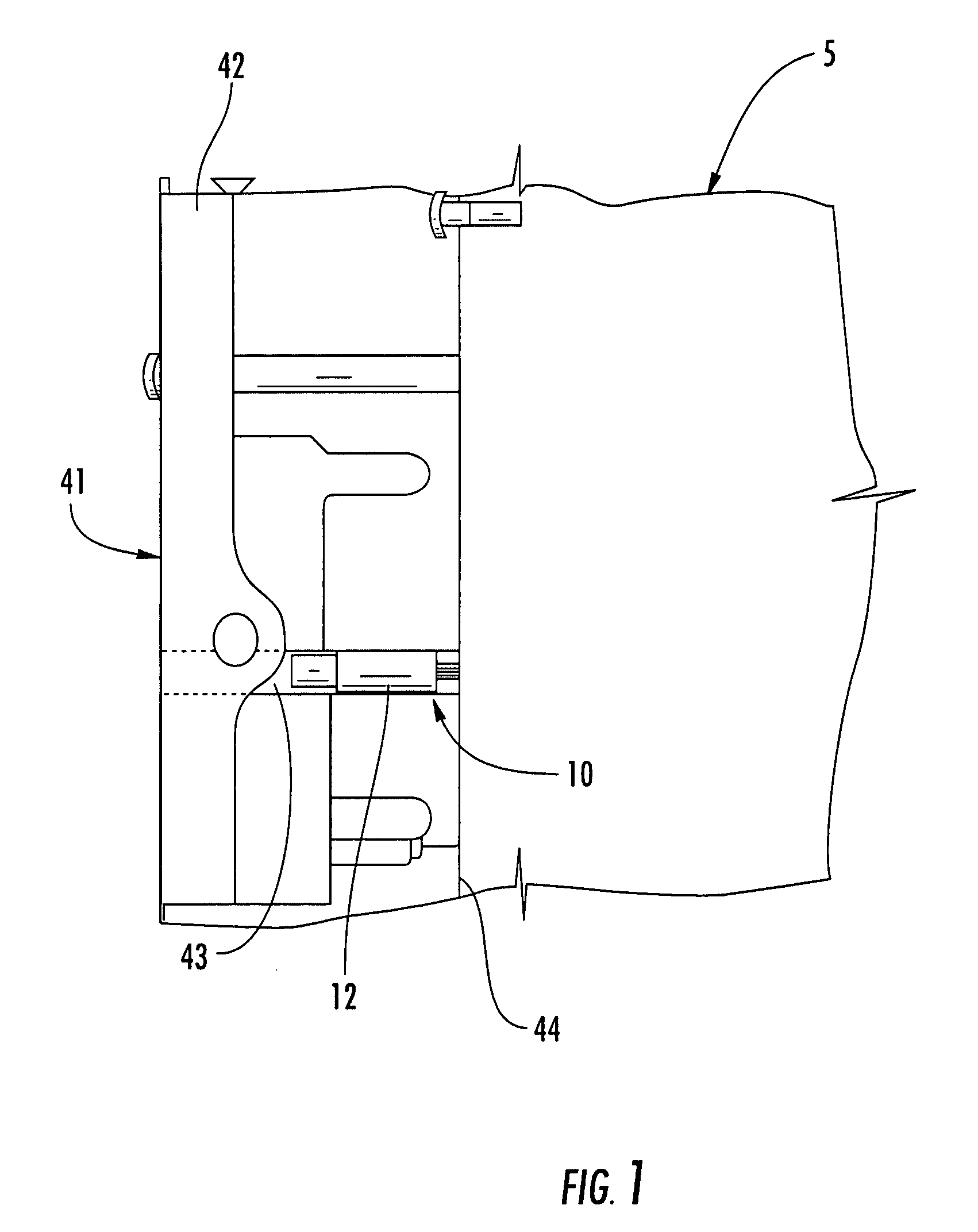
ActiveUS20120265153A1Less-prone to mechanical and elastic deformationReduce generation and built-upAmpoule syringesIntravenous devicesPistonDrug delivery
The present invention relates to a housing component of a drug delivery device (10) to accommodate a drive mechanism (11), wherein the drive mechanism (11) is operable to interact with a piston (20) of a cartridge(16) containing a medicinal product to be dispensed by the drug delivery device. The housing component comprises at least one radially inwardly extending flange portion (16) to be operably engaged with a piston rod (24) of the drive mechanism (11), characterized in that the flange portion (16) is at least partially structurally strengthened.
Owner:SANOFI AVENTIS DEUTSCHLAND GMBH
Method and system for accurate alignment and registration of array for DNA sequencing
ActiveUS20110268347A1High precisionImprove trustImage enhancementImage analysisGenomic sequencingPattern matching
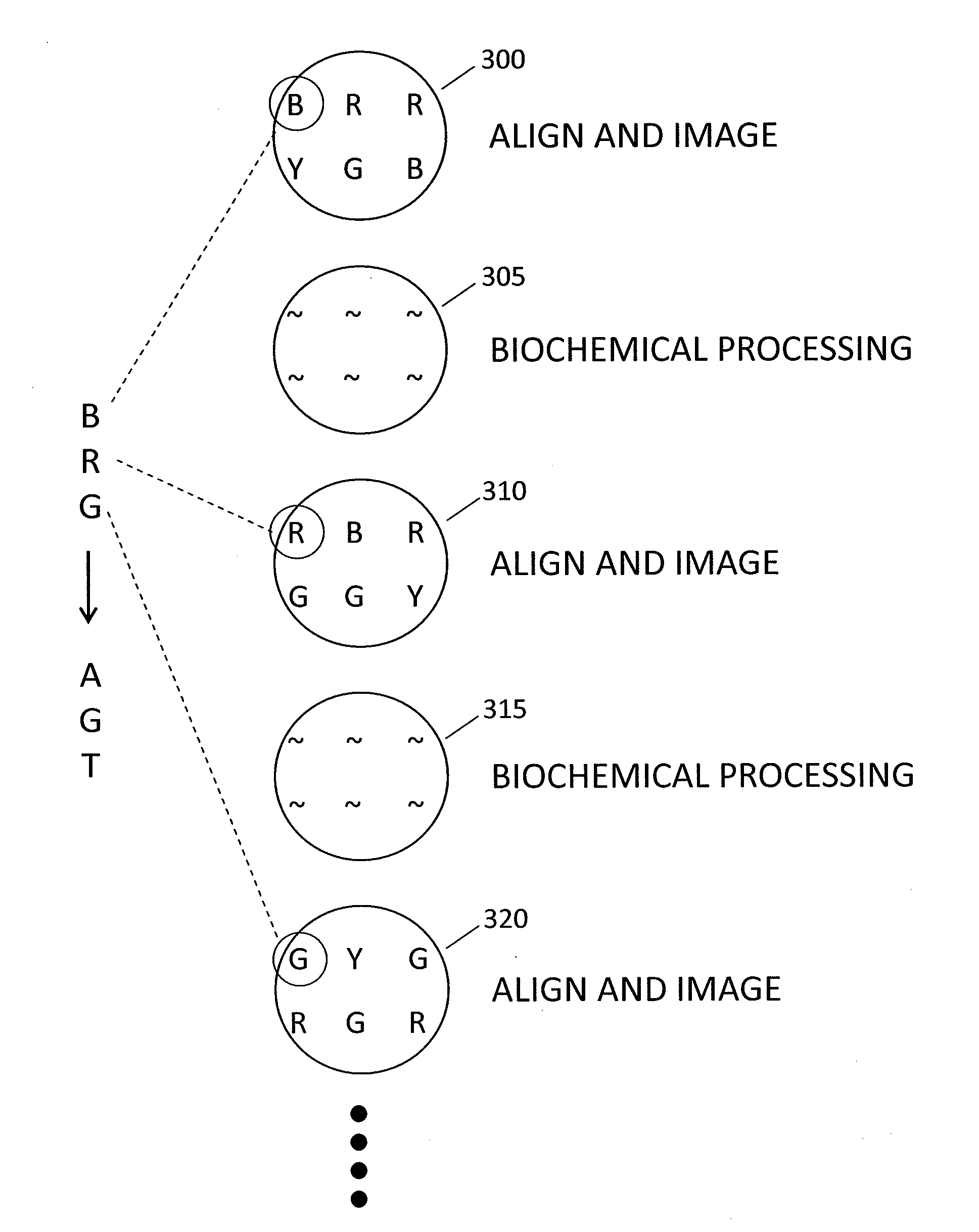
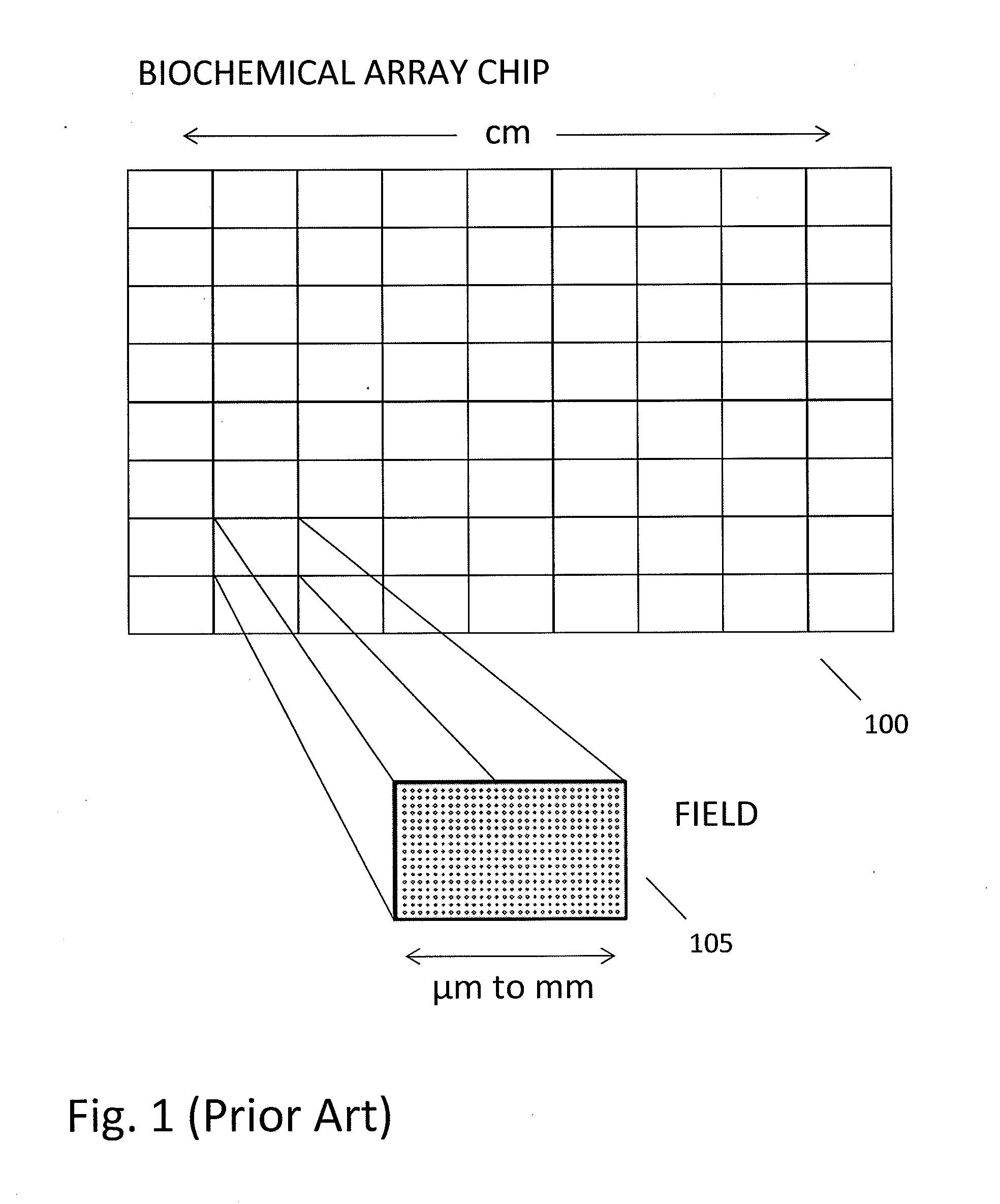
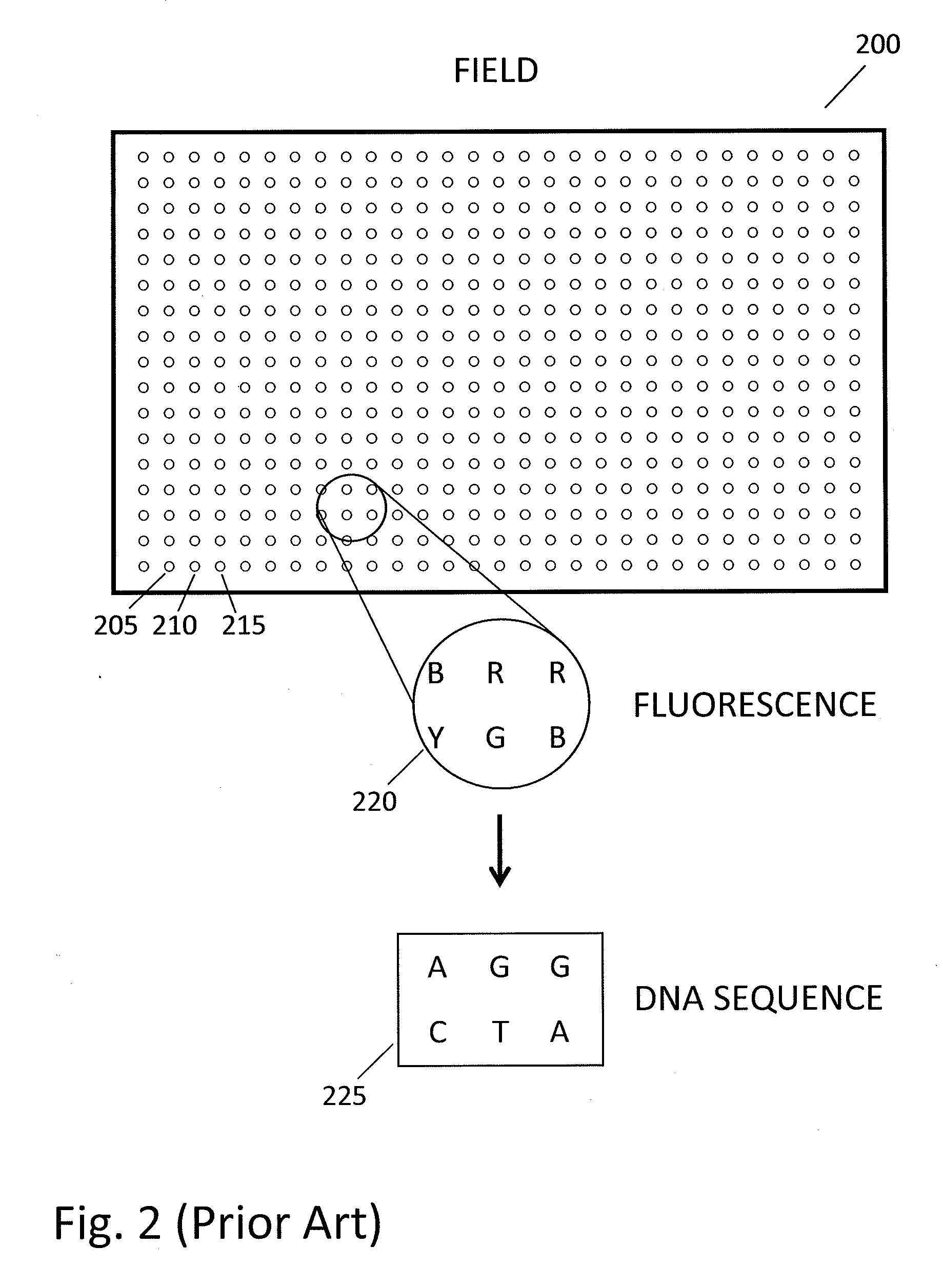
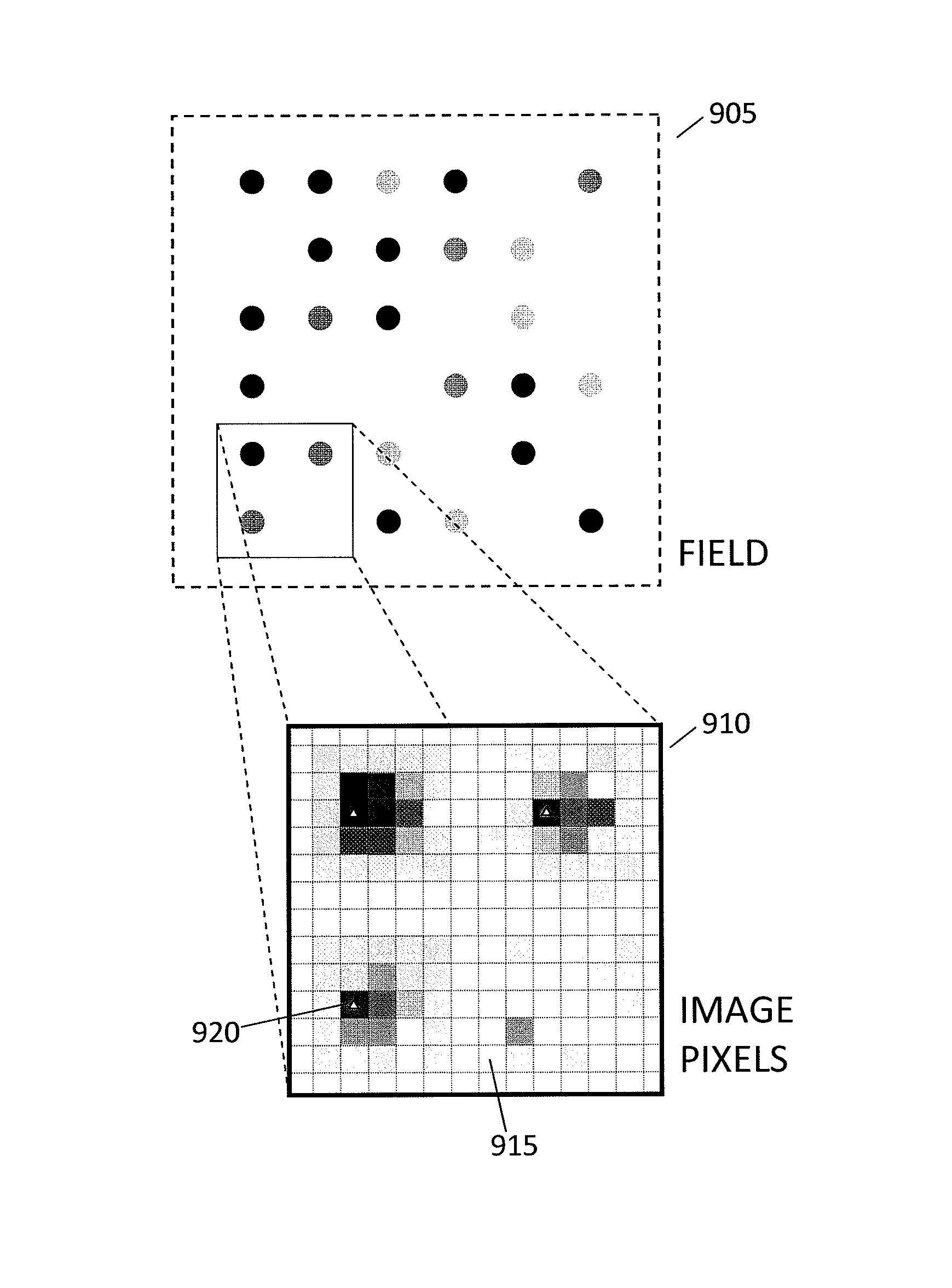
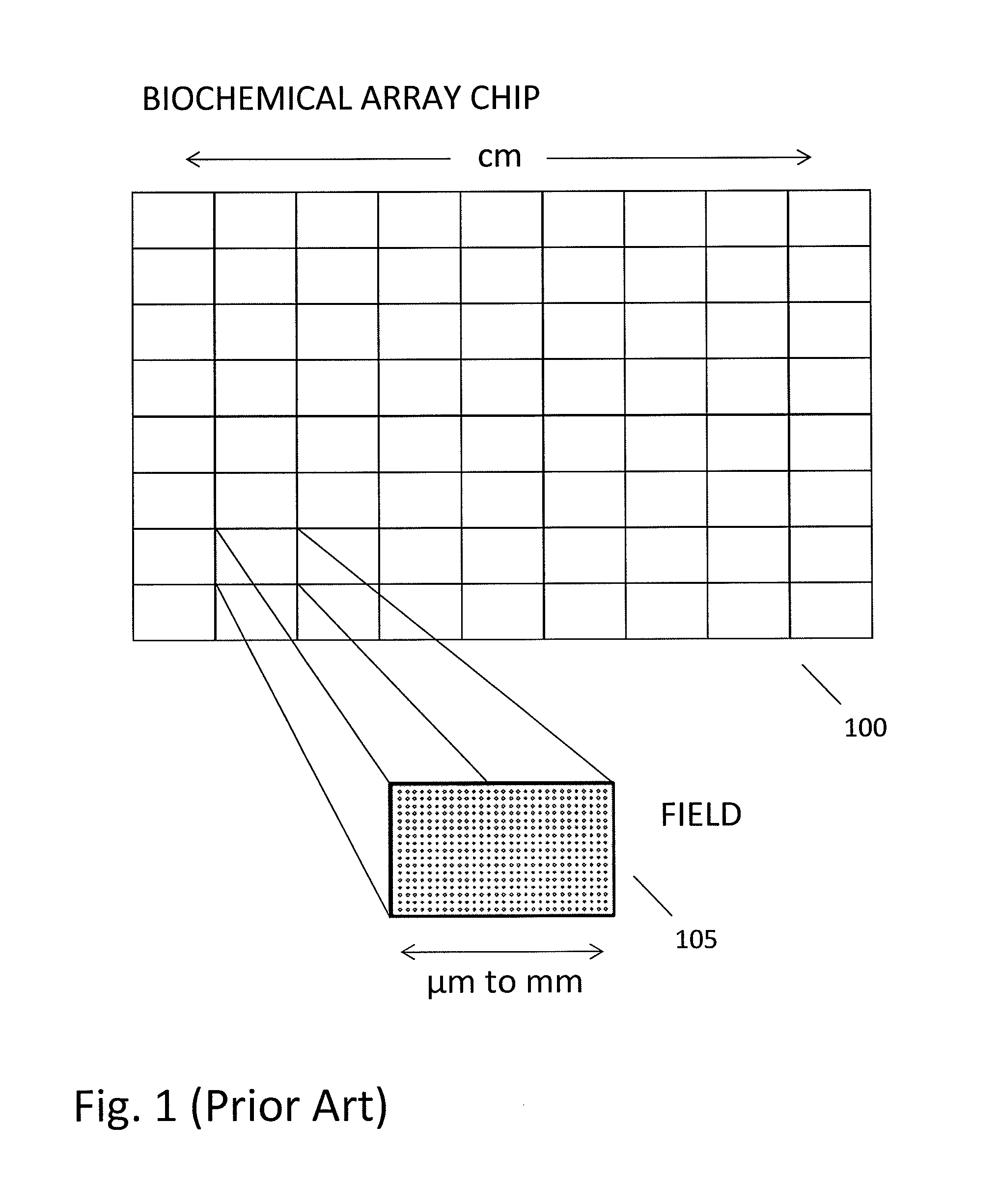
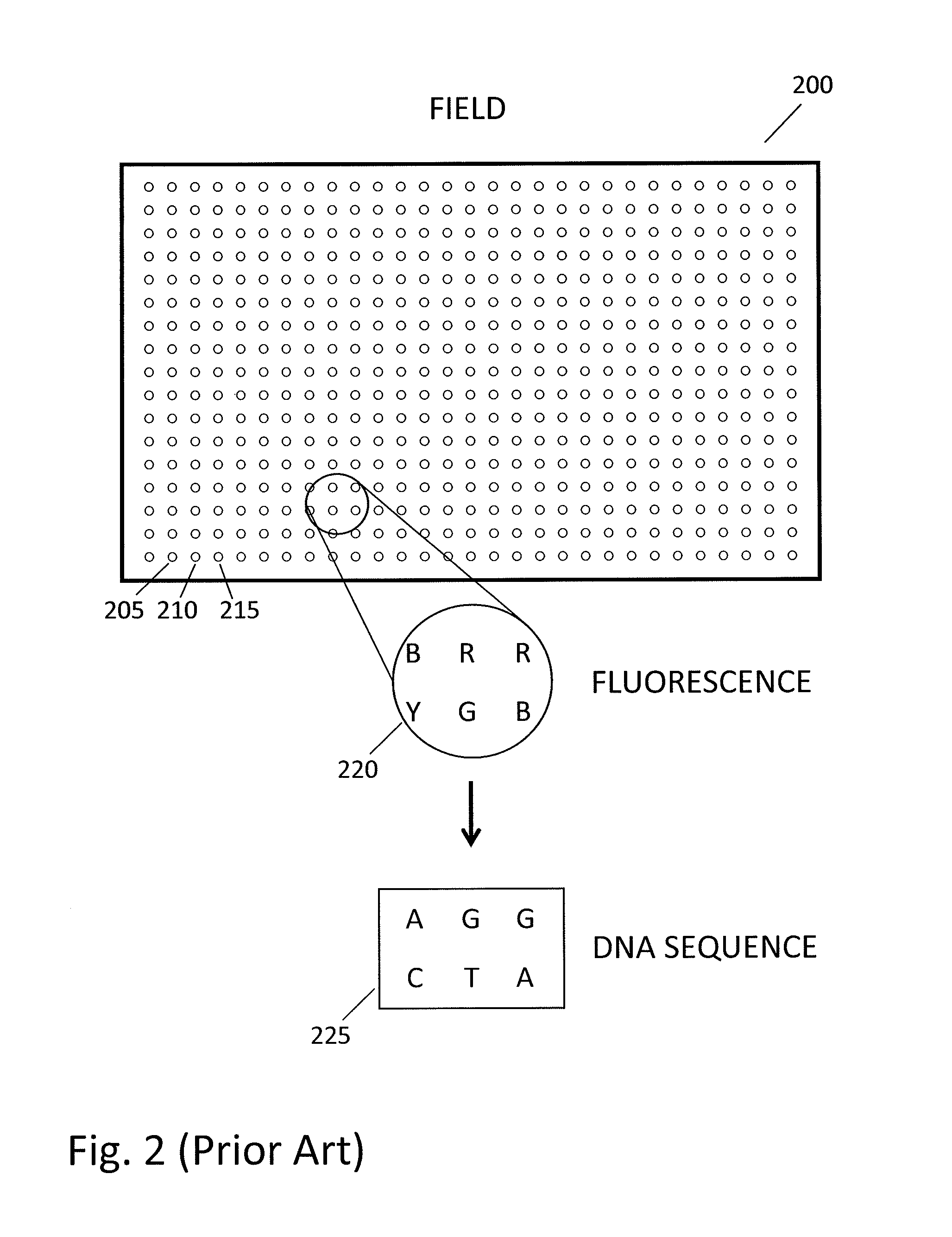
In a genome sequencing system and methodology, a protocol is provided to achieve precise alignment and accurate registration of an image of a planar array of nanoballs subject to optical analysis. Precise alignment correcting for fractional offsets is achieved by correcting for errors in subperiod x-y offset, scale and rotation by use of minimization techniques and Moiré averaging. In Moiré averaging, magnification is intentionally set so that the pixel period of the imaging element is a noninteger multiple of the site period. Accurate registration is achieved by providing for pre-defined pseudo-random sets of sites, herein deletion or reserved sites, where nanoballs are prevented from attachment to the substrate so that the sites of the array can be used in a pattern matching scheme as registration markers for absolute location identification. Information can be extracted with a high degree of confidence that it is correlated to a known location, while at the same time the amount of information that can be packed on a chip is maximized.
Owner:COMPLETE GENOMICS INC
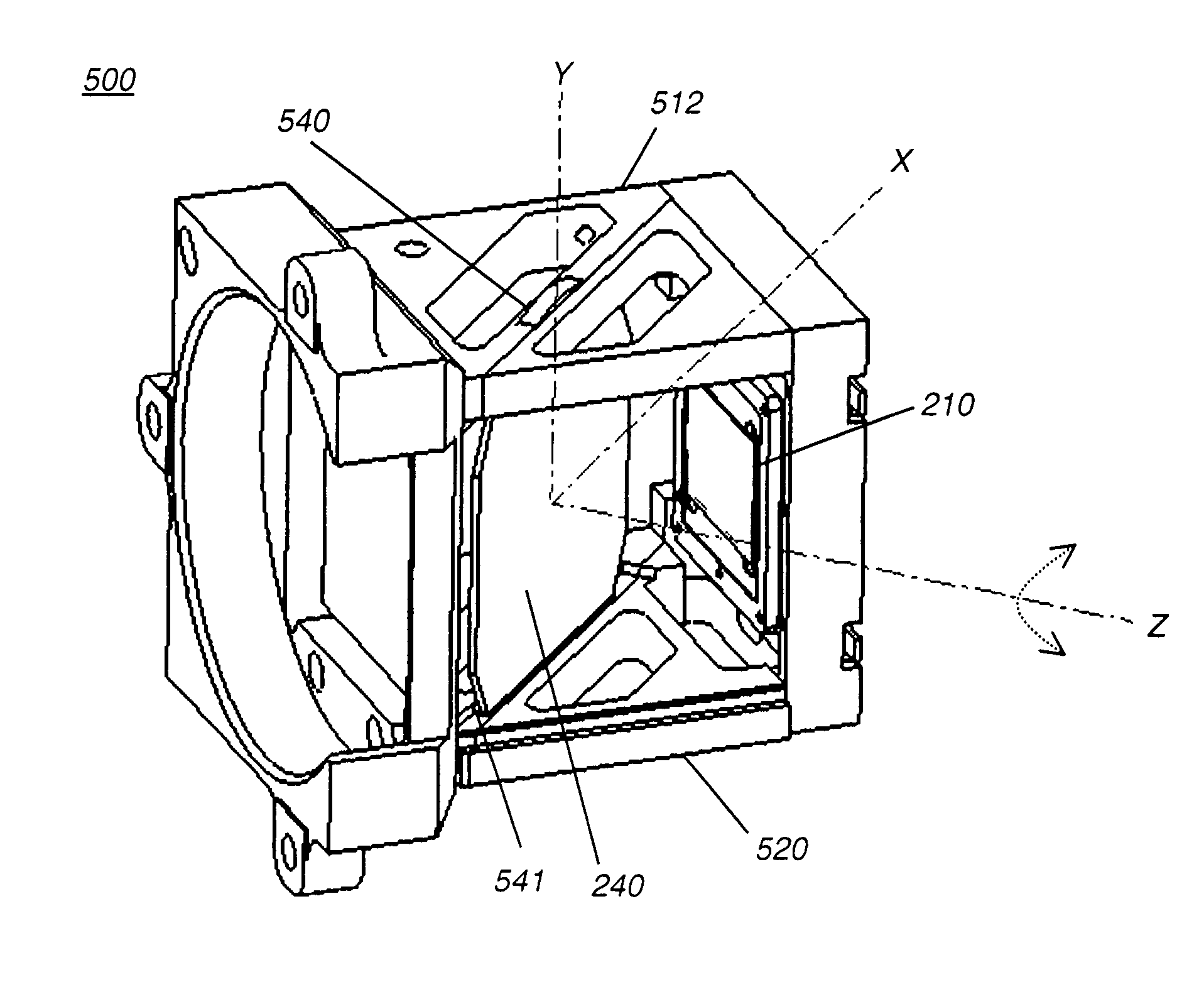
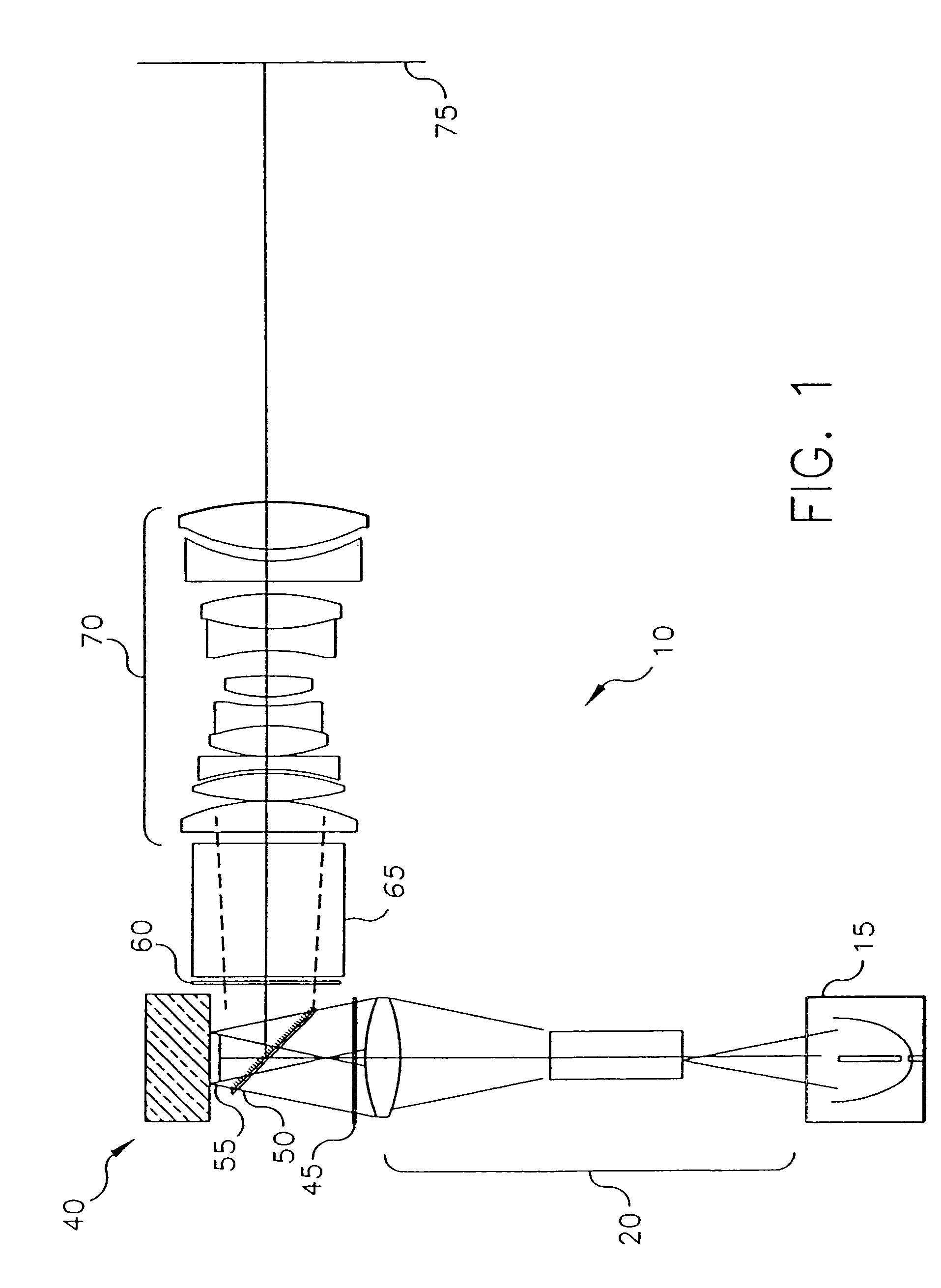
Housing for mounting a beamsplitter and a spatial light modulator with an output optical path
InactiveUS7131737B2Reduce image qualityMechanically robustTelevision system detailsProjectorsSpatial light modulatorWire grid
A housing (500) for a wire grid polarizing beamsplitter (240) and a spatial light modulator (210) in alignment with an output optical path has a front plate having an opening (502) for admitting incident illumination and a modulator mounting plate (506) for mounting the spatial light modulator (210) in the optical output path. First and second polarizer support plates (512, 520) extend between the front plate and the modulator mounting plate (506), with their respective facing inner surfaces providing coplanar support features for supporting the wire grid polarizing beamsplitter (240) within a fixed plane. The coplanar support features allow rotation of the wire grid polarizing beamsplitter (240) with respect to the output axis. The wire grid polarizing beamsplitter (240) has its surface at a fixed angle with respect to the surface of the spatial light modulator (210), the angle defining an output optical axis along the output optical path.
Owner:SONY CORP
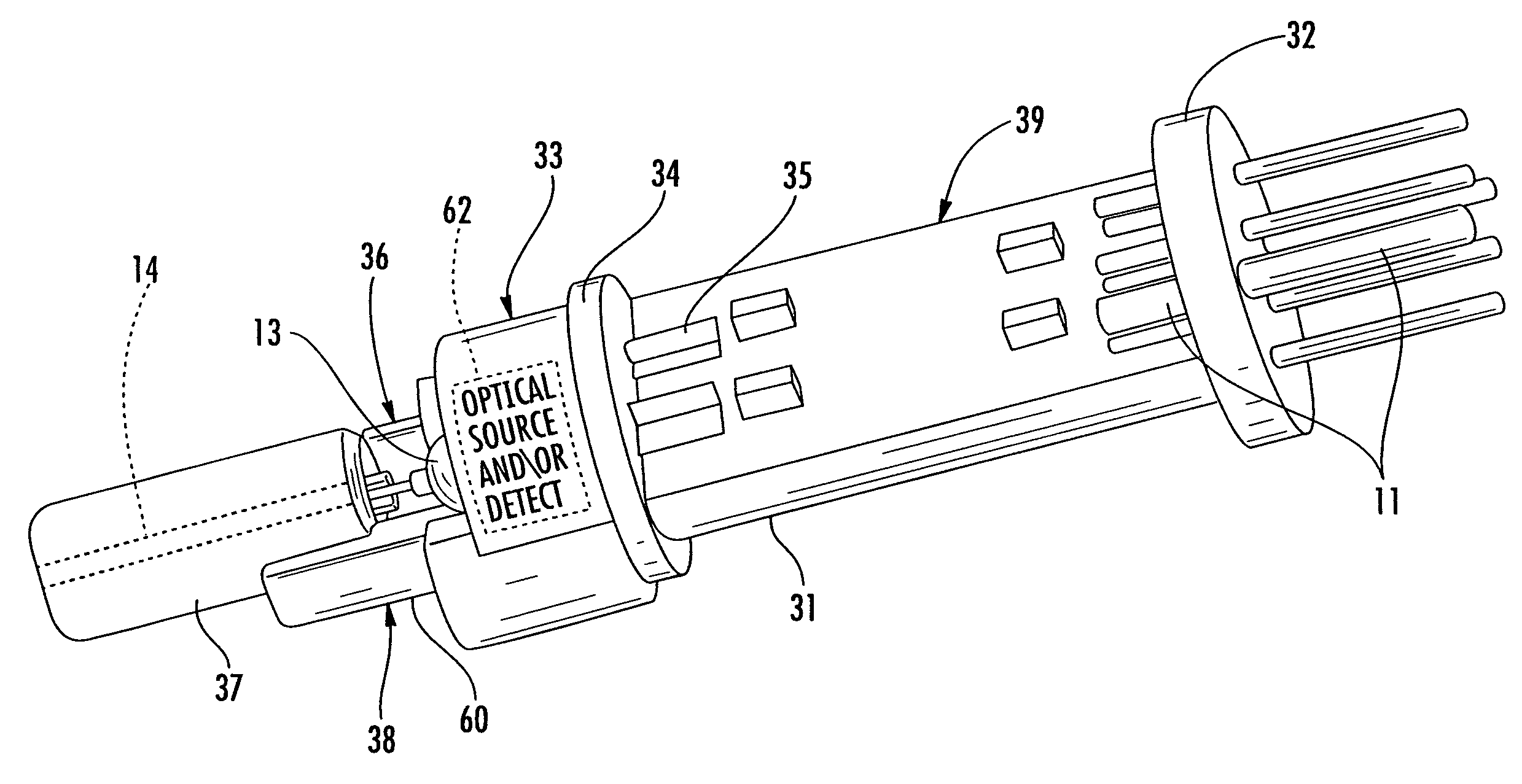
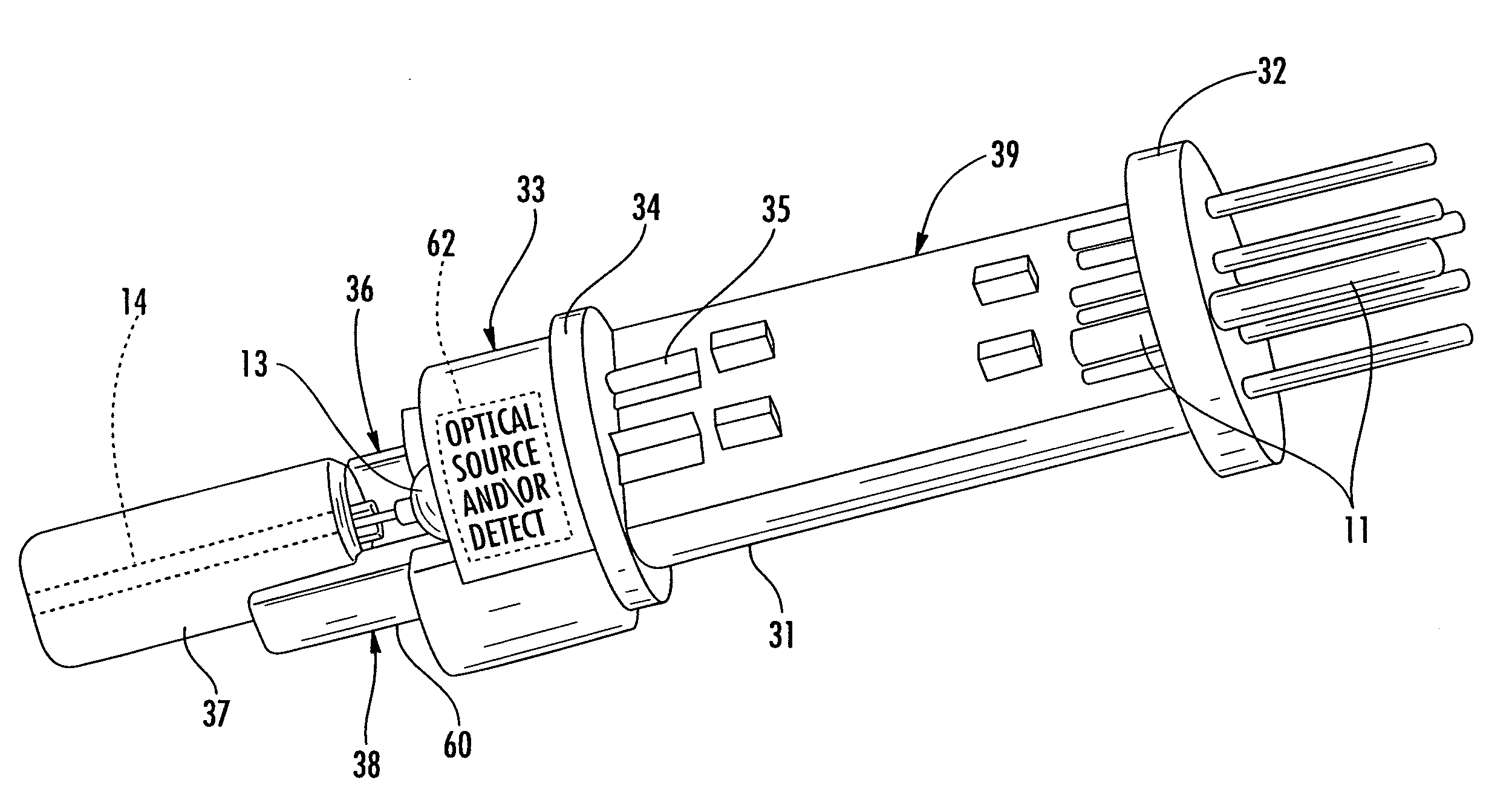
Fiber optic connector, active contact inserts therefor, and associated methods
ActiveUS7690849B2Easy to assembleEasy to manufactureCoupling light guidesElectromagnetic transmissionFiberEngineering
A fiber optic connector insert for a connector may include a housing to be received within a passageway of a connector body, a circuit board extending longitudinally within the housing and having opposing first and second ends, and a first base and first electrically conductive pins carried thereby. The first electrically conductive pins may be coupled to the circuit board adjacent the first end thereof. The fiber optic connector insert may also include an opto-electronic (O / E) converter having a second base and second electrically conductive pins extending outwardly therefrom. The second electrically conductive pins may be coupled to the circuit board adjacent the second end thereof.
Owner:MOOG INC
Method of forming stacked-die integrated circuit
ActiveUS7989959B1Mechanically robustSemiconductor/solid-state device detailsSolid-state devicesContact padElectrical connection
A stacked-die integrated circuit and a method of fabricating same. The stacked-die integrated circuit has circuitry formed in the first surface of a mother die, a plurality of through-die vias with at least one through-die via providing electrical connection between the circuitry of the mother die and the second surface and a plurality of contact pads formed in the second surface of the semiconductor die for mounting a daughter die wherein some of the contact pads are electrically isolated dummy pads.
Owner:XILINX INC
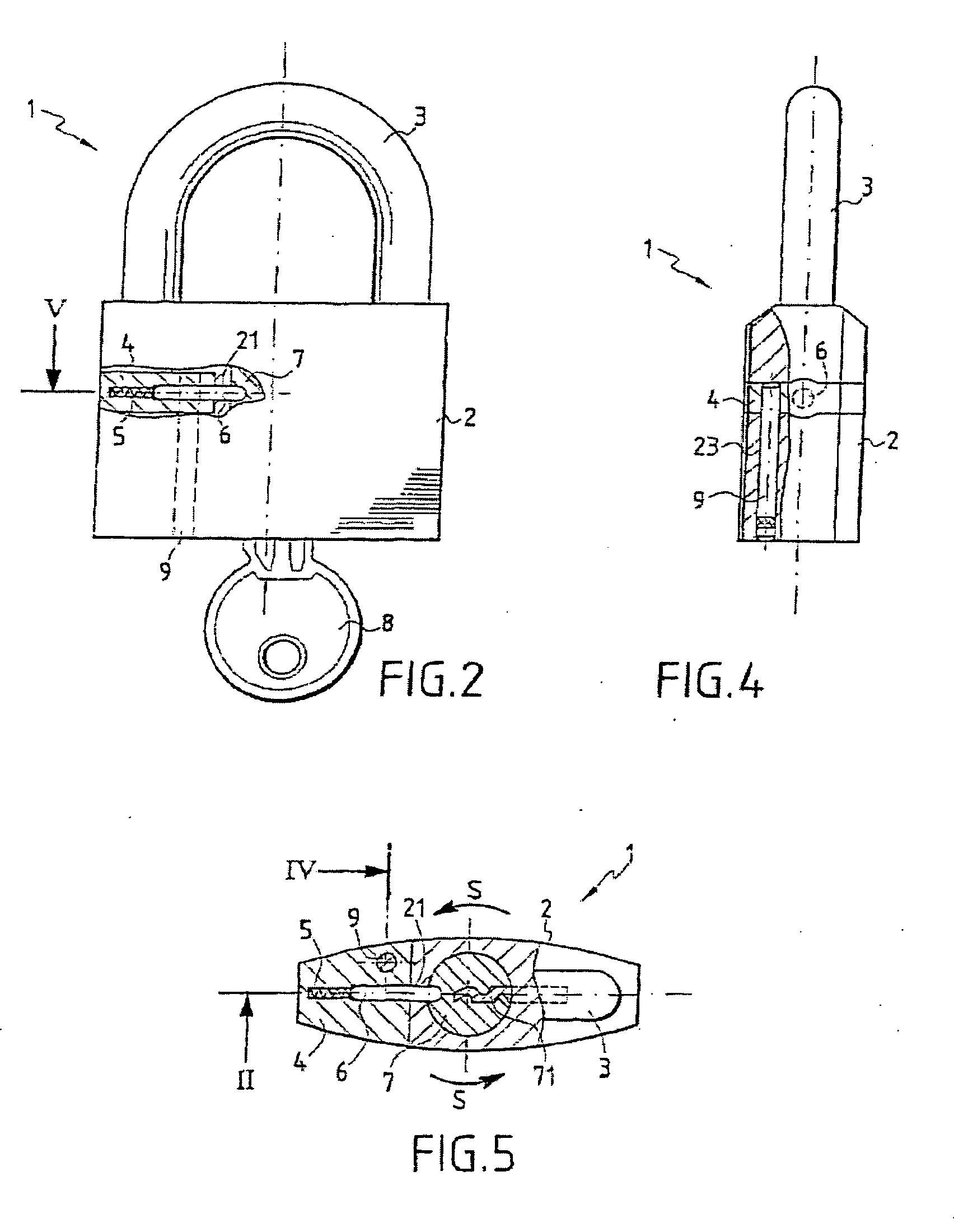
Multi-purpose seal with lock
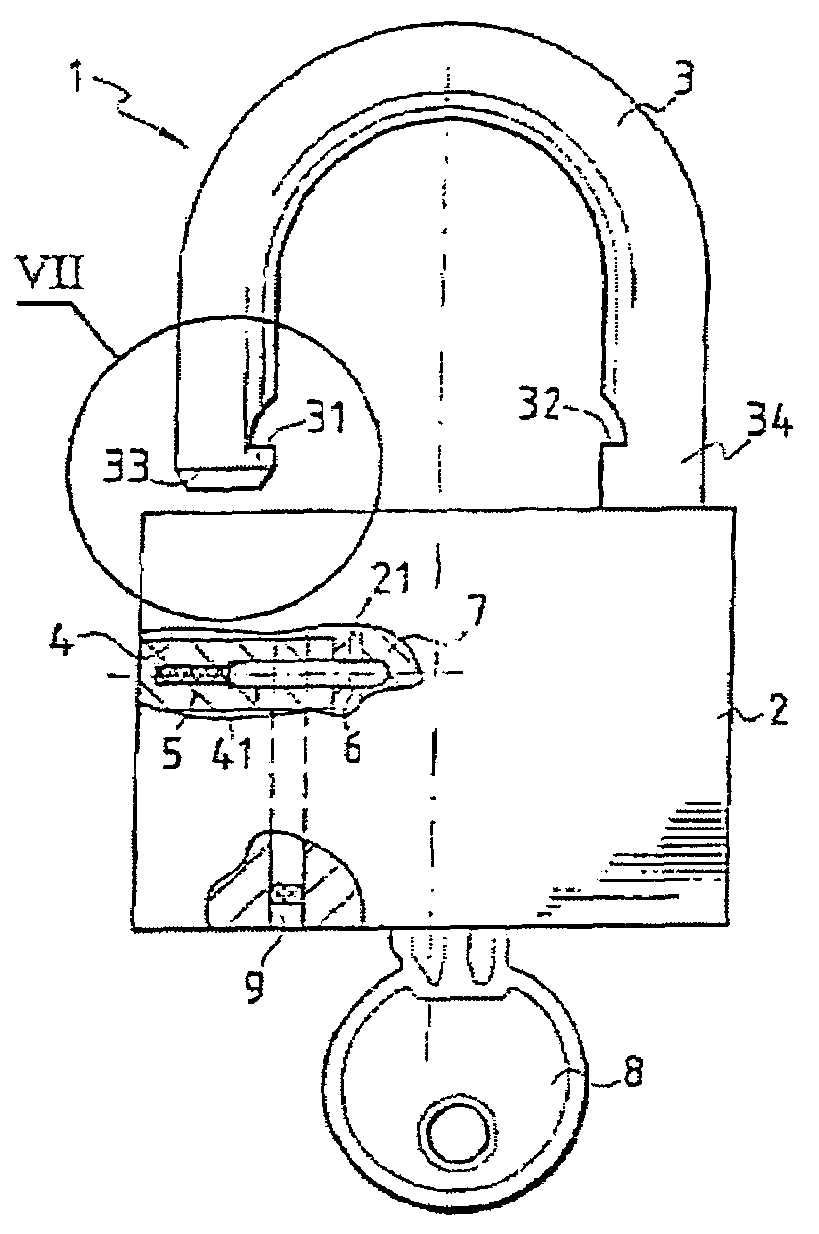
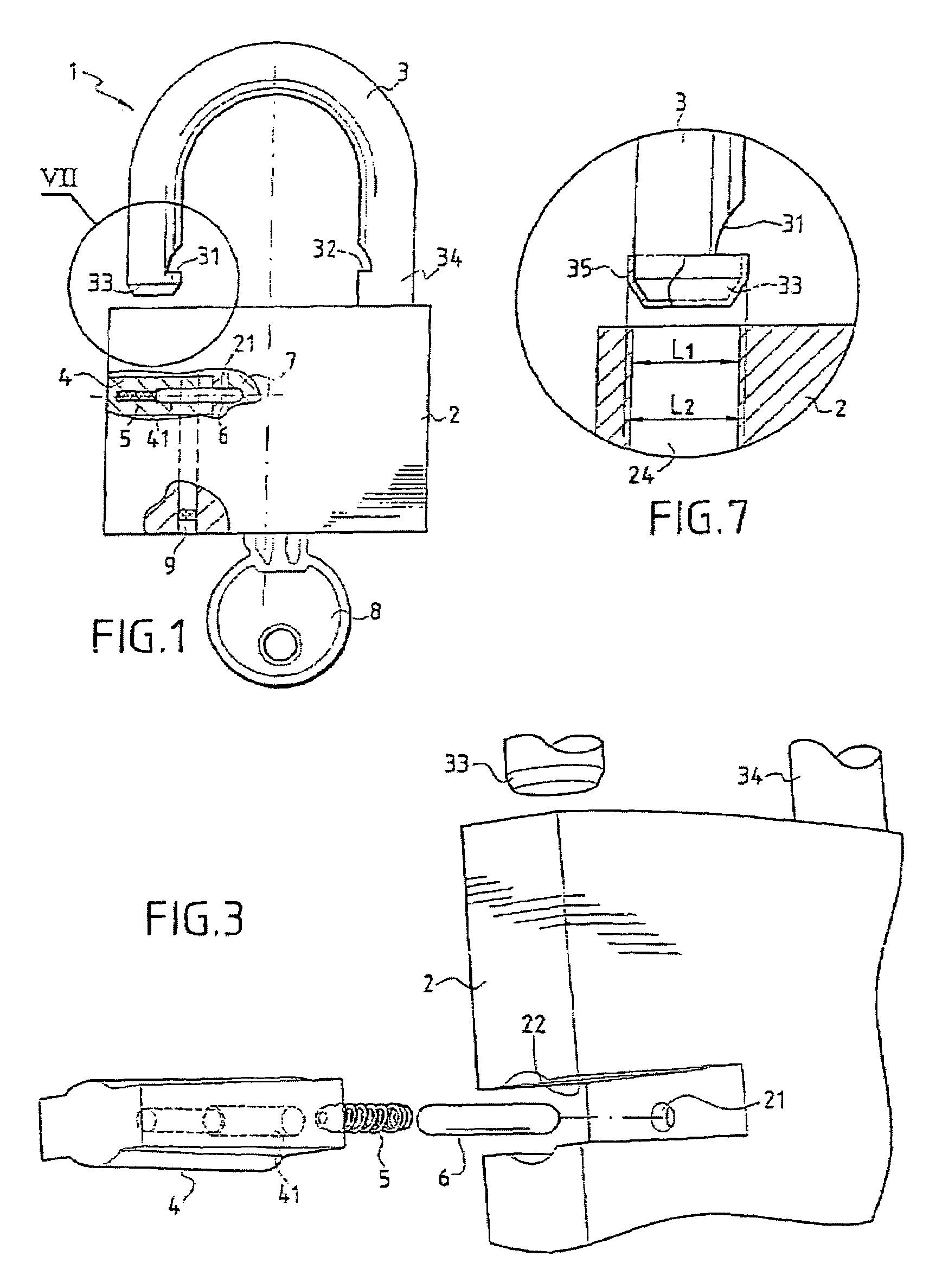
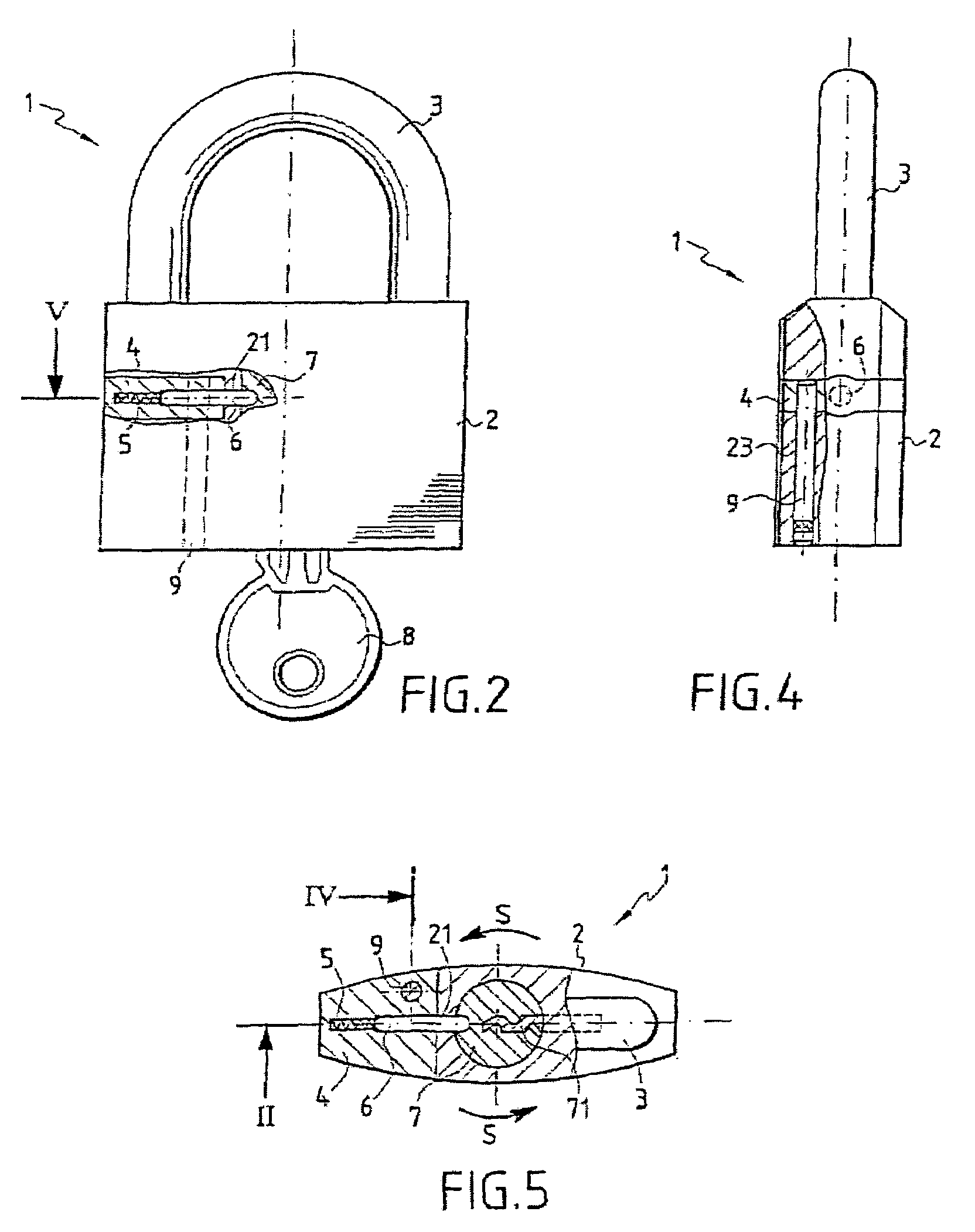
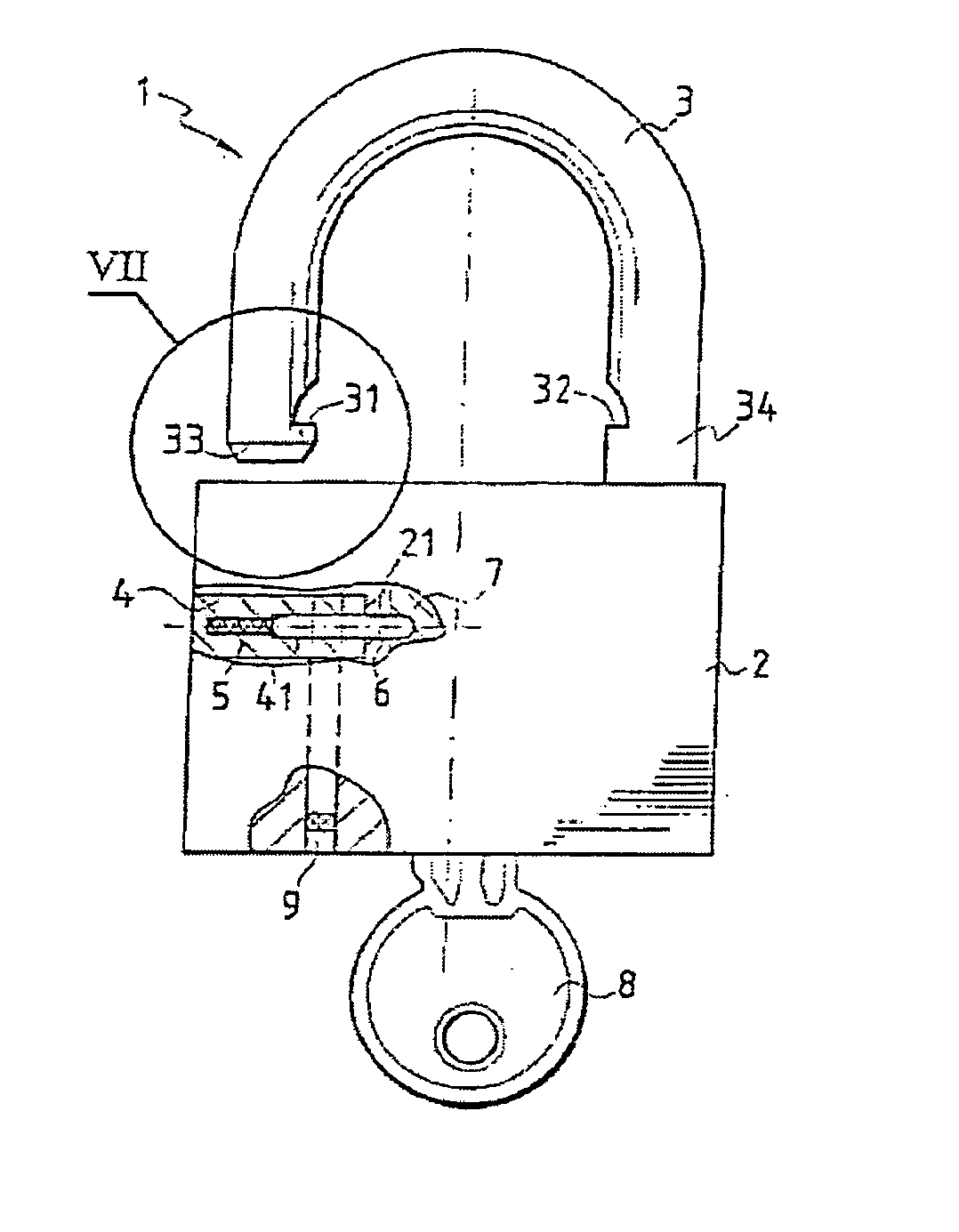
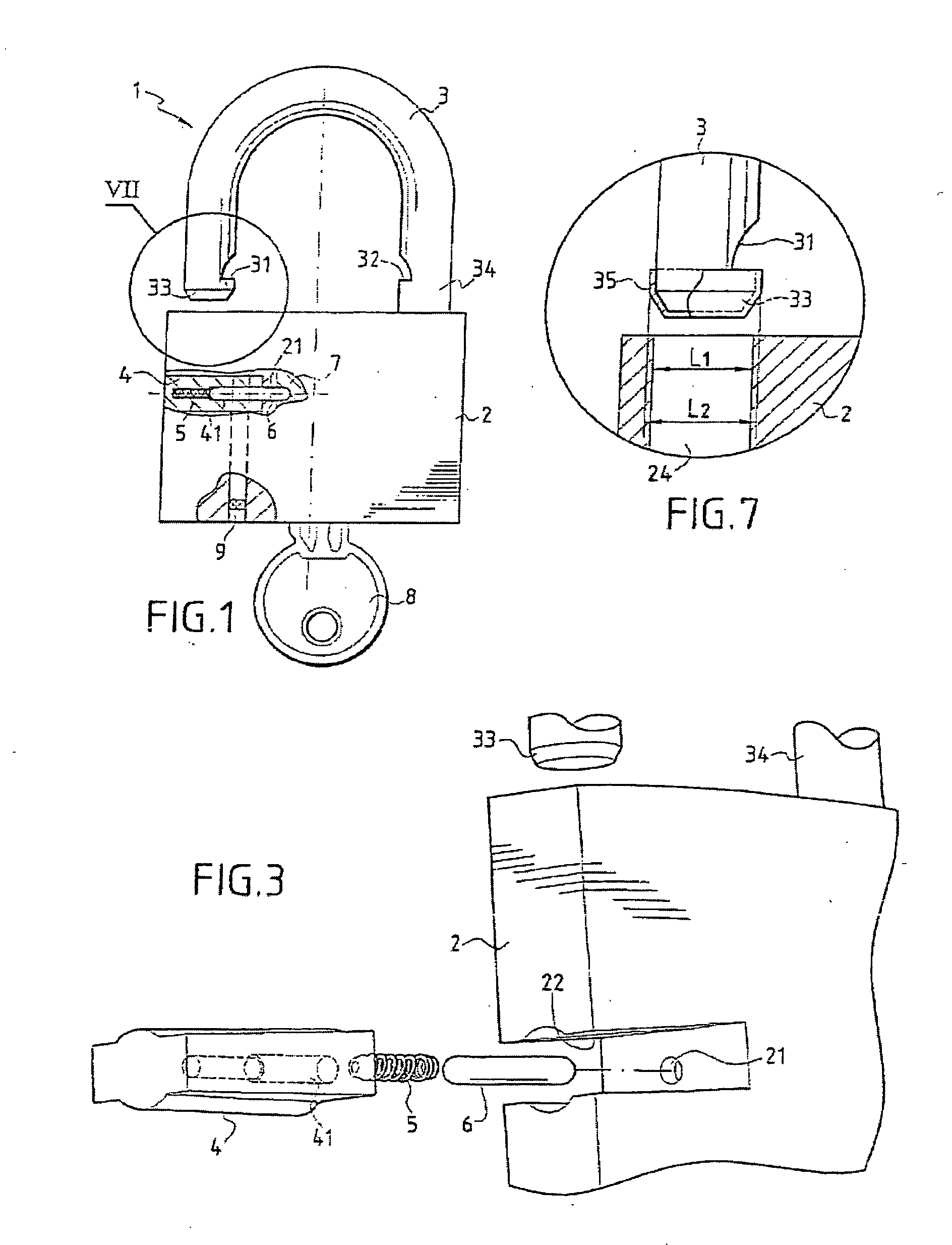
A multipurpose seal with a lock, including a mobile closure apparatus of a casing containing a lock and an attachment element designed to be locked in the casing. The lock includes a barrel configured to move in the casing between a locked position and an unlocked position. A remote-interrogatable electronic component including a data-storage is maintained in a housing, which passes through part of the casing and part of the lock barrel when the lock barrel is in the locked position. Thus, any rotation of the barrel from the locked position causes the component to break.
Owner:EURON COMMUNITY EC +1
Ceramic membranes with integral seals and support, and electrochemical cells and electrochemical cell stacks including the same
InactiveUS20070054169A1High mechanical strengthAvoid Diffusion LimitationFuel cells groupingActive material electrodesCeramic membranePorous electrode
Ceramic membranes with integral seals and support, and related electrochemical cells and cell stacks. The membrane comprises a thin electrolyte layer supported on a porous electrode layer which in turn is supported on a thick ceramic support layer, preferably a ceramic electrolyte support. The support layer is divided into a plurality of self-supporting thin membrane regions by a network of thicker integrated support ribs. The thin electrolyte layer and thick ceramic support layer preferably define a sealing perimeter surrounding the porous electrode layer.
Owner:NEXCERIS INNOVATION HLDG LLC
High density electrical connector and PCB footprint
ActiveUS20110212632A1Good signal integrityMechanically robustTwo-part coupling devicesElectrical connection printed elementsMechanical integrityPrinted circuit board
An interconnection system that includes a daughter card and backplane electrical connectors, each mounted to a printed circuit board at a connector footprint. The backplane connector has conductive elements with transition regions that allow the mating contact portions to be positioned on a uniform pitch while contact tail portions can be shaped to improve signal integrity or to provide a more compact and / or mechanically robust footprint. The conductive elements in both connectors are configured such that the contact tails of the ground conductors align from column to column, but the planar portions of the ground conductors in one column align with a pair of signal conductors in the other column, which improves mechanical and signal integrity. Mechanical integrity may be improved by forming the connector footprints with pads for the ground conductors that span multiple columns.
Owner:AMPHENOL CORP
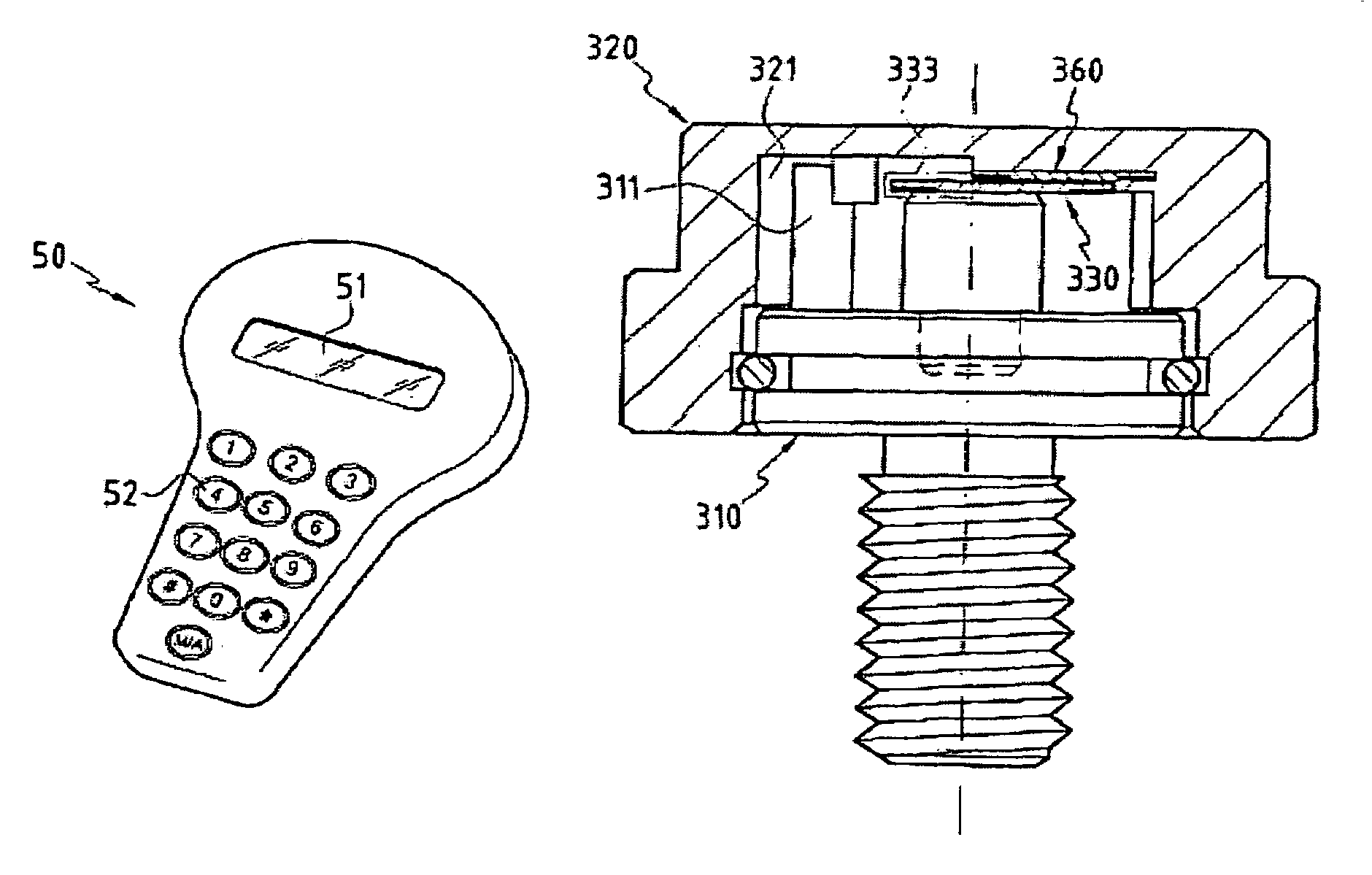
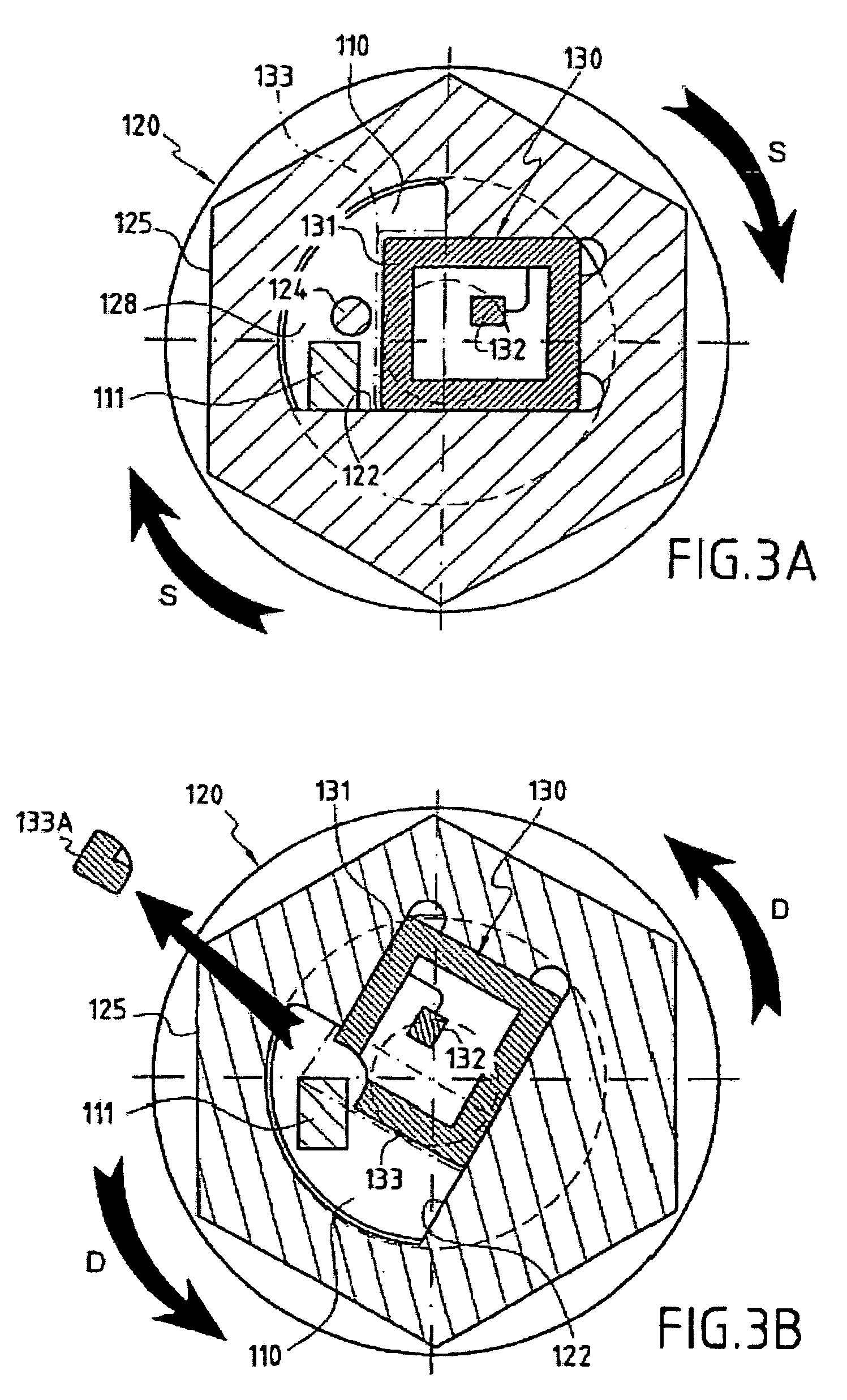
Fixing member comprising an integrity control system
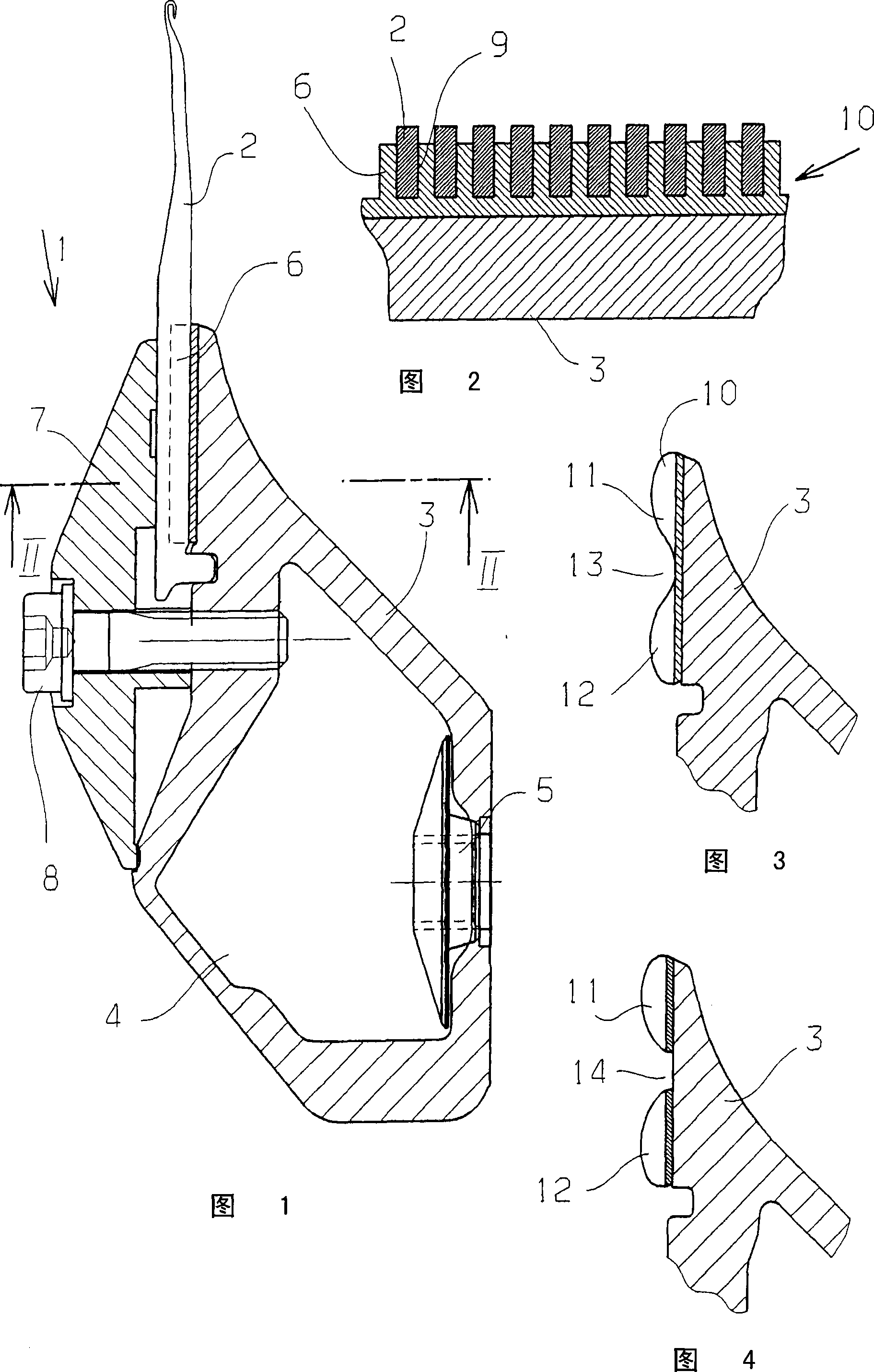
InactiveUS7246980B2Easily and quickly queriedMechanically robustStampsPinsControl systemElectronic component
The invention concerns a fastening member comprising a threaded part (150) and a head (100) with which the threaded part can be turned in order to tighten the fastening member in a corresponding element. The head (100) is formed of a plate (110) in one piece with the threaded part (150) and a cap (120) covering said plate. The cap (120) comprises a recessed portion (121) in which an offset is provided to form a housing (128). The plate (110) comprises a guide finger (111) located inside the housing (128) to form a stop against which the cap presses in order to impart a rotational movement to the plate in the tightening direction. The cap (120) further comprises a remotely interrogatable electronic component (130) comprising a data storage means (132). Said component is held in the recessed portion (121) with its end (133) protruding into the housing (128) so as to strike against the guide finger (111) and cause the fracture of the component when any attempt is made to unscrew or untighten the fastening member.
Owner:EURON COMMUNITY EC
Liquid cooled laser bar arrays incorporating diamond/copper expansion matched materials
ActiveUS7660335B2Improve thermal conductivityStress minimizationSemiconductor lasersLaser cooling arrangementsThermal expansionCopper
A laser diode array having a plurality of diode bars bonded by a hard solder to expansion matched spacers and mounted on a gas or liquid cooled heatsink. The spacers are formed of a material such as copper / diamond composite material having a thermal expansion that closely matches that of the laser bars.
Owner:LASERTEL
In-situ Forming foams with outer layer
ActiveUS20130317418A1Improve cohesionResistance to deformationStentsSurgical adhesivesMedical devicePolymer
Systems, methods and kits relating to in-situ forming polymer foams for the treatment of aneurysms or fluid filled spaces are disclosed. The systems include an insertable medical device and an in-situ forming foam of lava like materials with a fast forming outer skin and a slower hardening interior that is formed from a one-, two- or multi-part formulation. When used to treat an aneurysm, the foam is placed into contact with at least a portion of an exterior surface of the medical device and / or the tissue surface of the aneurysm.
Owner:ARSENAL MEDICAL
Method and system for accurate alignment and registration of array for DNA sequencing
ActiveUS8774494B2High precisionImprove trustImage enhancementImage analysisGenomic sequencingPattern matching
In a genome sequencing system and methodology, a protocol is provided to achieve precise alignment and accurate registration of an image of a planar array of nanoballs subject to optical analysis. Precise alignment correcting for fractional offsets is achieved by correcting for errors in subperiod x-y offset, scale and rotation by use of minimization techniques and Moiré averaging. In Moiré averaging, magnification is intentionally set so that the pixel period of the imaging element is a noninteger multiple of the site period. Accurate registration is achieved by providing for pre-defined pseudo-random sets of sites, herein deletion or reserved sites, where nanoballs are prevented from attachment to the substrate so that the sites of the array can be used in a pattern matching scheme as registration markers for absolute location identification. Information can be extracted with a high degree of confidence that it is correlated to a known location, while at the same time the amount of information that can be packed on a chip is maximized.
Owner:COMPLETE GENOMICS INC
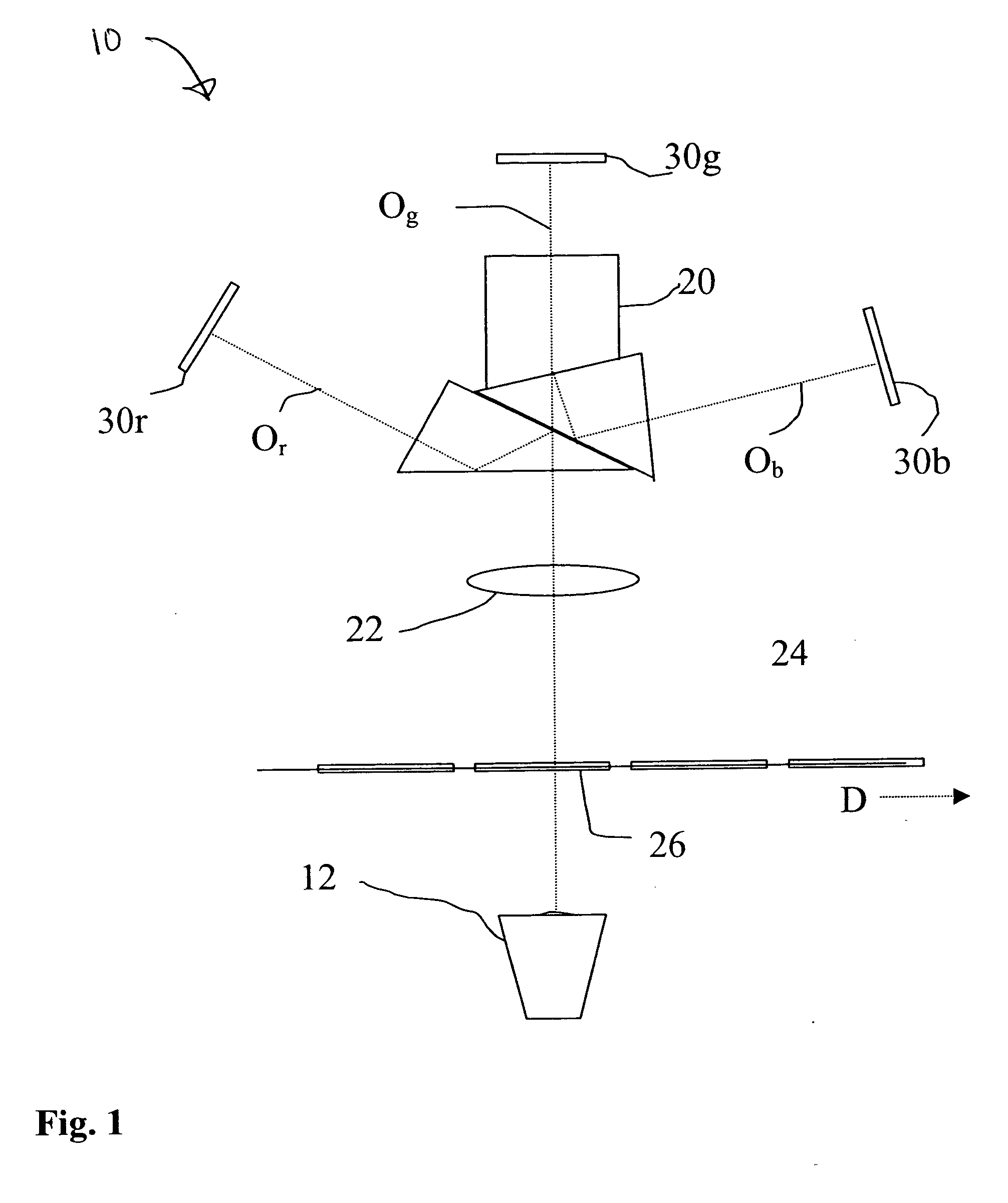
Scanner and method for operating a scanner
ActiveUS20060158662A1Manufacturing cost be reduceRobust constructionMirrorsCharacter and pattern recognitionSurface reliefLight beam
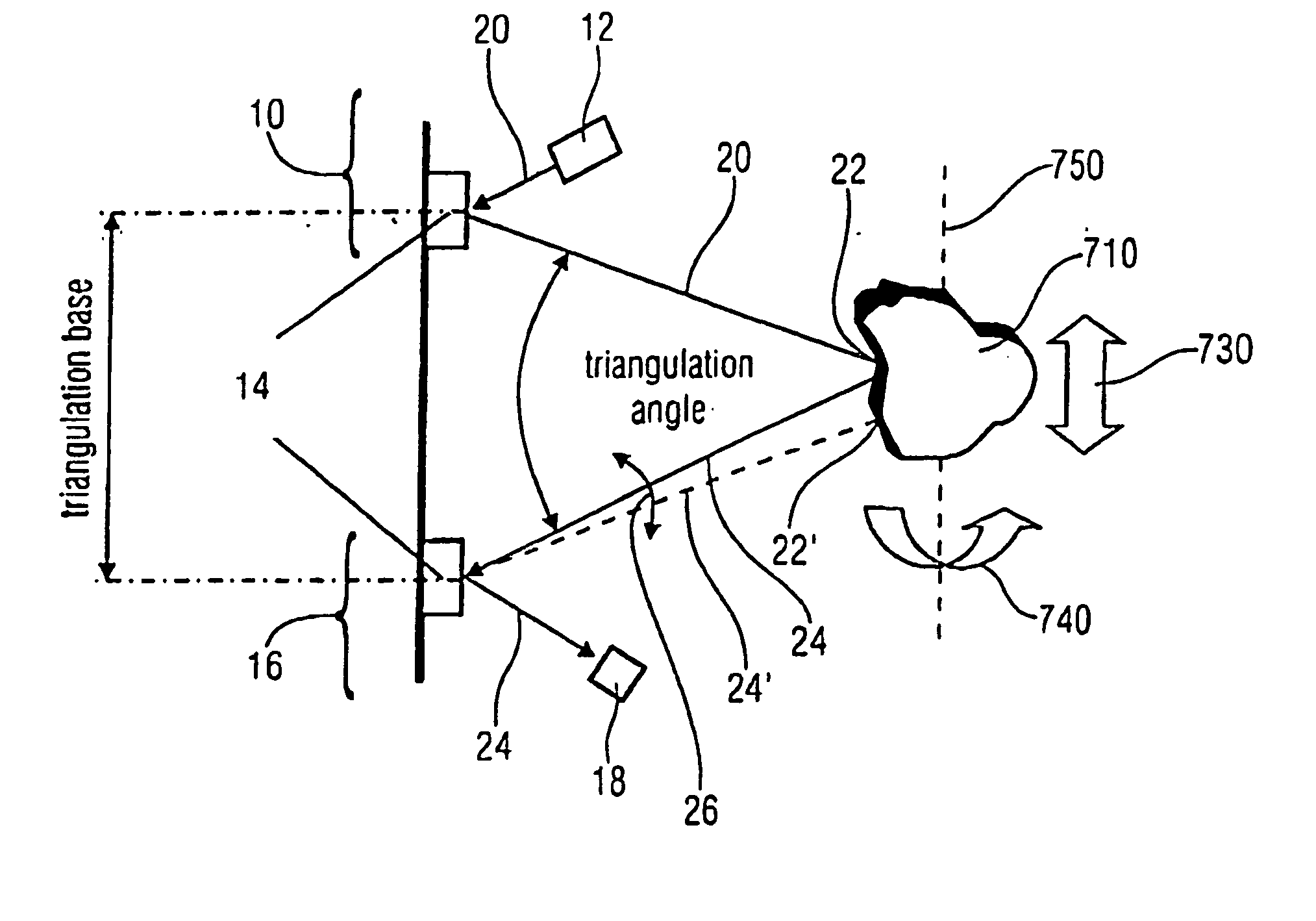
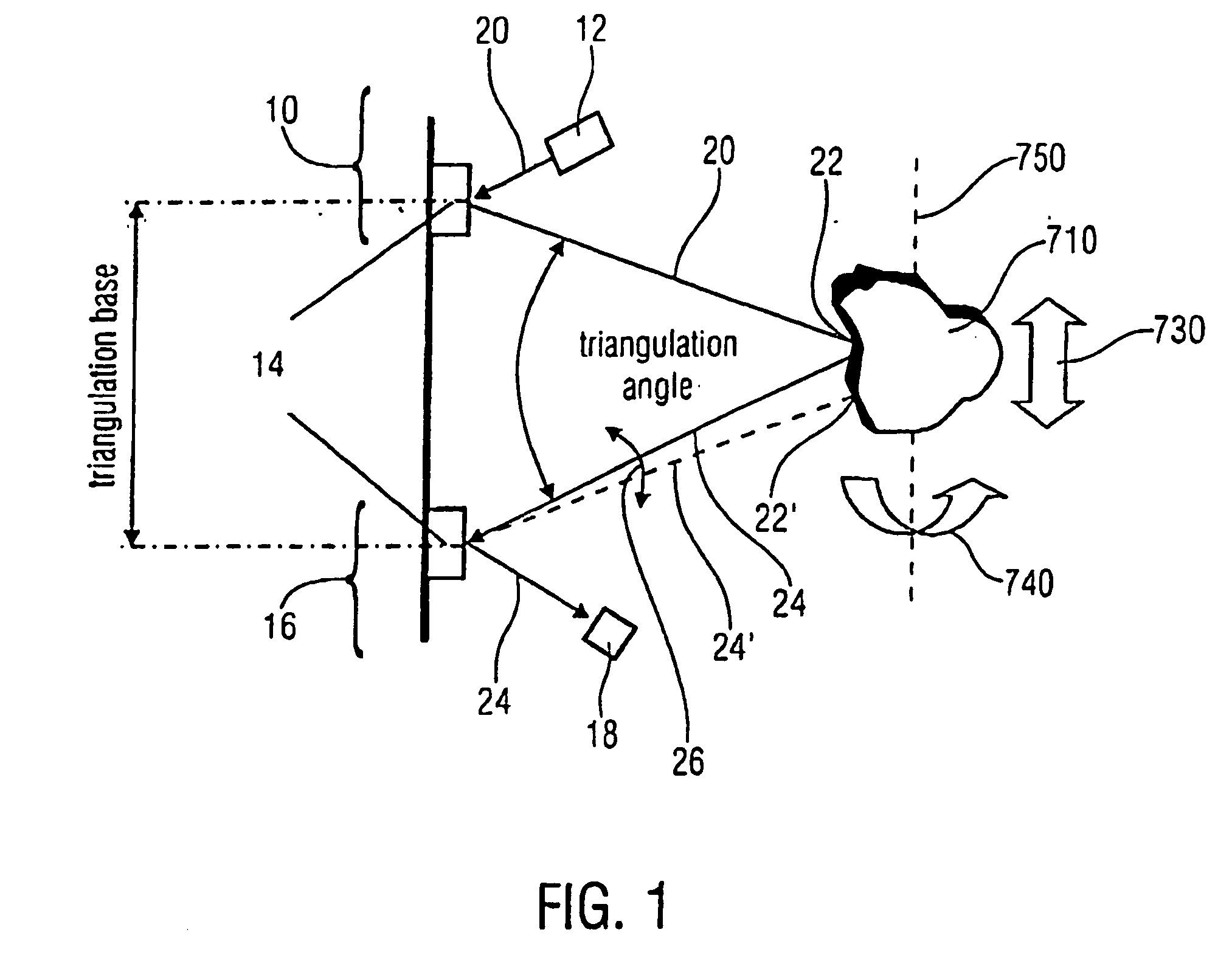
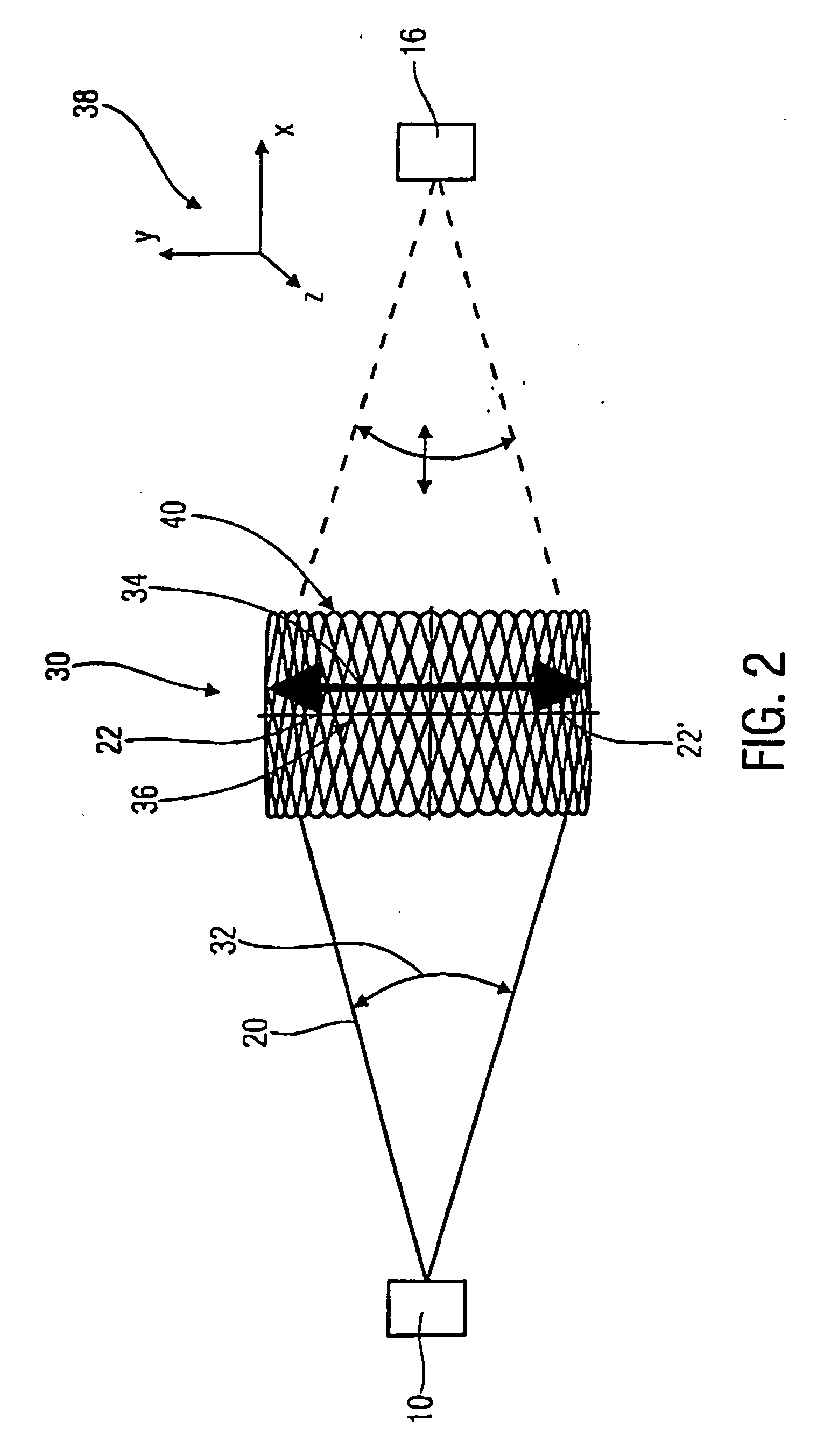
A scanner for providing a possibility of detecting a surface relief of an object includes a projector configured to guide a light beam in an illumination line over the surface relief to obtain an illuminated location on the surface relief, the projector being further configured to output a projection signal from which a position of the light beam in the illumination line is derivable. Additionally, the scanner includes a collector having a collector micro mirror stimulatable to oscillate in two dimensions and a point-shaped light detector, the collector micro mirror being oscillatingly arranged in a first direction of the illumination line and in a second direction differing from the first direction such that a reflection of the illuminated location within a scan area of the micro scanner mirror is imagable by it onto the point-shaped light detector, and the collector being configured to output a detection signal from which a position of the illuminated location in the first and second directions is derivable.
Owner:FRAUNHOFER GESELLSCHAFT ZUR FOERDERUNG DER ANGEWANDTEN FORSCHUNG EV
Multi-purpose seal with lock
A multipurpose seal with a lock, including a mobile closure apparatus of a casing containing a lock and an attachment element designed to be locked in the casing. The lock includes a barrel configured to move in the casing between a locked position and an unlocked position. A remote-interrogatable electronic component including a data-storage is maintained in a housing, which passes through part of the casing and part of the lock barrel when the lock barrel is in the locked position. Thus, any rotation of the barrel from the locked position causes the component to break.
Owner:EURON COMMUNITY EC +1
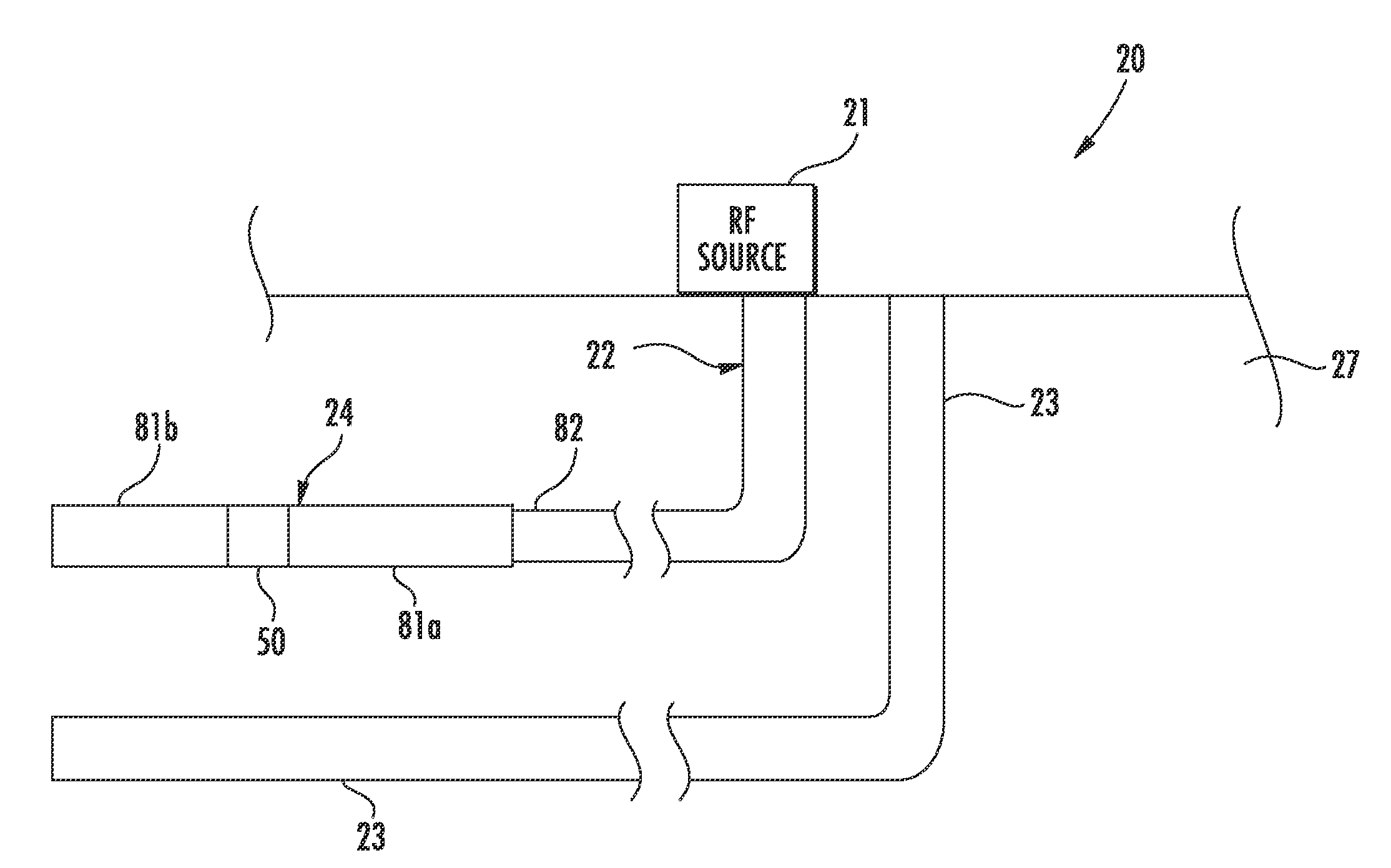
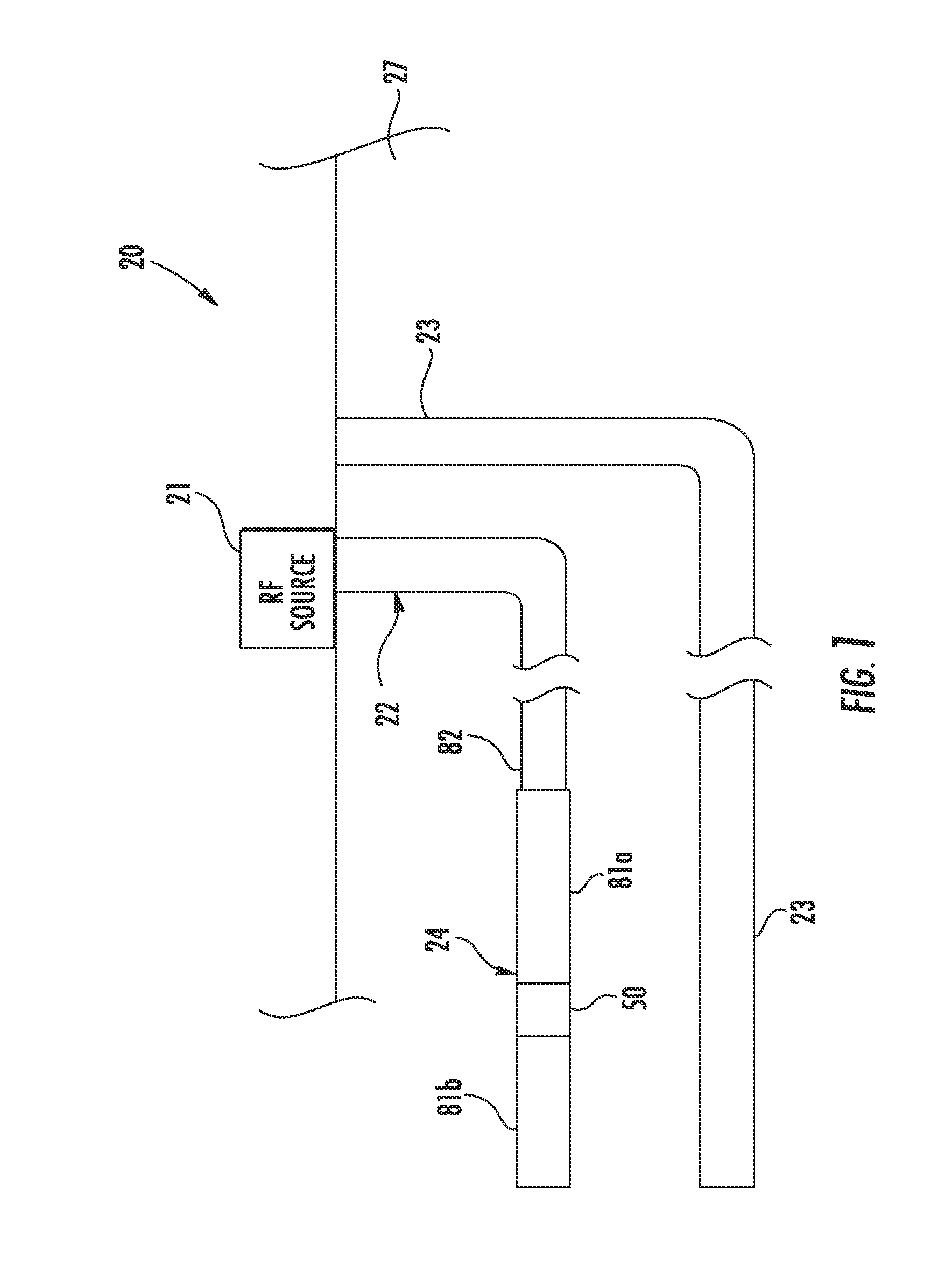
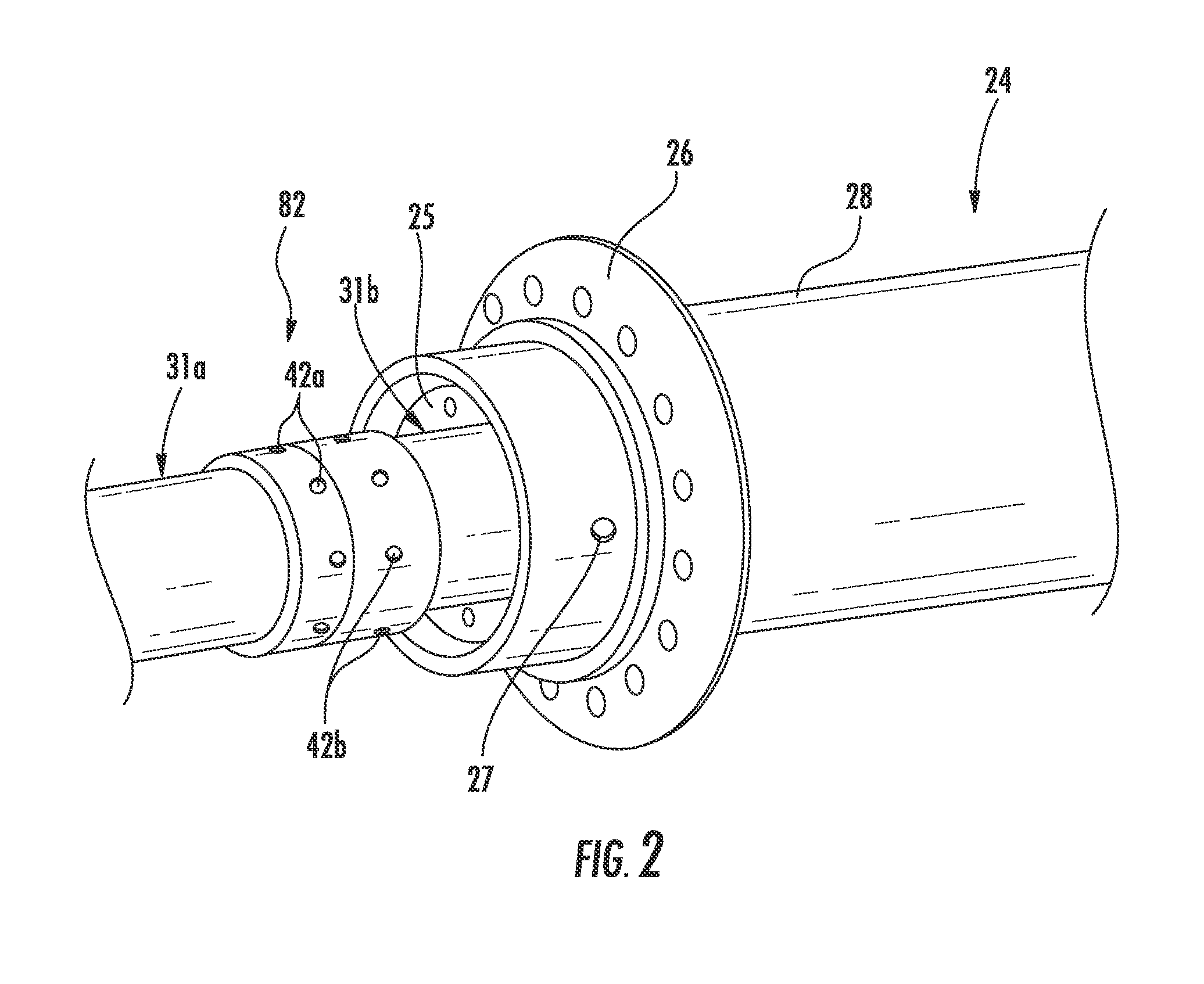
RF antenna assembly with dielectric isolator and related methods
ActiveUS20140262223A1Physically flexiblePhysically robustConstructionsInsulationElectrical conductorEngineering
An RF antenna assembly is positioned within a wellbore in a subterranean formation for hydrocarbon resource recovery. The RF antenna assembly includes first and second tubular conductors and a dielectric isolator therebetween. The dielectric isolator includes a dielectric tube having opposing first and second open ends, a first tubular connector having a first slotted recess receiving therein the first open end of the dielectric tube, and a second tubular connector having a second slotted recess receiving therein the second open end of the dielectric tube.
Owner:HARRIS CORP
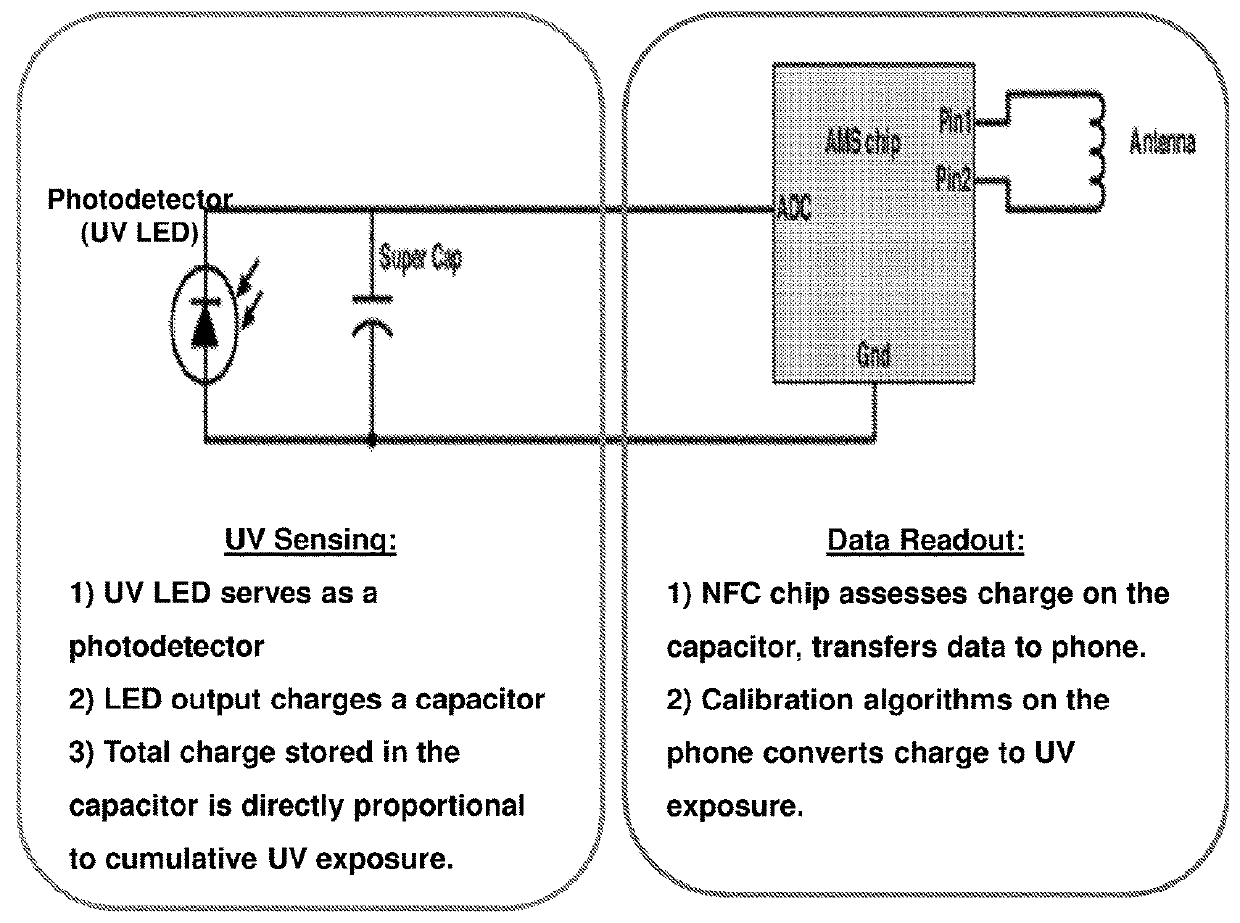
Alternative approach for UV sensing
ActiveUS20180274973A1Operation and useImprove performanceCircuit arrangementsDiagnostics using spectroscopyFluenceUltraviolet
The invention provides systems and methods for wearable and tissue-mounted electronics for monitoring exposure of a subject or object to electromagnetic radiation, particularly electromagnetic radiation in the visible, ultraviolet and infrared portions of the electromagnetic spectrum. In some embodiments, the devices are purely passive devices where absorption of incident electromagnetic radiation by the device provides at least a portion of the power for the measurement of the radiant exposure or flux of the incident electromagnetic radiation. Devices of the invention may include near field communication components, for example, for enabling readout by an external device, such as a computer or mobile device.
Owner:THE BOARD OF TRUSTEES OF THE UNIV OF ILLINOIS
Fiber optic connector, active contact inserts therefor, and associated methods
ActiveUS20080285923A1Easy to assembleEasy to manufactureCoupling light guidesElectromagnetic transmissionFiberOpto electronic
A fiber optic connector insert for a connector may include a housing to be received within a passageway of a connector body, a circuit board extending longitudinally within the housing and having opposing first and second ends, and a first base and first electrically conductive pins carried thereby. The first electrically conductive pins may be coupled to the circuit board adjacent the first end thereof. The fiber optic connector insert may also include an opto-electronic (O / E) converter having a second base and second electrically conductive pins extending outwardly therefrom. The second electrically conductive pins may be coupled to the circuit board adjacent the second end thereof.
Owner:MOOG INC
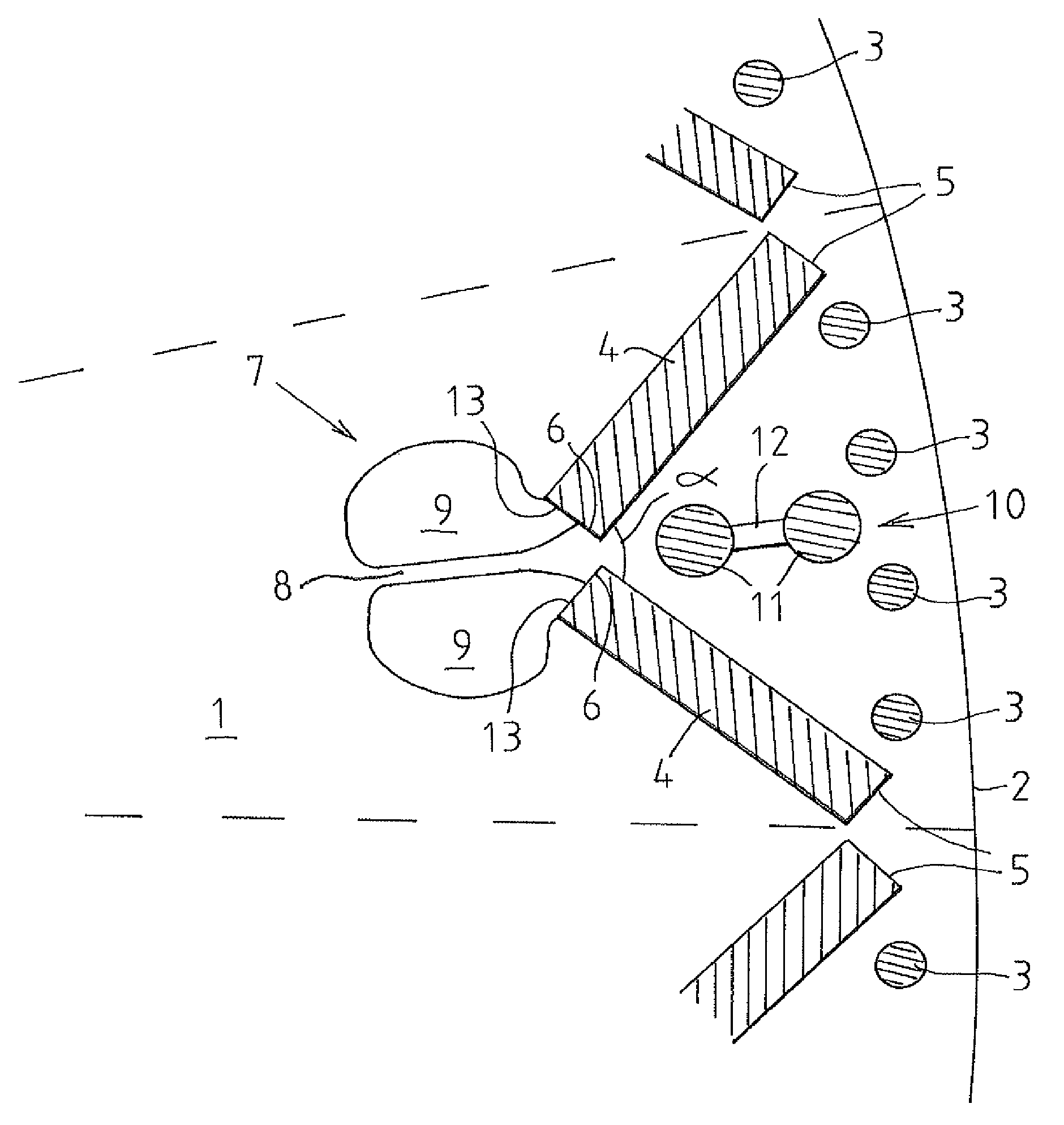
Laminated rotor structure for a permanent magnet synchronous machine
ActiveUS9178399B2Avoid overall overheatingMinimize leakage fluxMagnetic circuit rotating partsCooling/ventillation arrangementPermanent magnet synchronous machineEngineering
The invention relates to a laminated rotor structure for a permanent magnet synchronous machine wherein disks (1) of a ferromagnetic material constitute the body of the rotor. The body includes bars (3) of a damper winding that extend axially close to the surface (2) from one end thereof to the other, and a circle of permanent magnets (4) disposed in a V-shaped configuration inside the circle formed by the bars, in which permanent magnets the first ends (5) are disposed close to the outer perimeter of the rotor and the second ends (6) are disposed closer to the central axis of the rotor. According to the invention, two permanent magnets (4) constitute a pair of permanent magnets wherein the magnets are disposed at an angle (a) to each other so that their first ends (5) are disposed at a distance from each other and their second ends (6) in proximity to each other. In addition, a pair of permanent magnets includes an air channel (7) that extends axially through the laminar structure of the rotor in direct heat transfer contact with the second ends (6) of the magnets in the pair of magnets.
Owner:DANFOSS EDITRON OY
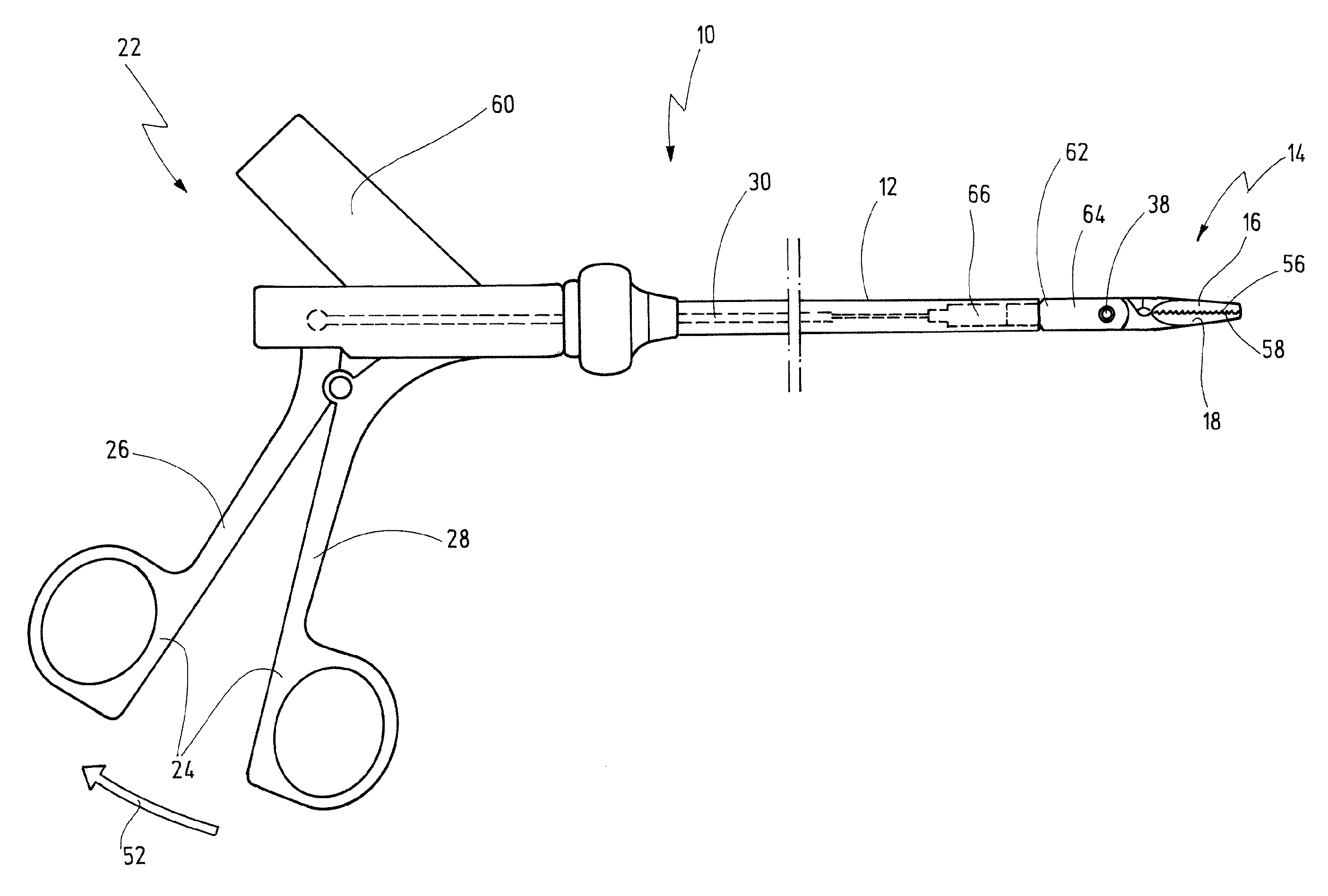
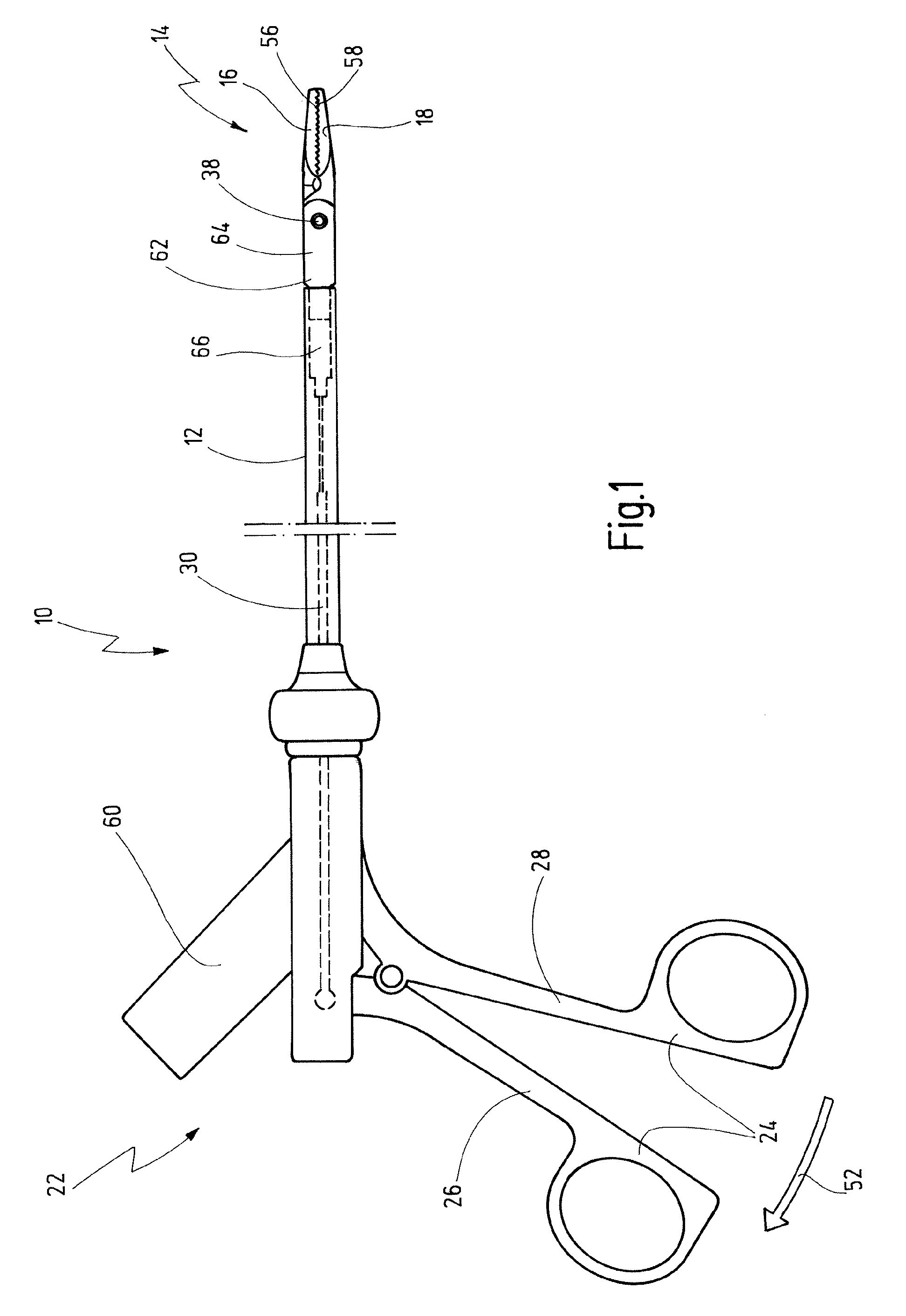
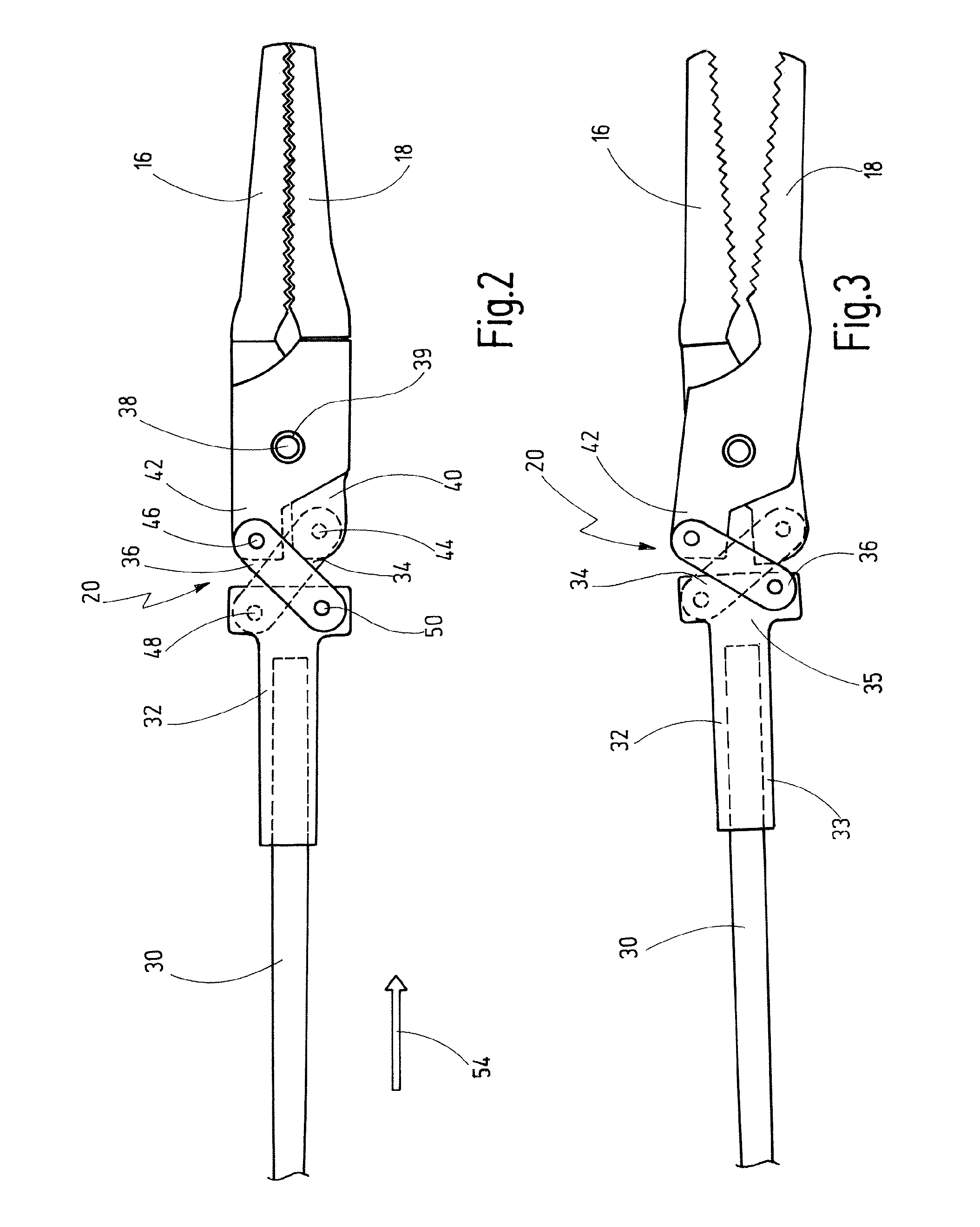
Bipolar Medical Instrument
ActiveUS20090082768A1Large separationAvoid voltage flashoverSurgical instruments for heatingSurgical forcepsEngineeringContact element
A bipolar medical instrument has a shaft and two jaw parts arranged at said distal end thereof. An axially movable force transmission element is disposed with said shaft and is surrounded by an isolating element. Said isolating element is connected to at least one of said two jaw parts in an articulated manner. Said two jaw parts are electrically isolated from one another and each provides an electrode to which high frequency current can be applied. A first electrical supply line is formed along said shaft via an electrically conductive first contact element to one of said two jaw parts. A second electrical supply line is formed along said axially movable force transmission element via an electrically conductive second contact element to the other of said two jaw parts. The first contact element is fixed at said isolating element surrounding said distal end area of said force transmission element.
Owner:KARL STORZ GMBH & CO KG
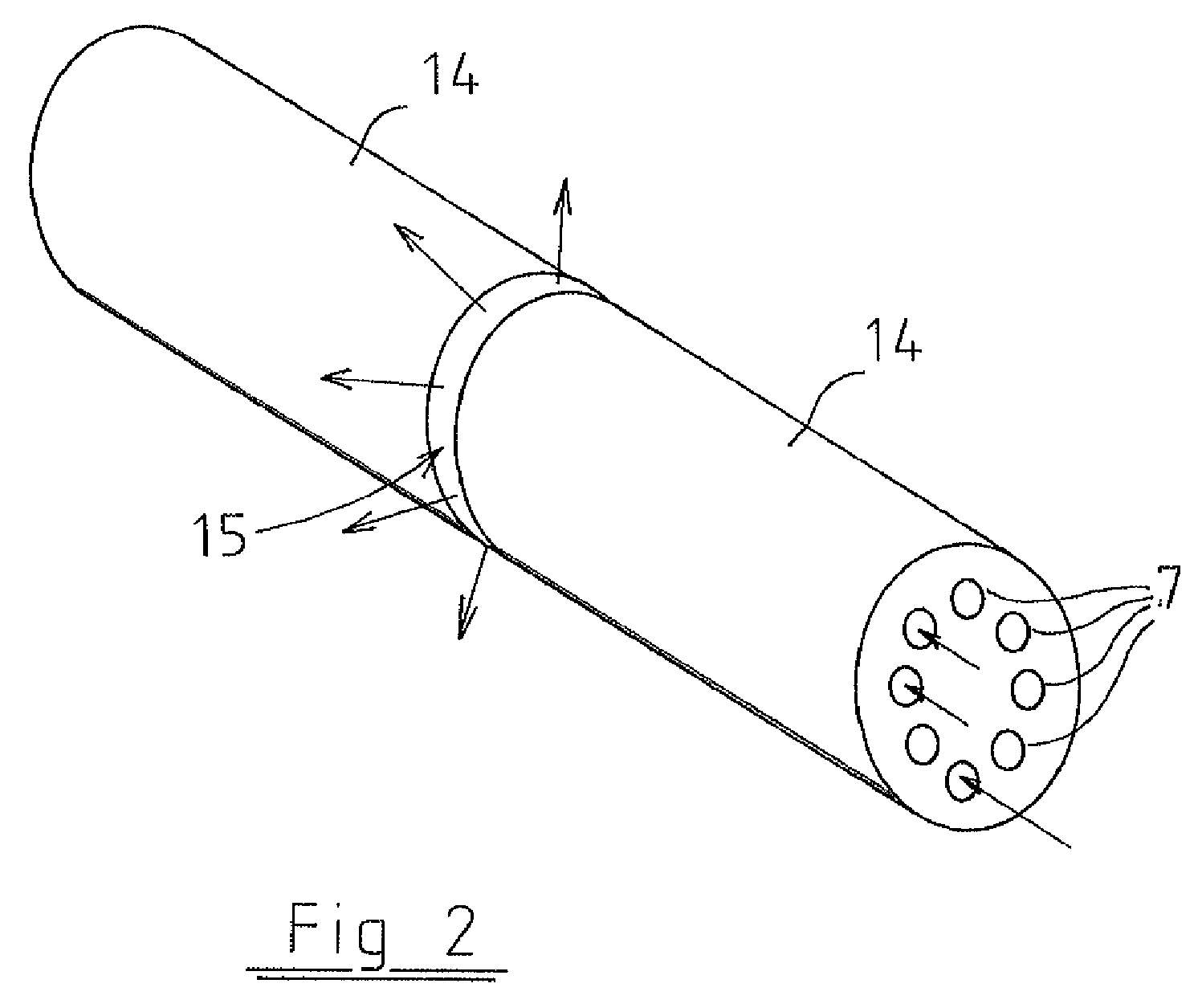
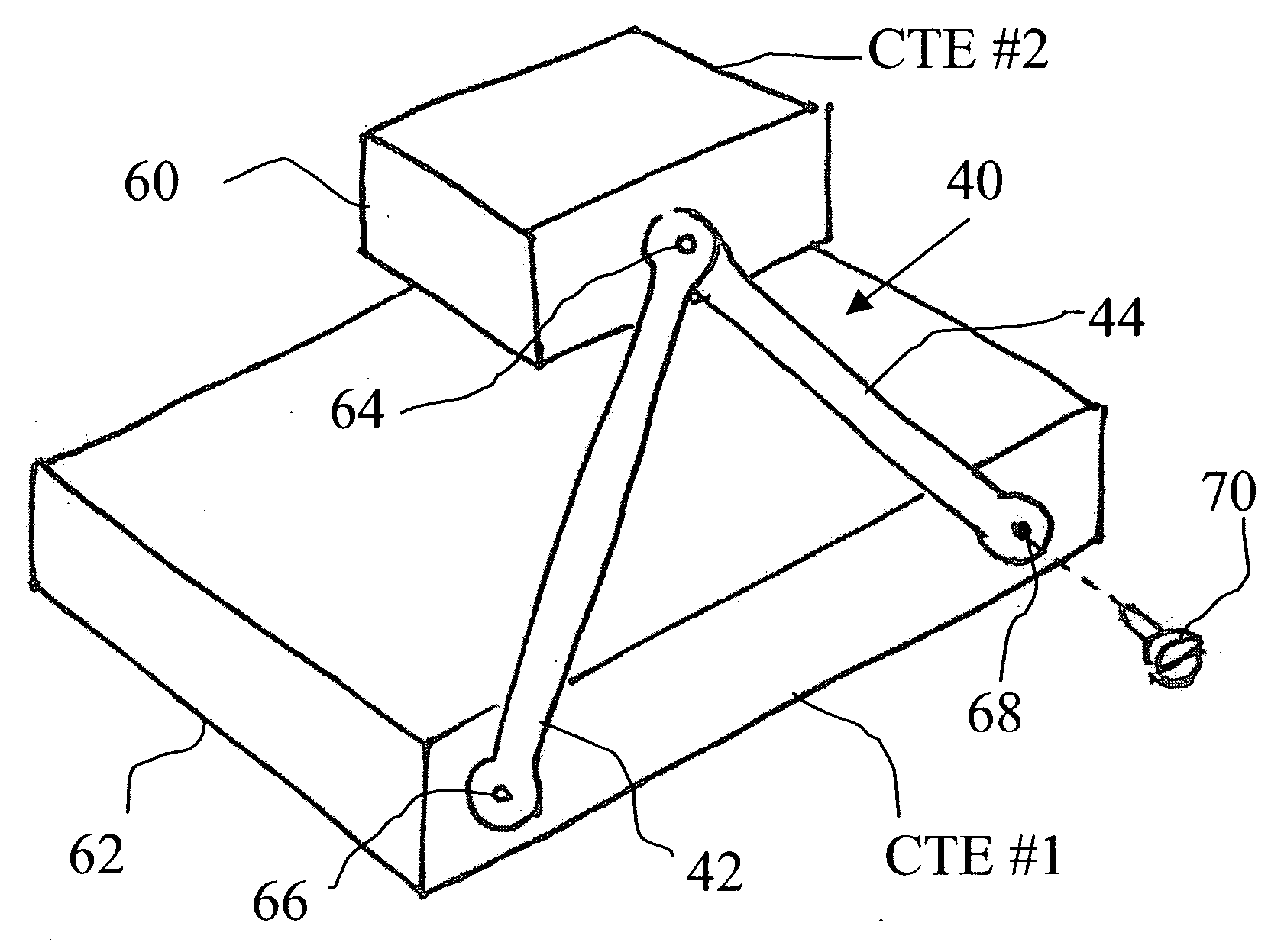
Compound coupling
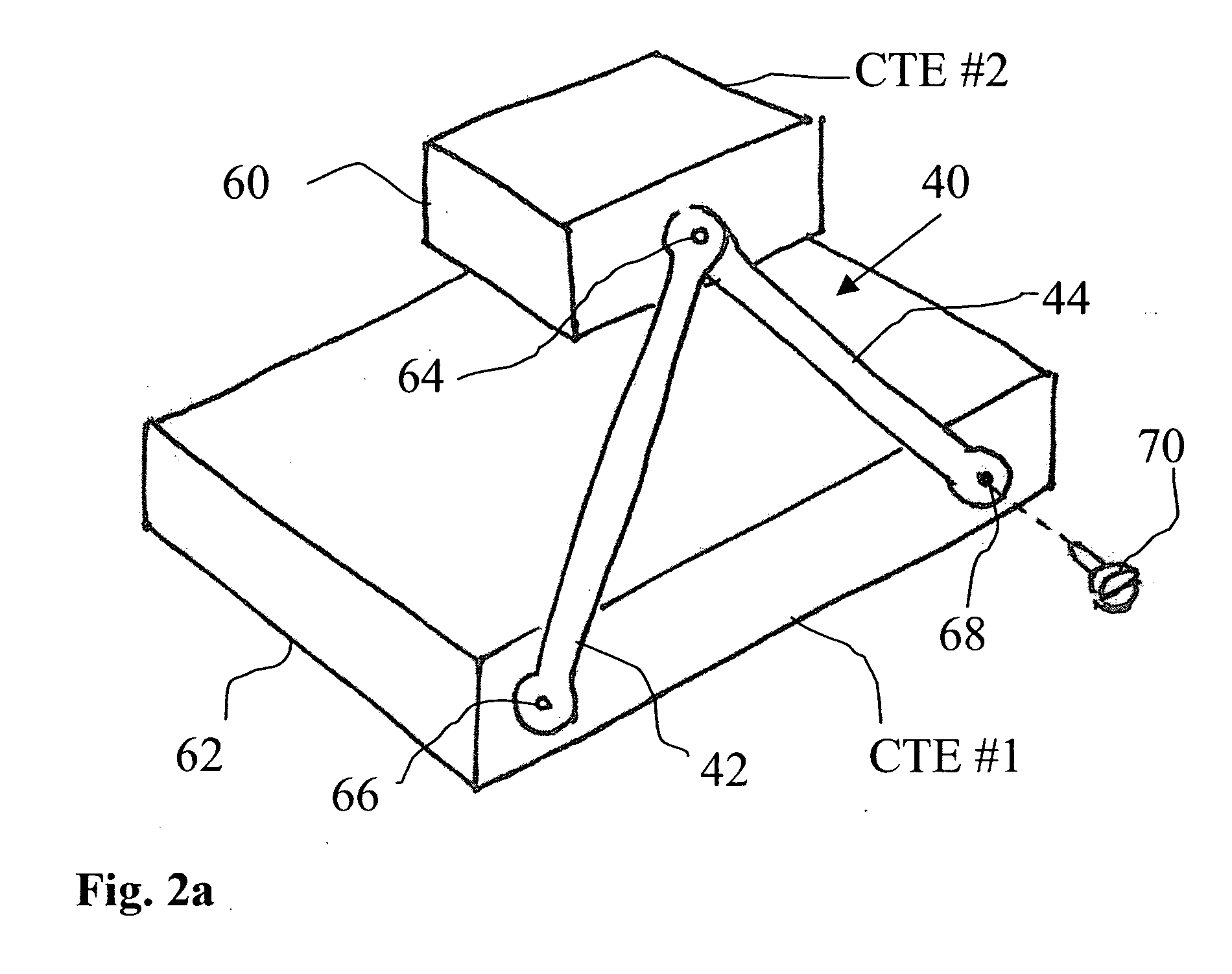
InactiveUS20050069376A1Minimal obstructionMechanically simple and robustRopes and cables for vehicles/pulleySecuring devicesEngineeringThermal expansion
A compound coupling for mounting a component (60) having a first coefficient of thermal expansion (CTE) to a base (62) having a second CTE, the compound coupling comprising a first flexure coupling (40), a second flexure coupling (40), and a third flexure coupling (40), each flexure coupling (40) extends from the base (62) to the component (60) and attached to the base (62) at a first and a second mount point (66, 68) associated with that flexure coupling (40). Each flexure coupling (40) is also attached to the component (60) at an associated component mount point (64). Each flexure coupling (40) has a flexure CTE substantially equal to the second CTE of base (62).
Owner:EASTMAN KODAK CO
Three-dimensional thin-film semiconductor substrate with through-holes and methods of manufacturing
InactiveUS8551866B2Eliminate and reduce disadvantageEliminate and reduce and problemFinal product manufactureSemiconductor/solid-state device manufacturingPorous semiconductorsThin film semiconductors
Owner:BEAMREACH SOLAR INC
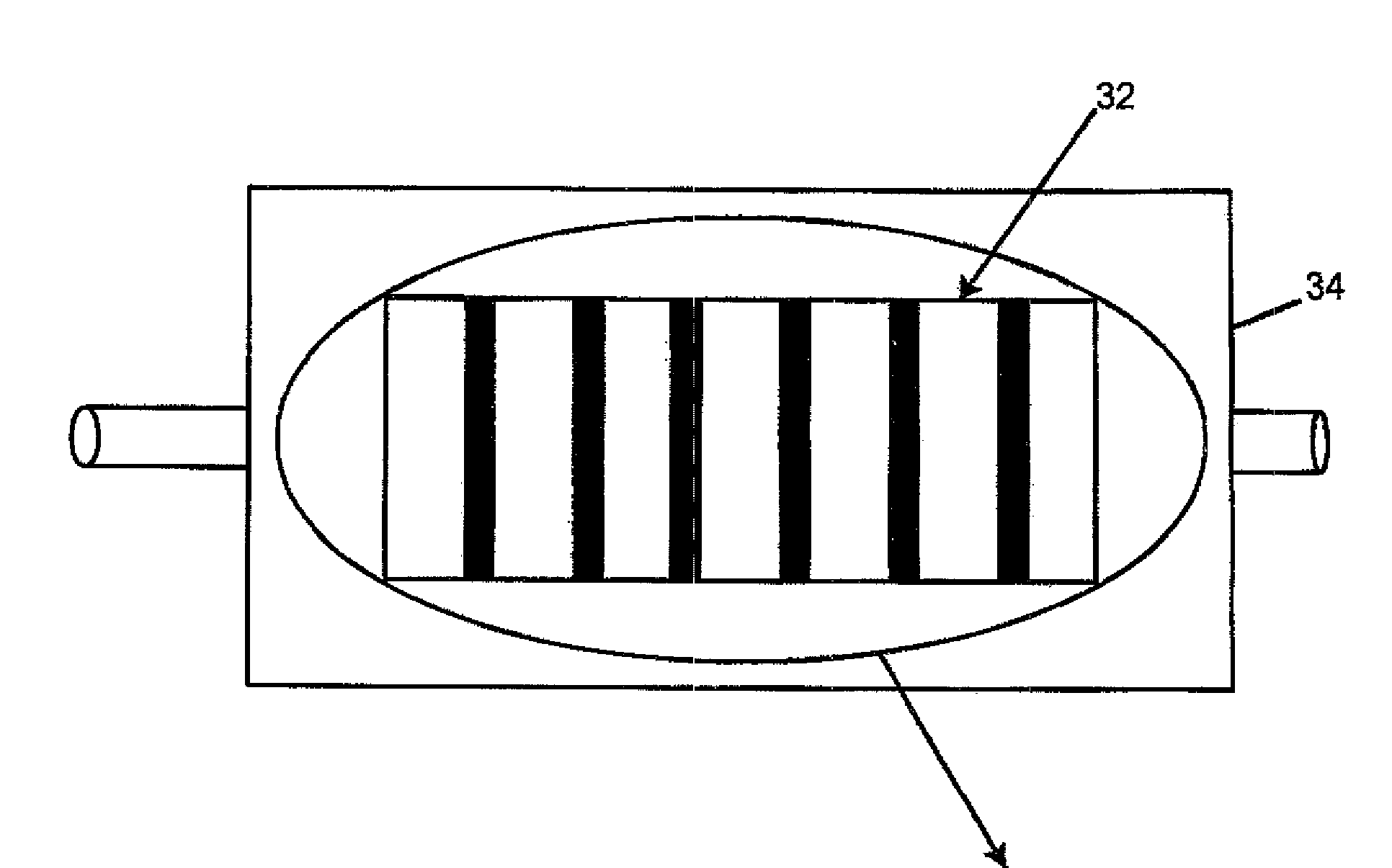
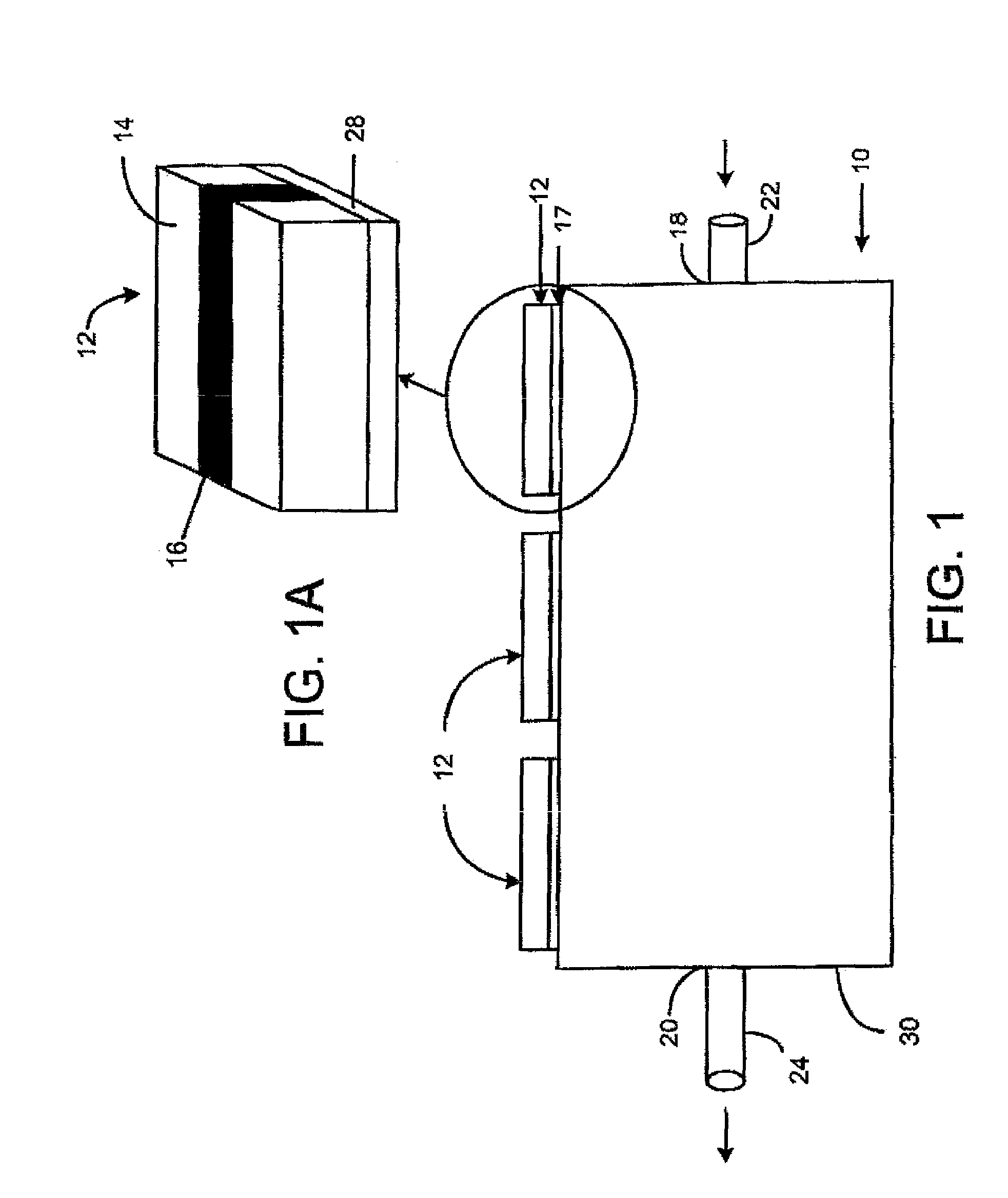
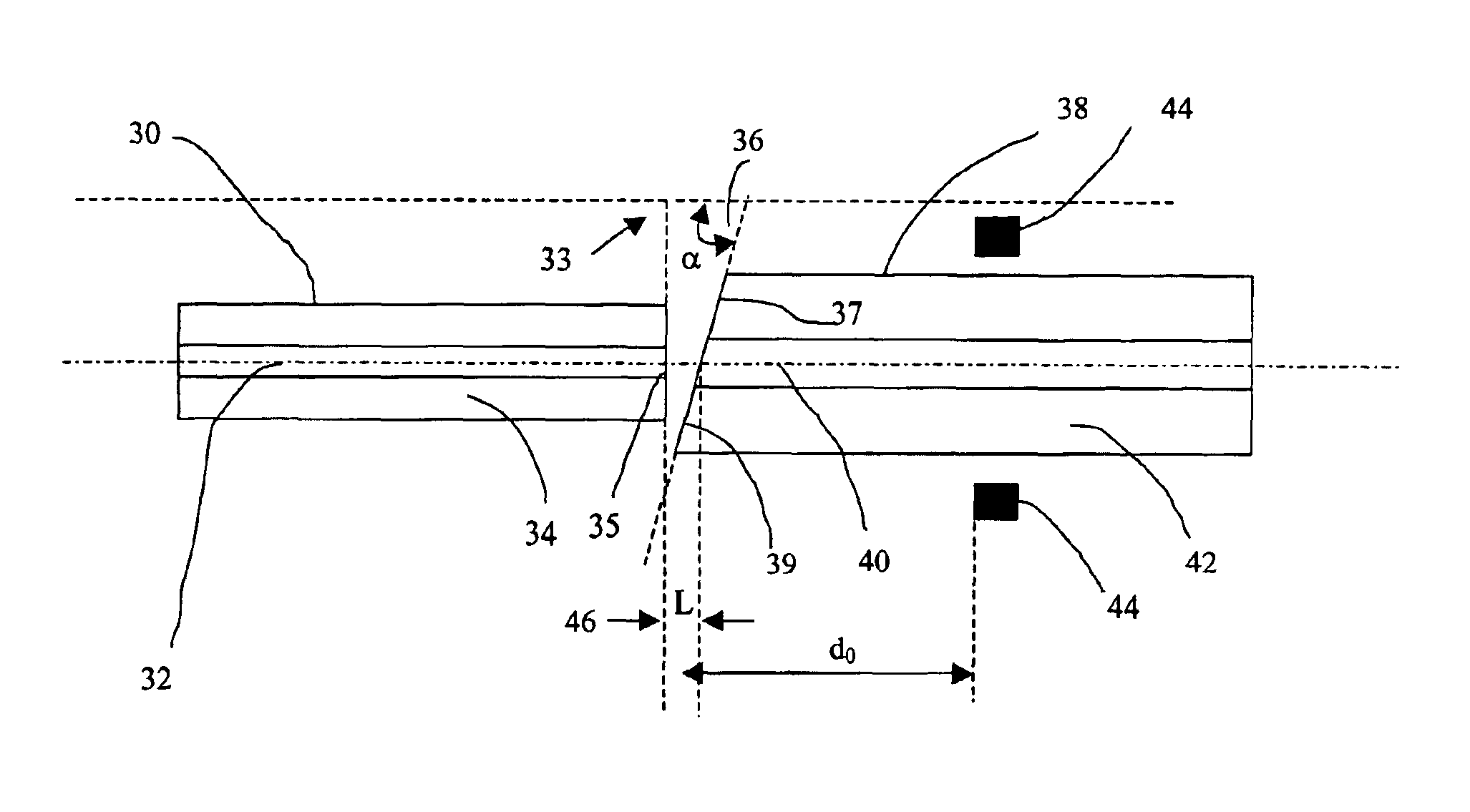
Method of angle fusion splicing silica fiber with low-temperature non-silica fiber
InactiveUS6866429B2Reduce back reflectionMechanically robustGlass making apparatusCoupling light guidesRange of motionSilica fiber
A low-cost approach is provided for forming a low splice loss, low back reflection loss and mechanically robust angle-fusion splice between a standard silica fiber and a low-temperature non-silica glass fiber. This is accomplished by angle cleaving the silica fiber, square cleaving the non-silica fiber and then asymmetrically heating the fibers to form an angle fusion splice. A matched angle at the end of the non-silica fiber is generated in situ during the splicing process. The tip of the angle-cleaved silica fiber may be polished flat back to the edge of the core to reduce the range of motion of the non-silica fiber during splicing thereby further reducing splice loss and enhancing the mechanical strength of the joint.
Owner:NP PHOTONICS A CORP OF DELAWARE
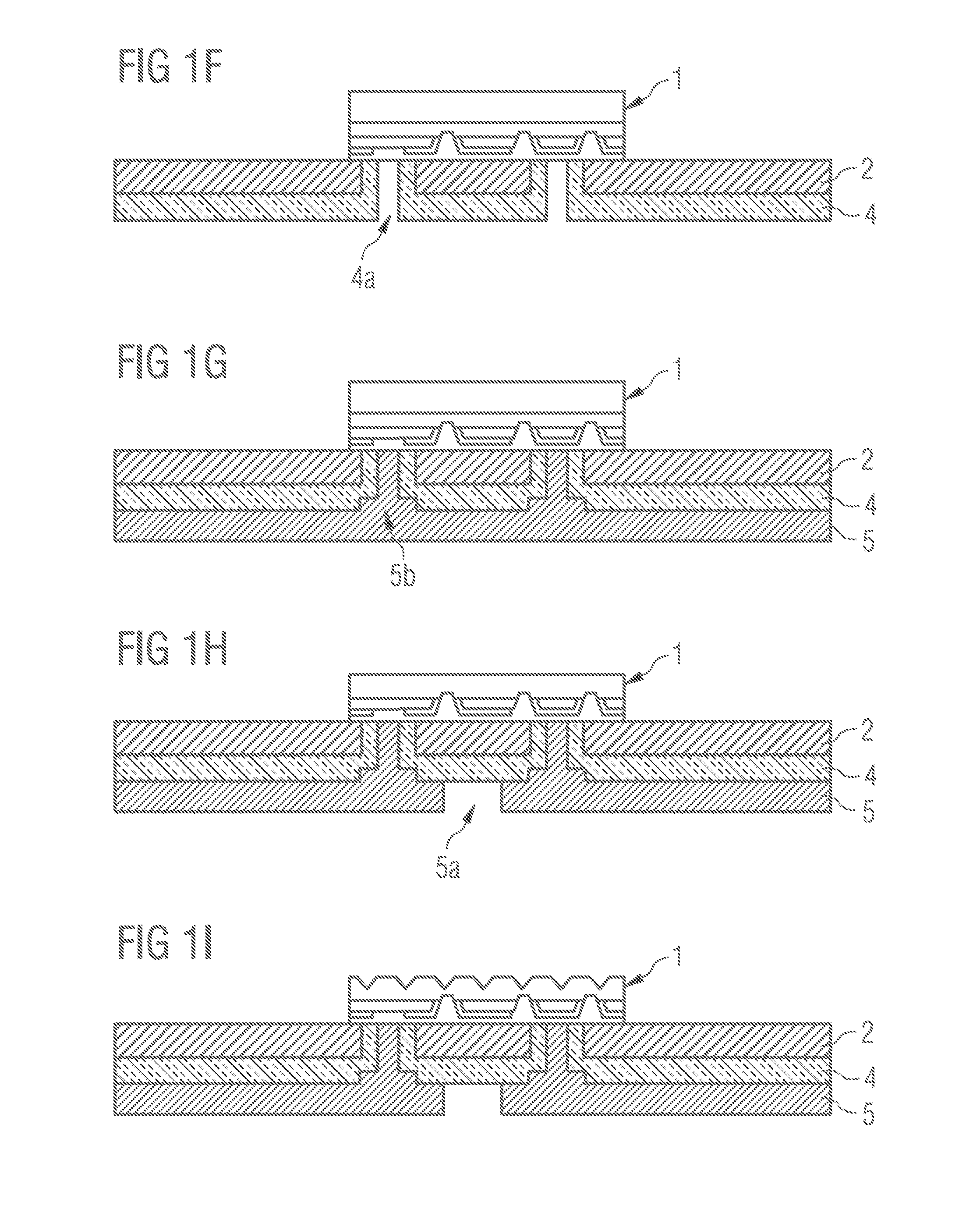
Method for Producing an Optoelectronic Semiconductor Component and Such a Semiconductor Component
InactiveUS20140151724A1Simplify chip adjustmentMechanically robustSolid-state devicesSemiconductor/solid-state device manufacturingElectrically conductiveEngineering
A method for producing a semiconductor component is disclosed. A carrier substrate includes a mounting region and an opening, which is formed in the mounting region of the carrier substrate. After mounting a semiconductor chip, an electrically insulating layer is applied to the carrier substrate in such a way that the electrically insulating layer completely fills the first opening in the carrier substrate. A second opening is formed in the electrically insulating layer. An electrically conductive layer is then applied to the electrically insulating layer in such a way that the second opening is filled with the electrically conductive layer in the form of a via. A semiconductor component produced in this way is also provided.
Owner:OSRAM OPTO SEMICON GMBH & CO OHG
Boat that can be converted into a gymnastics apparatus
ActiveUS8986057B2Low production costEasy to buySteering ruddersMuscle power acting propulsive elementsDriver/operatorEngineering
The present invention relates to a boat propelled by two submerged independent wings actuated by means of a step simulator mechanism that imparts an vertical alternating movement to the wings, which are articulated around a shaft located forward of the center of pressure thereof, such that the wings assume angles of inclination that generates propulsive force in the upward and downward movements. In the analogous position of a conventional bicycle, the driver actuates the footrests while seated. A handlebar, connected to a rudder, is used to maneuver the vehicle. The catamaran configuration offers hydrodynamic efficiency and stability. When disassembled, the mainframe of the boat can be converted into a step machine, simply fitting a pair of dampers to the structure, which absorb and dissipate the energy generated during exercising. Thus, the present invention has a dual use—boat and gymnastics apparatus—and can be transported in the car trunk.
Owner:CATARINA MATEUS FROIS SANTA
Features
- R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
Why Patsnap Eureka
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Social media
Patsnap Eureka Blog
Learn More Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com