Patents
Literature
Hiro is an intelligent assistant for R&D personnel, combined with Patent DNA, to facilitate innovative research.
321 results about "Full length cdna" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Full-length plant cDNA and uses thereof
Owner:NAT INST OF AGROBIOLOGICAL SCI +1
Acid sphingomyelinase protein and methods of treating type B Niemann-Pick disease
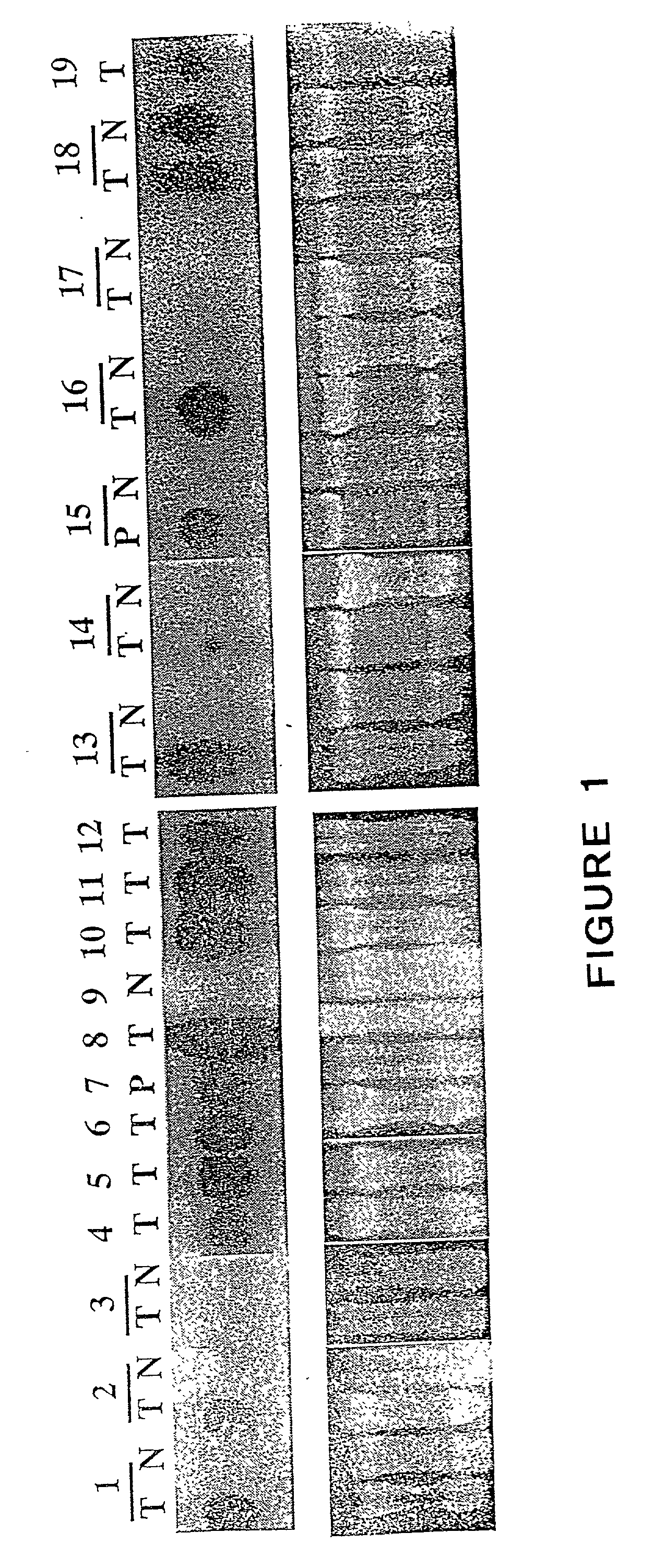
The present invention relates to the acid sphingomyelinase gene and to methods of diagnosing Niemann-Pick disease. It is based, at least in part, on the cloning and expression of the full-length cDNA encoding acid sphingomyelinase and on the discovery of mutations in the acid sphingomyelinase gene of Ashkenazi Jewish Niemann-Pick disease patients.
Owner:MT SINAI SCHOOL OF MEDICINE
Tumor antigen useful in diagnosis and therapy of prostate and colon cancer
InactiveUS7037667B1Microbiological testing/measurementBiological material analysisCell membraneTumor antigen
Owner:AGENSYS
Methods and compositions for diagnosis and treatment of breast cancer
InactiveUS6399328B1Control metastatic potentialOrganic active ingredientsTumor rejection antigen precursorsAbnormal tissue growthLymphatic Spread
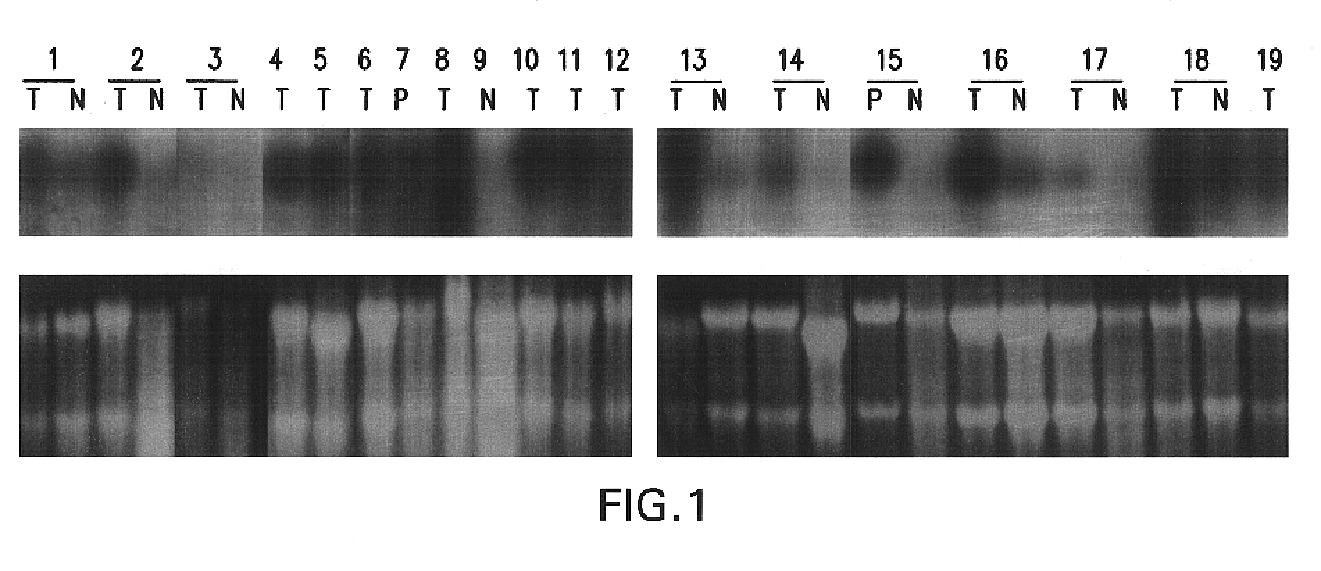
The present invention relates to a novel gene, Di12, that is differentially expressed as a 1.35 kb RNA in breast cancer tissues and cell lines, and in several normal tissues. The full length cDNA encodes a protein of 339 amino acids. Antibodies to the gene product were developed to investigate the expression of Di12 in breast cancer cell-lines and tumors. The Di12 protein was found in tissue sections of infiltrating ductal carcinomas (IDCs), but not in benign or normal breast specimens. Di12 wag also present in IDC-breast cancer patient sera, and its expression level increased markedly if IDC was accompanied by lymph node or distal metastases. As IDC constitutes ~70% of breast cancers seen clinically, the level of Di12 expression is useful for diseases diagnosis predicting disease progression and monitoring a therapeutic treatment.
Owner:MUSC FOUND FOR RES DEV
Method of synthesizing cdna
ActiveUS20060246453A1High yieldIncrease chanceSugar derivativesMicrobiological testing/measurementHeteroduplexReverse transcriptase
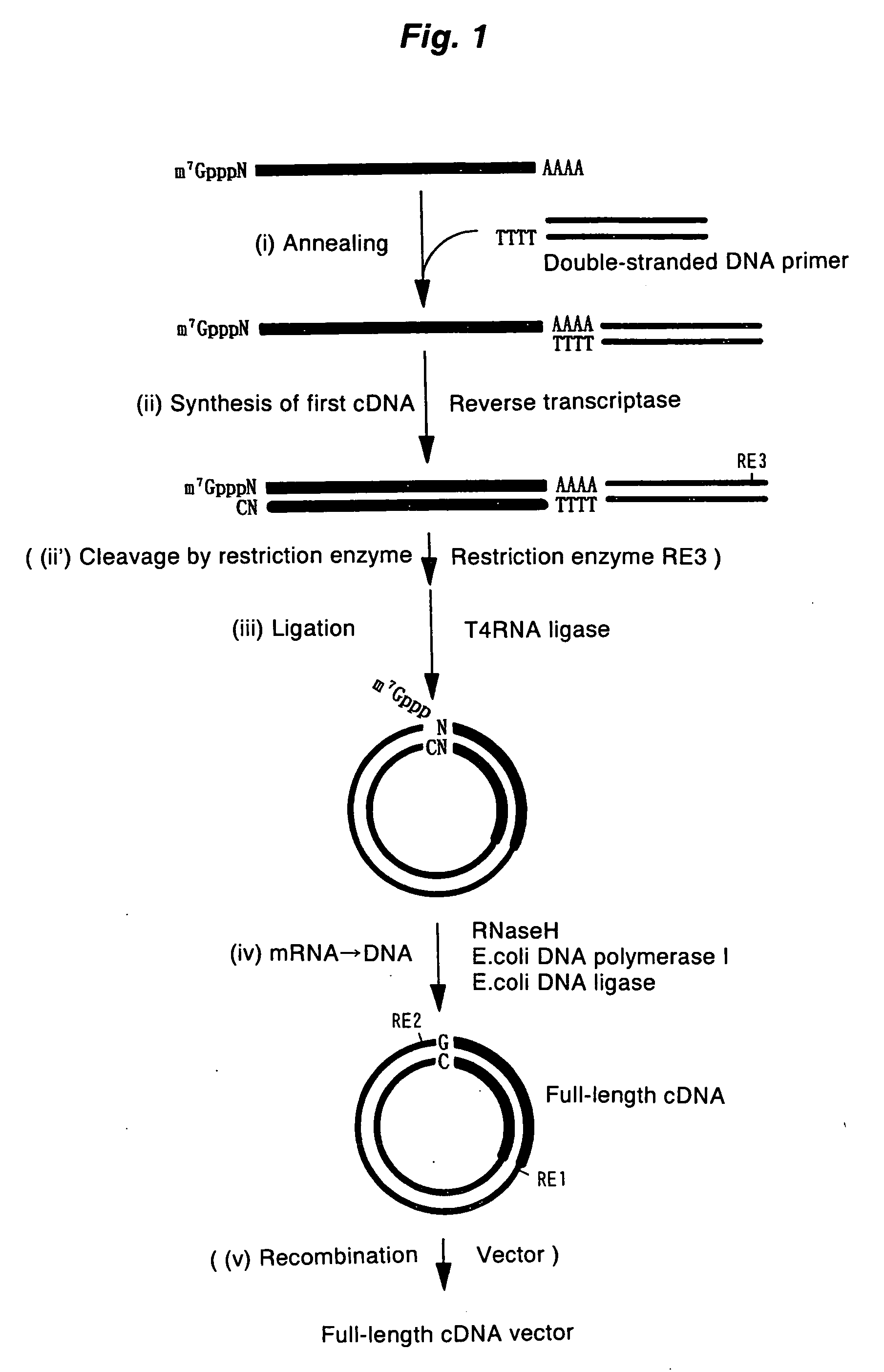
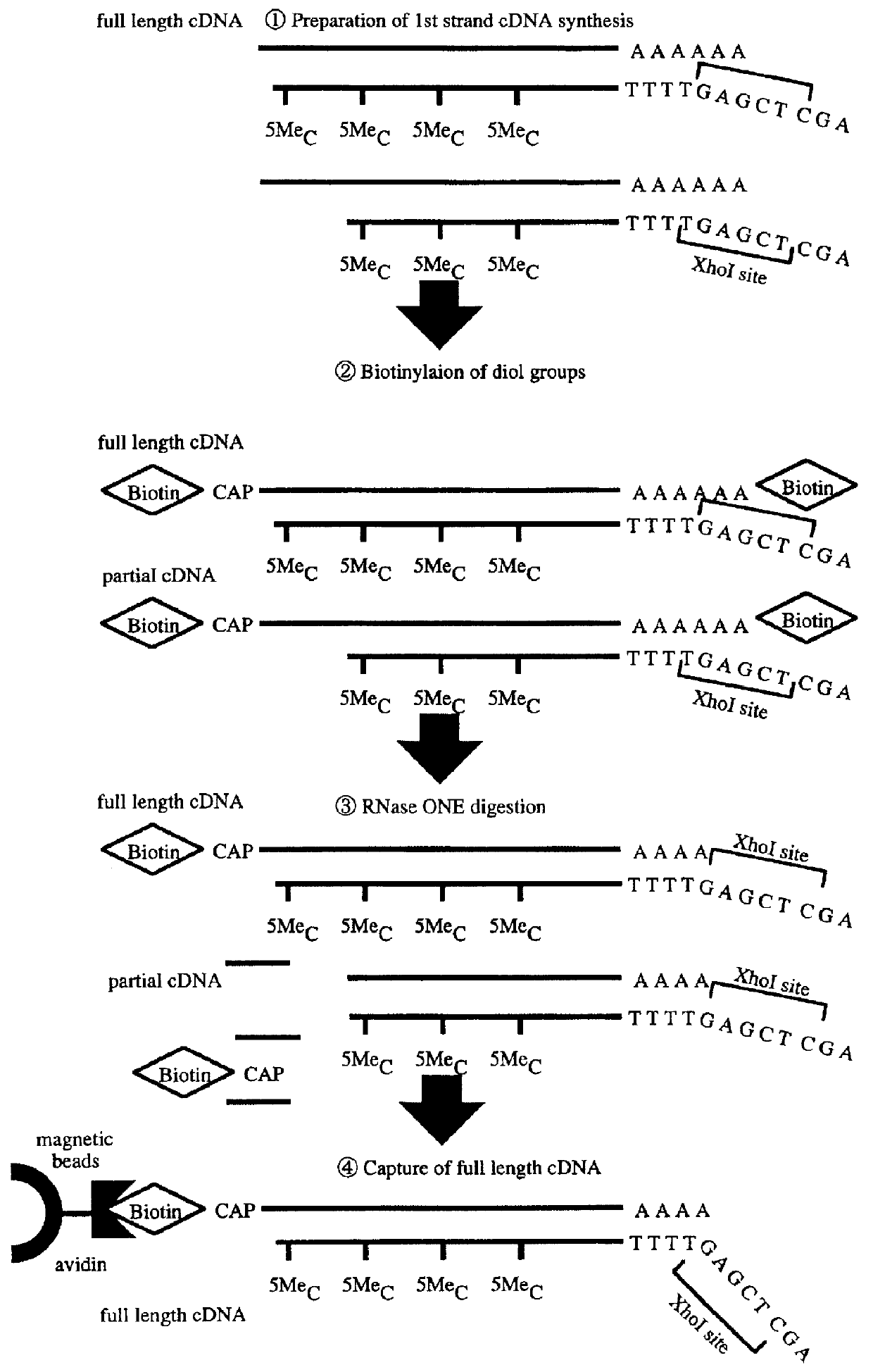
A method for synthesizing cDNA possessing a consecutive sequence starting with a nucleotide adjacent to a cap structure of mRNA, which comprises (i) a process for annealing a double-stranded DNA primer and an RNA mixture containing mRNA possessing a cap structure, (ii) a process for preparing a conjugate of an mRNA / cDNA heteroduplex and a double-stranded DNA primer by synthesizing the first-strand cDNA primed with the double-stranded DNA primer using reverse transcriptase, and (iii) a process for circularizing the conjugate of the mRNA / cDNA heteroduplex and the double-stranded DNA primer by joining the 3' and 5' ends of the DNA strand containing cDNA using ligase. This method enables us to efficiently synthesize a full-length cDNA possessing a consecutive sequence starting with a transcription-start-site nucleotide from a small amount of RNA by small processes.
Owner:KOKURITSU SHINTAI SHIYOUGAISHI +2
Method for making full-length cDNA libraries
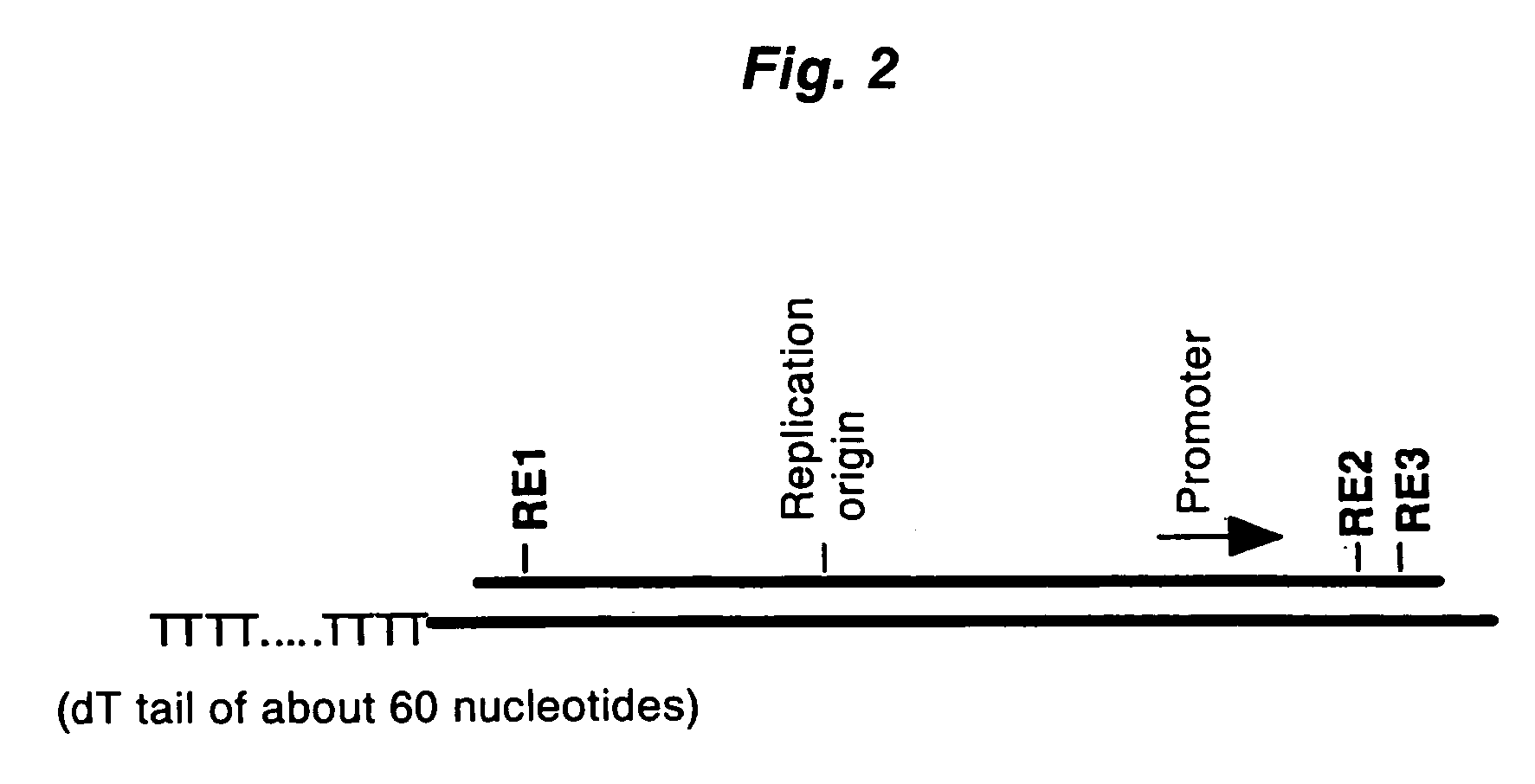
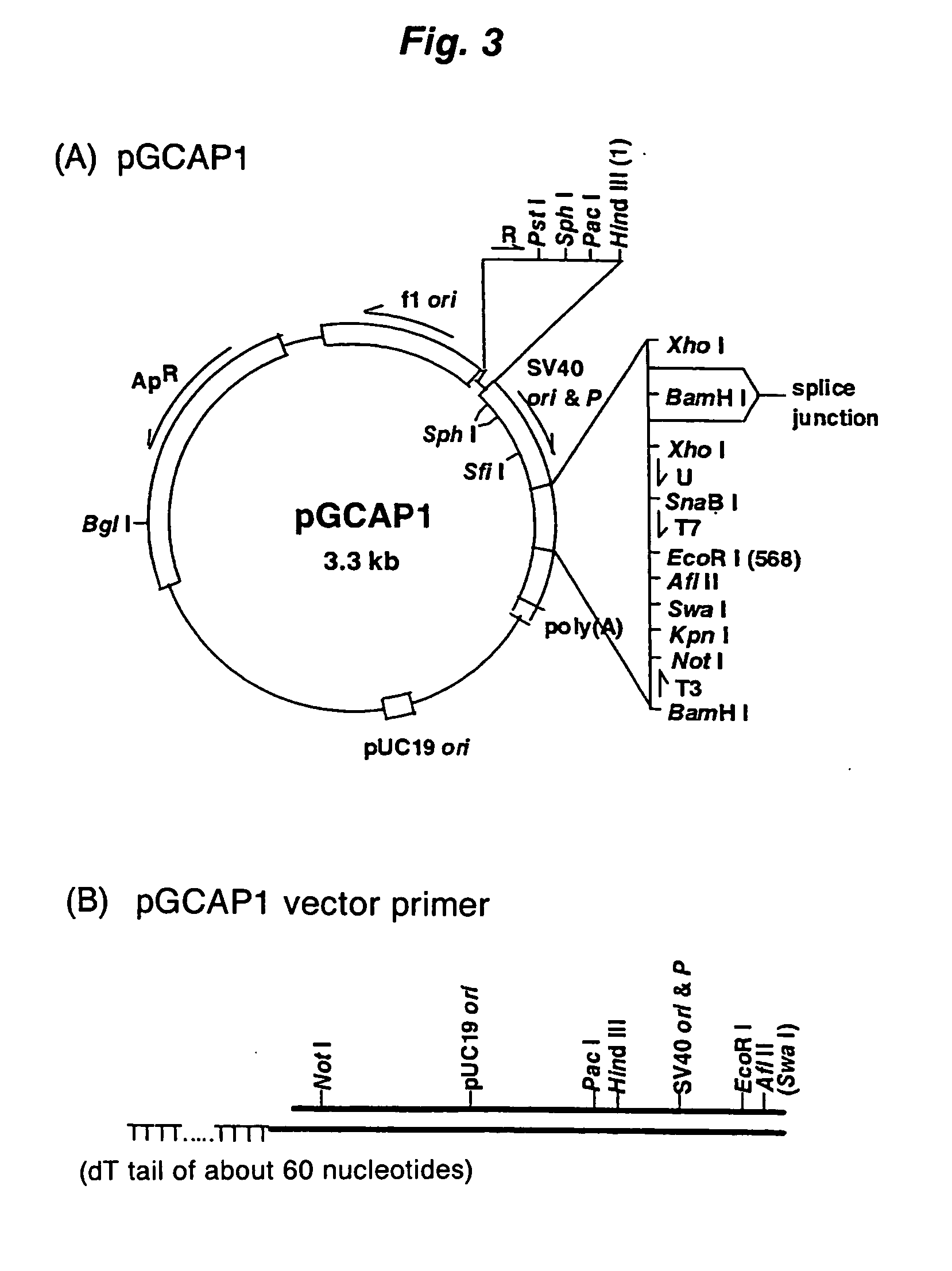
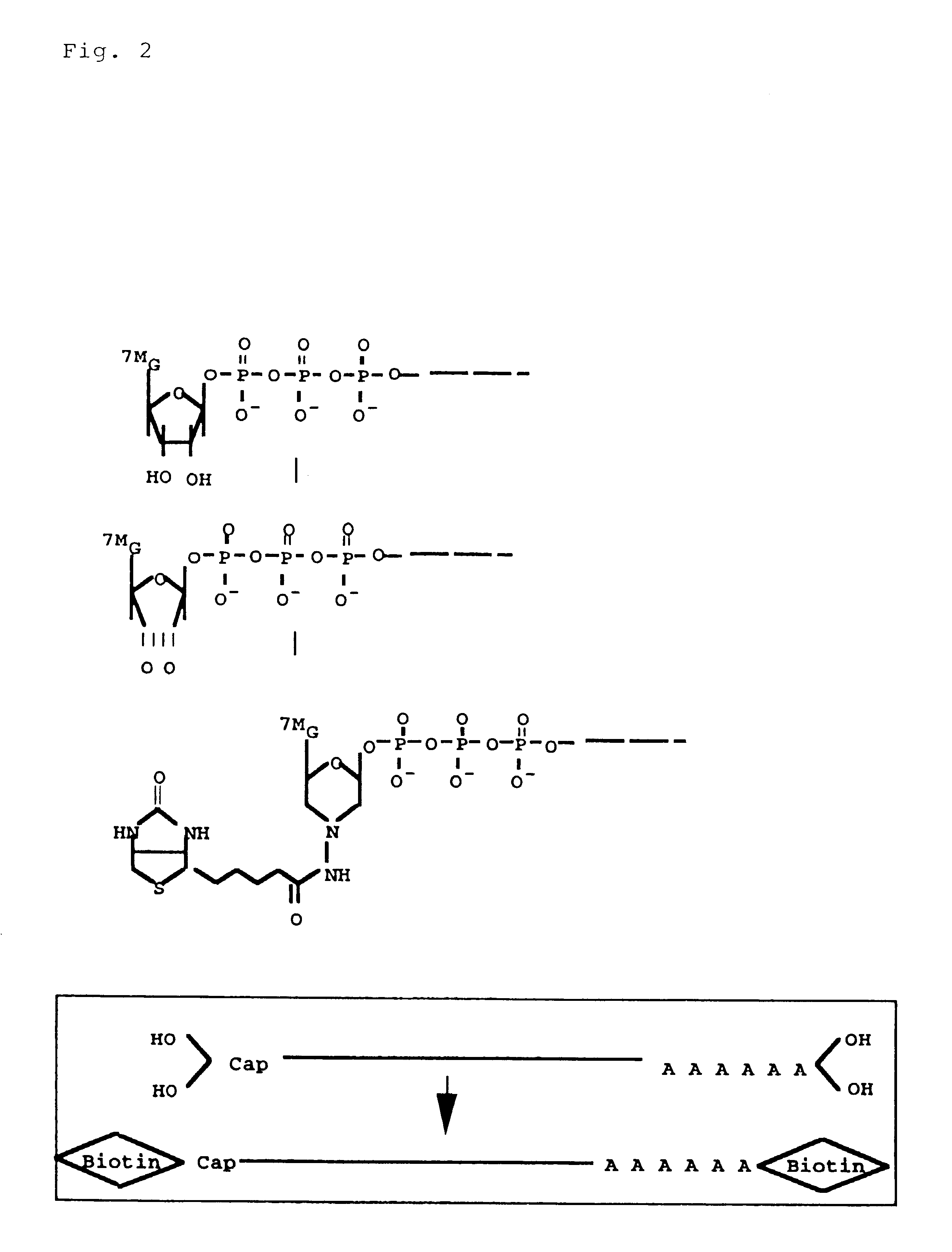
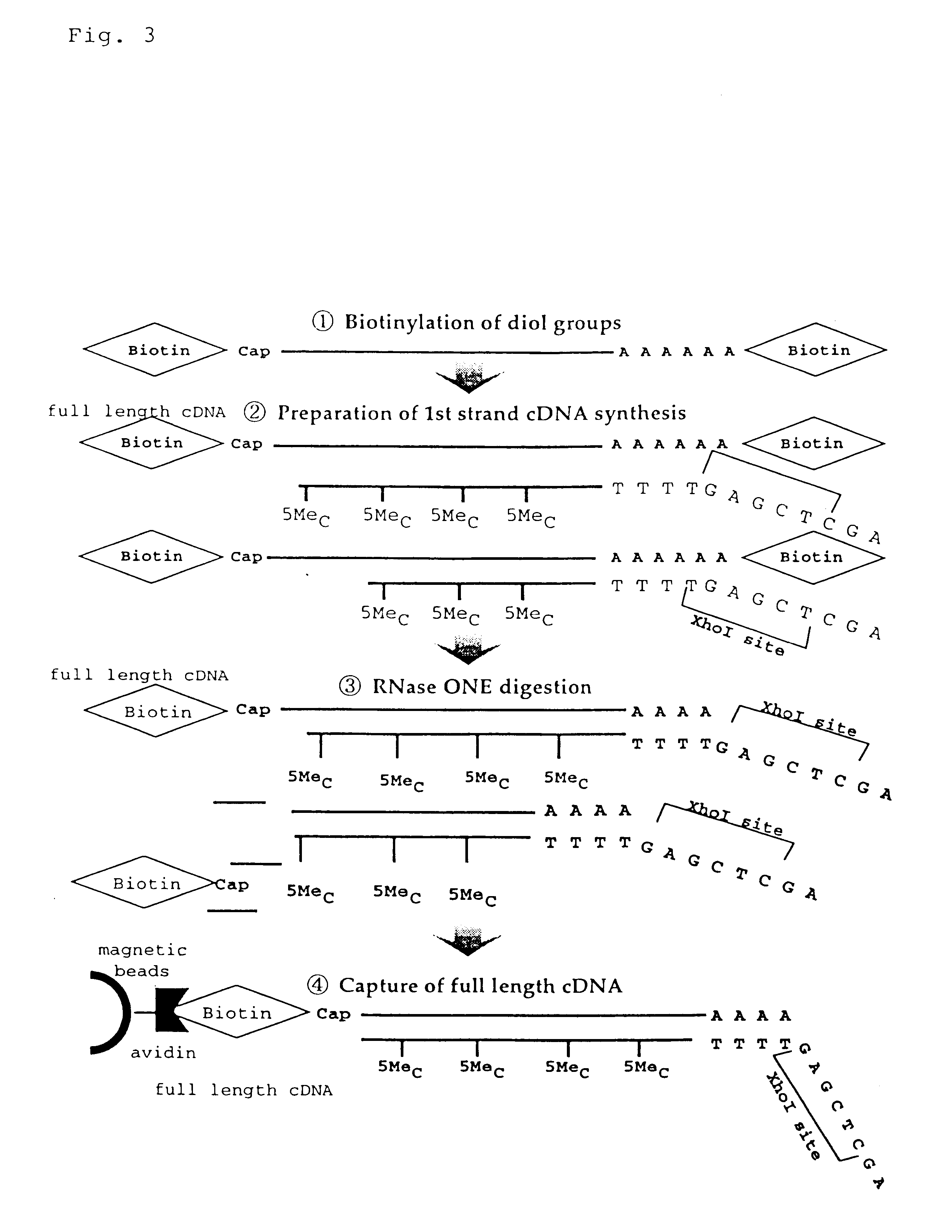
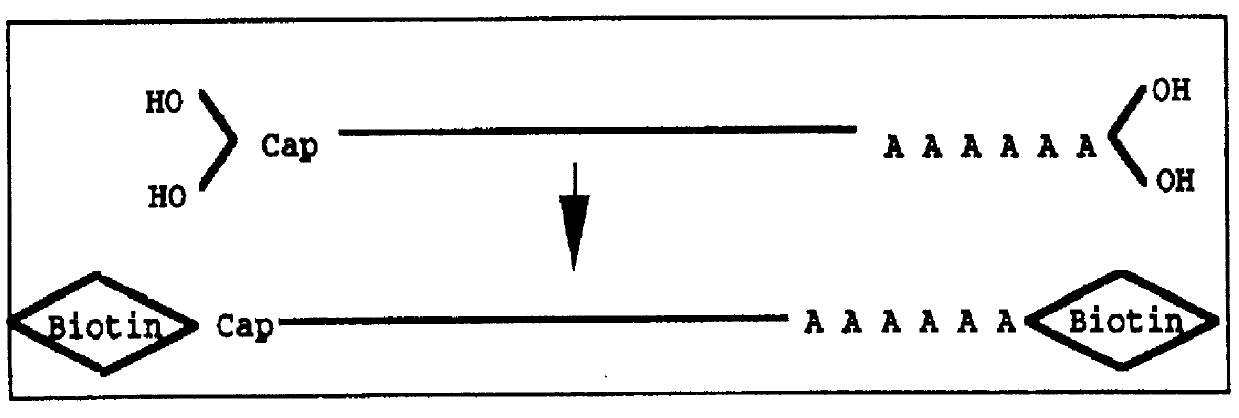
Disclosed is a method for making full-length CDNA libraries, which is for making libraries of cDNAs having a length corresponding to a full-length of mRNAs and comprises the following steps of; binding a tag molecule to a diol structure present in 5' Cap (7MeGpppN) sites of mRNAs, forming RNA-DNA hybrids by reverse transcription using primers such as oligo dT and the mRNAs connected with the tag molecule as templates, and separating RNA-DNA hybrids carrying a DNA corresponding to a full-length of mRNAs from the RNA-DNA hybrids formed above by using function of the tag molecule. To obtain mRNA connected with a tag molecule, the diol structure present in 5' Cap site of mRNA is subjected to a ring-open reaction by oxidation with sodium periodate to form a dialdehyde and the dialdehyde is reacted with a tag molecule having a hydrazine terminus. According to the present invention, there are provided a novel method capable of efficiently labeling 5' Cap site and a method for making full-length cDNA libraries utilizing the labeling method.
Owner:THE INST OF PHYSICAL & CHEM RES WAKO
Methods and compositions for diagnosis and treatment of cancer
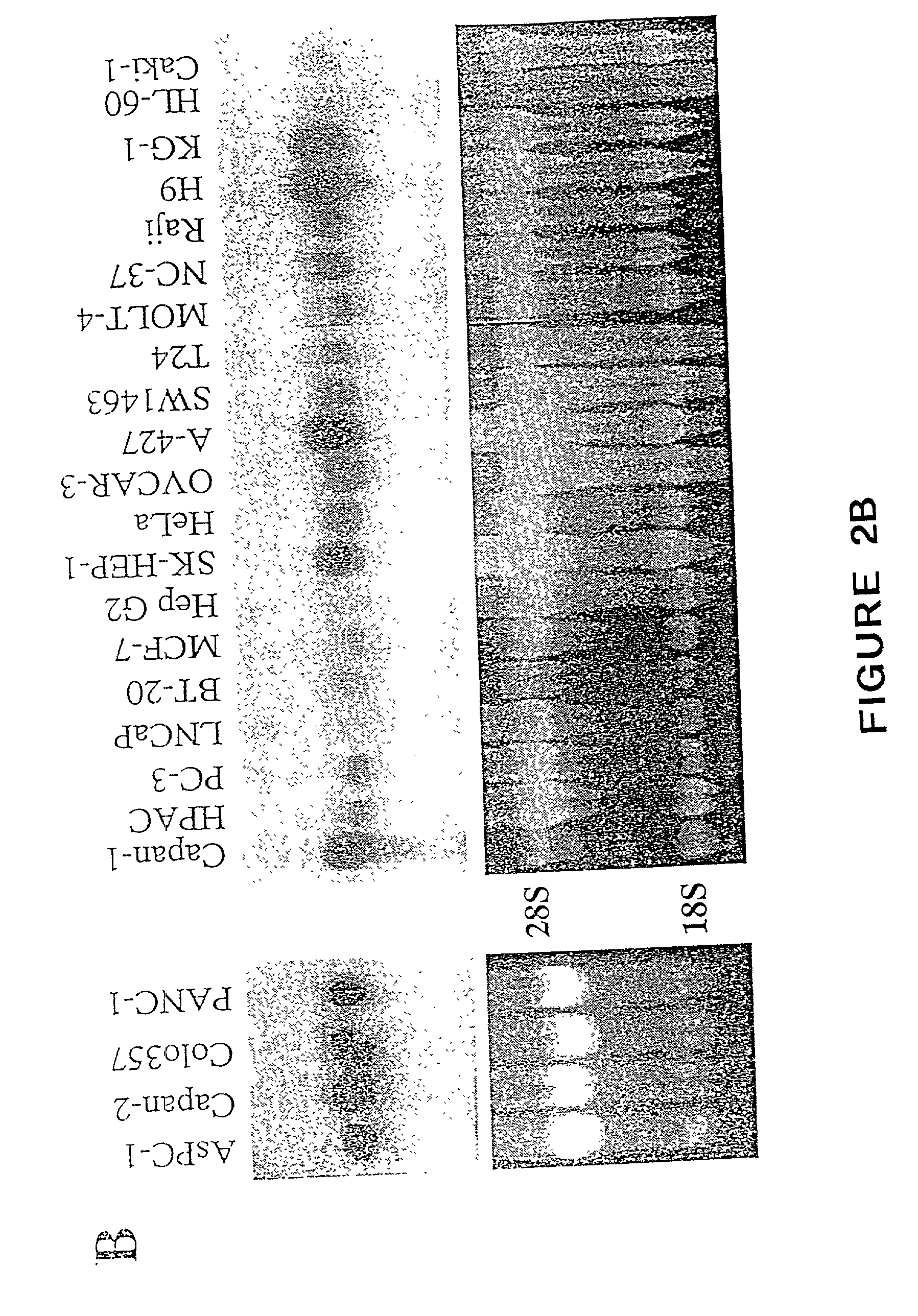
The present invention relates to a novel gene, CaSm, that is highly expressed in cancer tissues and cell lines, especially pancreatic cancer. The full length cDNA of CaSm encodes a protein of 133 amino acids. CaSm contains the two Sm motifs found in the common snRNP proteins, with the greatest homology to the Sm G protein (60% similarity). The present invention further encompasses CaSm peptides, fusion proteins, host cell expression systems, antibodies to CaSm, antisense CaSM molecules, and compounds that modulate CaSm gene expression or CaSm activity. Antisense CaSm RNA is able to alter the transformed phenotype of pancreatic cancer cells by reducing their ability to form large colonies in soft agar when compared to untransfected cells. The present invention also encompasses methods for disease diagnosis, drug screening and the treatment of cancer.
Owner:MUSC FOUND FOR RES DEV
Method of preparing normalized and/or subtracted cDNA
InactiveUS20020106666A1Avoid disadvantagesImproving and subtractionSugar derivativesMicrobiological testing/measurementSingle-Stranded RNAFull length cdna
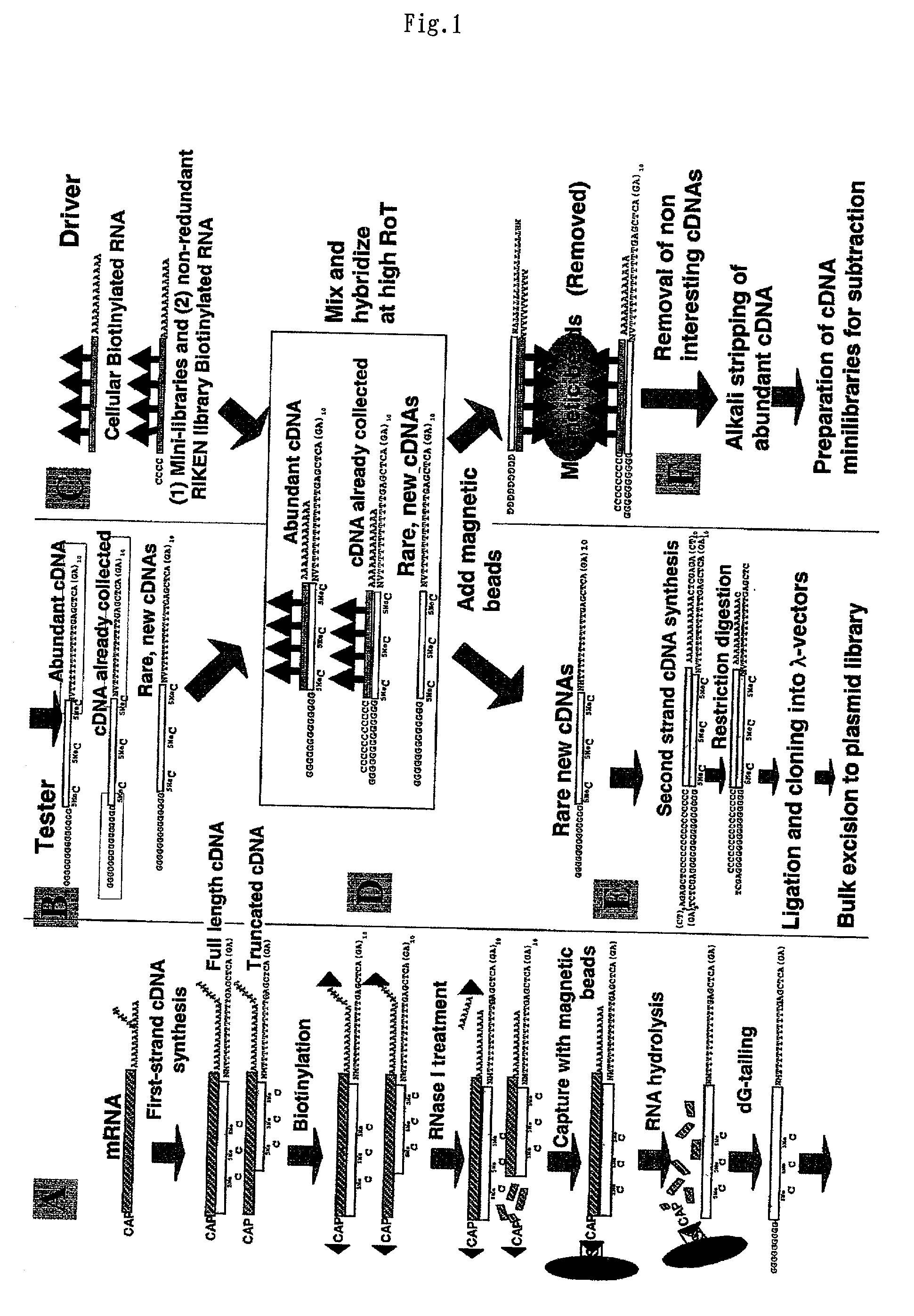
A method of preparing normalized and / or subtracted cDNA; a method in which the cDNA that is normalized and / or subtracted is in the form of uncloned cDNA (cDNA tester); a method of preparing normalized and / or subtracted cDNA comprising the steps of: (a) preparing cDNA tester; (b) preparing normalization and / or subtraction RNA driver; (c) conducting normalization and / or subtraction in two steps in any order, or conducting normalization / subtraction as a single step and mixing the normalization / subtraction RNA driver with said cDNA tester; (d) adding an enzyme capable of cleaving single strand sites on RNA drivers nonspecifically bound to cDNA tester; (e) removing said single strand RNA driver cleaved in step d) from the tester and removing tester / driver hybrids; and (f) recovering the normalized and / or subtracted cDNA; and a method of efficiently preparing normalized and / or subtracted long-chain, full-coding, and full-length cDNA libraries are provided.
Owner:YOSHIHIDE HAYASHIZAKI +2
Porcine DC-SIGN, ICAM-3 and LSECtin and uses thereof
The present invention relates to the cloning, identification and characterization of the unique and entire genomic sequences encoding new porcine DC-SIGN and LSECtin proteins, including the novel nucleotide sequences of the full-length cDNA and genes of both pDC-SIGN gene and pLSECtin. Also provided are the nucleic acid molecules encoding newly discovered porcine ICAM-3 isoforms from porcine monocyte-derived dendritic cells and the use thereof. Specifically, the invention is drawn to an isolated nucleic acid molecule comprising a nucleotide sequence encoding one or more of porcine DC-SIGN, porcine ICAM-3, porcine LSECtin, a complement of the nucleotide sequence or a functional, defined portion of the nucleotide sequence or a protein fusion product linked to a protein that may be of porcine or human origin. Methods for isolating and cloning the new porcine genes and for using the new nucleotide sequences in improved methods for propagating viruses, particularly enveloped viruses, are additionally described herein. The invention further includes new transfected cells or cell lines that can stably express the porcine proteins, new antibodies and the like.
Owner:VIRGINIA TECH INTPROP INC
Plant flavonoid synthesis regulation gene and its application
Owner:WUHAN BOTANICAL GARDEN CHINESE ACAD OF SCI
Method for forming full-length cDNA libraries
InactiveUS6143528AEasy to prepareEfficiently labeledSugar derivativesMicrobiological testing/measurementPresent methodFull length cdna
PCT No. PCT / JP97 / 03992 Sec. 371 Date Jul. 16, 1999 Sec. 102(e) Date Jul. 16, 1999 PCT Filed Oct. 31, 1997 PCT Pub. No. WO98 / 20122 PCT Pub. Date May 14, 1998Disclose is a method for making full-length cDNA libraries, which is for making libraries of cDNAs having lengths corresponding to full lengths of mRNAs and comprises the following steps of; forming RNA-DNA hybrids by reverse transcription starting from primers using mRNAs as templates, chemically binding a tag molecule to a diol structure present in the 5' Cap (7MeGpppN) site of a mRNA which is forming a RNA-DNA hybrid, and separating RNA-DNA hybrids carrying a DNA corresponding to a full-length mRNA from the RNA-DNA hybrids formed above by using a function of the tag molecule. The present method is a method for preparing full-length cDNA libraries utilizing a method for labeling the 5' Cap site more efficiently than protein enzyme reactions, which is avoidable a decrease of a full-length cDNA synthesis efficiency caused by cleavage of mRNA, and can synthesize a full-length cDNA more efficiently.
Owner:RIKEN
Methods and compositions for diagnosis and treatment of cancer
InactiveUS20020086812A1Reducing tumorigenicityReduce dosageOrganic active ingredientsBiocideDiseaseNovel gene
The present invention relates to a novel gene, CaSm, that is highly expressed in cancer tissues and cell lines, especially pancreatic cancer. The full length cDNA of CaSm encodes a protein of 133 amino acids. The present invention further encompasses CaSm peptides, fusion proteins, host cell expression systems, antibodies to CaSm, antisense CaSm molecules, and compounds that modulate CaSm gene expression or CaSm activity. The present invention also encompasses methods for disease diagnosis, drug screening and the treatment of cancer. In particular, the combined use of a CaSm antagonist with a therapeutic agent to treat cancer is encompassed.
Owner:THE MEDICAL UNIV OF SOUTH CAROLINA
Method for extensively screening scallop SNP
InactiveCN101845489AThe principle is safe and reliableReliable Large-Scale Screening RunsMicrobiological testing/measurementMolecular geneticsPhenol
The invention belongs to a method for extensively screening scallop SNP in the technical field of the scallop DNA molecular genetic marker, which comprises the following steps that: solution D and phenol-chloroform are firstly used for extracting a plurality of Chlamys farreri RNA, and a ultraviolet specrophotometer is used for measuring the concentration of the mixture of solution D and the phenol-chloroform which are uniformly mixed; a scallop full-length cDNA sample is re-established and comprises the synthesis of a cDNA first chain and the synthesis and augmentation of a cDNA second chain; then homogenization of full-length cDNA is performed, and ultrasonic breaking is performed; then sequence measuring joints are connected, and terminal repair, joint connection and sample augmentation are included; finally a biological software is used for analyze the data to obtain an SNP marker; then an SNP marker to be screened is selected to design a primer; and DNA of individual scallop is extracted to be equivalently mixed to be used as a template, conventional colony sequence measuring is undertaken after the PCR augmentation, and the position point with the occurring frequency of single base difference being 10 percent or more is defined as an SNP marker. The method has safe and reliable principle, simple and controllable flow procedures, reliable running of large-scale screening, excellent effect and strong practicability.
Owner:OCEAN UNIV OF CHINA
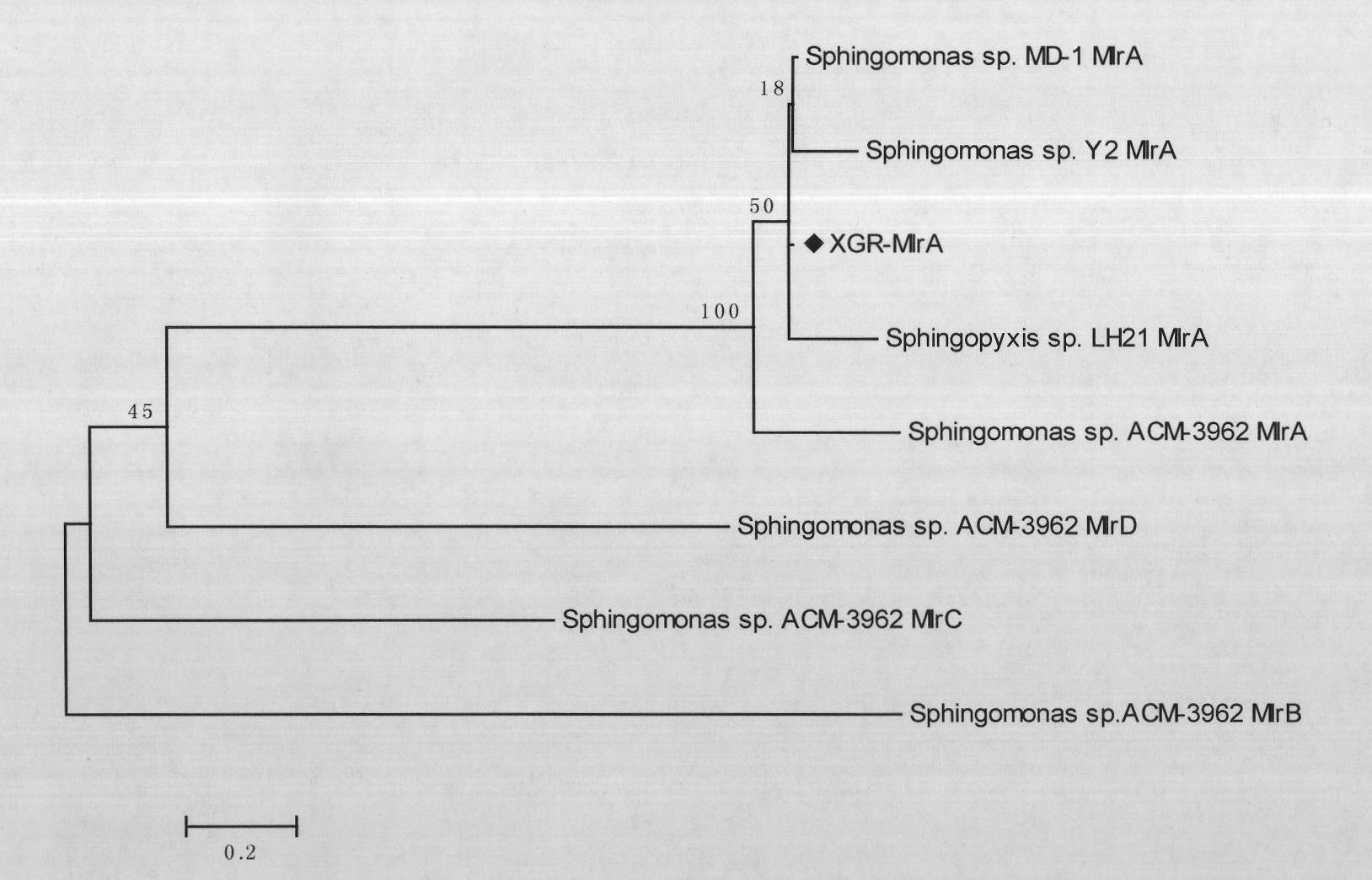
Overall length cDNA sequence of micro-capsule algae toxins degrading enzyme MlrA, coded amino acid and application
InactiveCN101775403AAvoid damageImprove food safetyBacteriaAnimal feeding stuffPhylum CyanobacteriaCyanophages
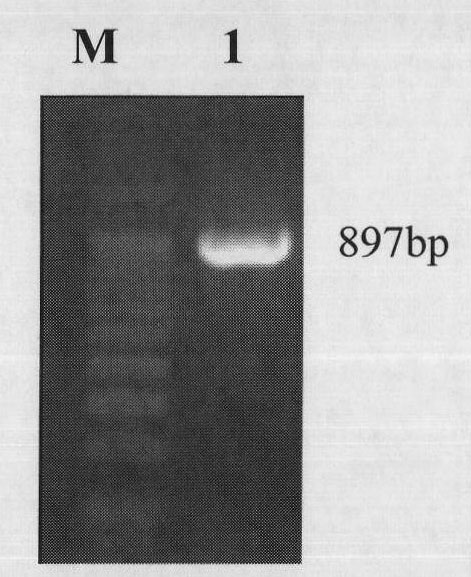
The invention provides an overall length cDNA sequence of micro-capsule algae toxins degrading enzyme MlrA, a coded protein and application. The nucleotide sequence of the overall length cDNA sequence of micro-capsule algae toxins degrading enzyme MlrA is shown as SEQ IDNO:1, and the coded amino acid sequence s shown as SEQ ID NO:3. The overall length cDNA sequence of micro-capsule algae toxins degrading enzyme MlrA in the invention has uniqueness and specifity for degradation of algae toxins, and can be used for detecting the situation of micro-capsule algae toxins in water body or fish products. The micro-capsule algae toxins degrading enzyme MlrA can be used as crude enzyme preparation in optimizing aquaculture water to effectively prevent and control damage of algae toxins on eating safety of bred aquatic products, and also can be used as fish bait additive to mitigate damage of micro-capsule algae toxins on fish hepatic cell DNA, maintain detoxication capacity of hepatic cells on algae toxins during blue-green algae bloom outbreak, and greatly improve eating safety of bred fish.
Owner:JINAN UNIVERSITY
Functional mutant of human plasminogen, its preparation method and application
InactiveCN102199587AGood for renaturationExtended half-lifeSenses disorderNervous disorderPichia pastorisEukaryotic plasmids
A functional mutant of human plasminogen disclosed in the invention, respectively, is hPLG-delta K: human plasminogen protein Pro<544>-Asn<791> polypeptide; Pro<559> in RGD-hPLG-delta K: hPLG-delta K is mutated to Asp<559>; and Gly<560> in RHP-hPLG-delta K: hPLG-delta K polypeptide is mutated to His<560>. The invention also discloses a preparation method for the functional mutant. The product is obtained by using a plasmid containing a full-length cDNA sequence of human plasminogen as a template to carry out a PCR to construct a plasmid and using Pichia Pastoris expression. The functional mutant has a dual-function of fibrinolysis and inhibiting platelet aggregation or inhibiting fibrin monomer polymerization.
Owner:GUANGDONG PHARMA UNIV
Method for constructing hog-cholera virus infectious cDNA carrier having molecule mark
ActiveCN101864445AFermentationVector-based foreign material introductionTotal rnaTechnological system
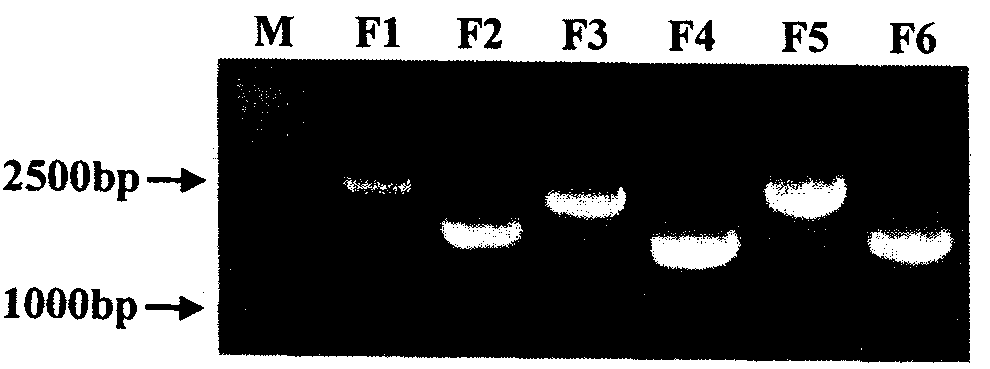
The invention relates to a virus antibody research and provides a method for constructing a hog-cholera virus infectious cDNA carrier having a molecule mark. The method comprises the following steps of: (1) diluting a hog-cholera live vaccine to 1 portion per milliliter as the experimental material, after extracting the head RNA, amplifying 6 corresponding cDNA fragments through RT-PCR section according to designed 6 pairs of primers, (2) leading in the corresponding molecule marks from F1 and F5 of cDNA fragments, (3) reforming plasmids required by overall length cDNA cloning, and (4) constructing the overall length cDNA carrier. A hog-cholera virus lapinized vaccine C stem infectious cDNA carrier pA-FL22 having a molecule mark can be acquired through the method of the invention. The permissive cells of RNA electro transferred hog-cholera virus acquired by the carrier can save the infectious progeny virus which can stably inherit the molecule mark led in the construction. By using the saving technique system of the infectious cDNA carrier and the progeny virus, the copy stem and the pathopoiesia reason of the virus can be deeply researched and the foundation for developing new marked vaccines can be settled.
Owner:ZHEJIANG UNIV
cDNA specific molecular tag and application thereof
ActiveCN106754904AQuantitatively accurateHigh precisionMicrobiological testing/measurementLibrary creationInformation analysisNucleotide
The invention discloses a cDNA specific molecular tag and application thereof. The specific molecular tag comprises a universal primer sequence as shown in SEQ ID NO.1, a transposase joint sequence P7 as shown in SEQ ID NO.2, random sequences (N)n and a poly(T) sequence with 30 T alkalis, wherein N refers to the random sequences in a quantity of n, and a specific nucleotide sequence is as shown in SEQ ID NO.3. By application of the specific molecular tag to cDNA library construction, the specific molecular tag can be added to a 5' terminal of each cDNA to make each gene unique in a process of reverse transcription of RNA into full-length cDNA, and each gene in samples can be quantified according to sequences of the specific molecular tag in subsequent information analysis.
Owner:VAZYME BIOTECH NANJING
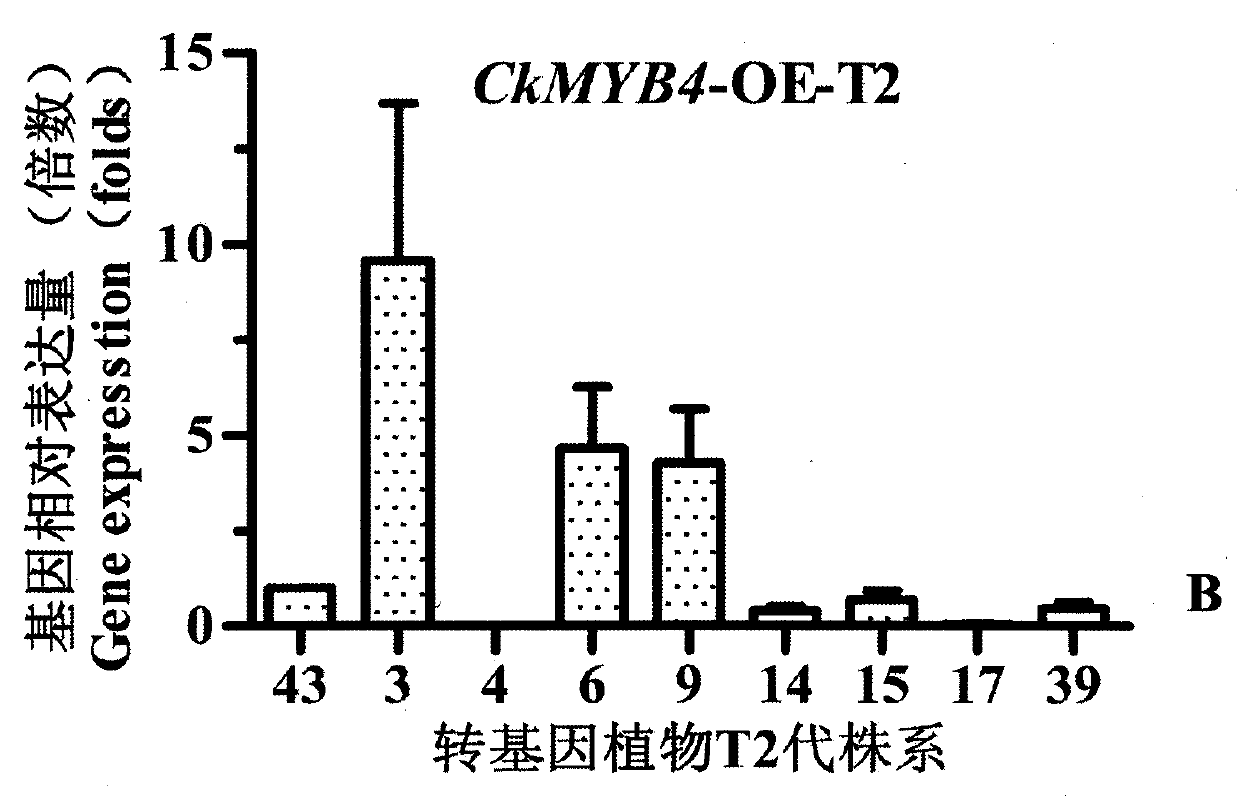
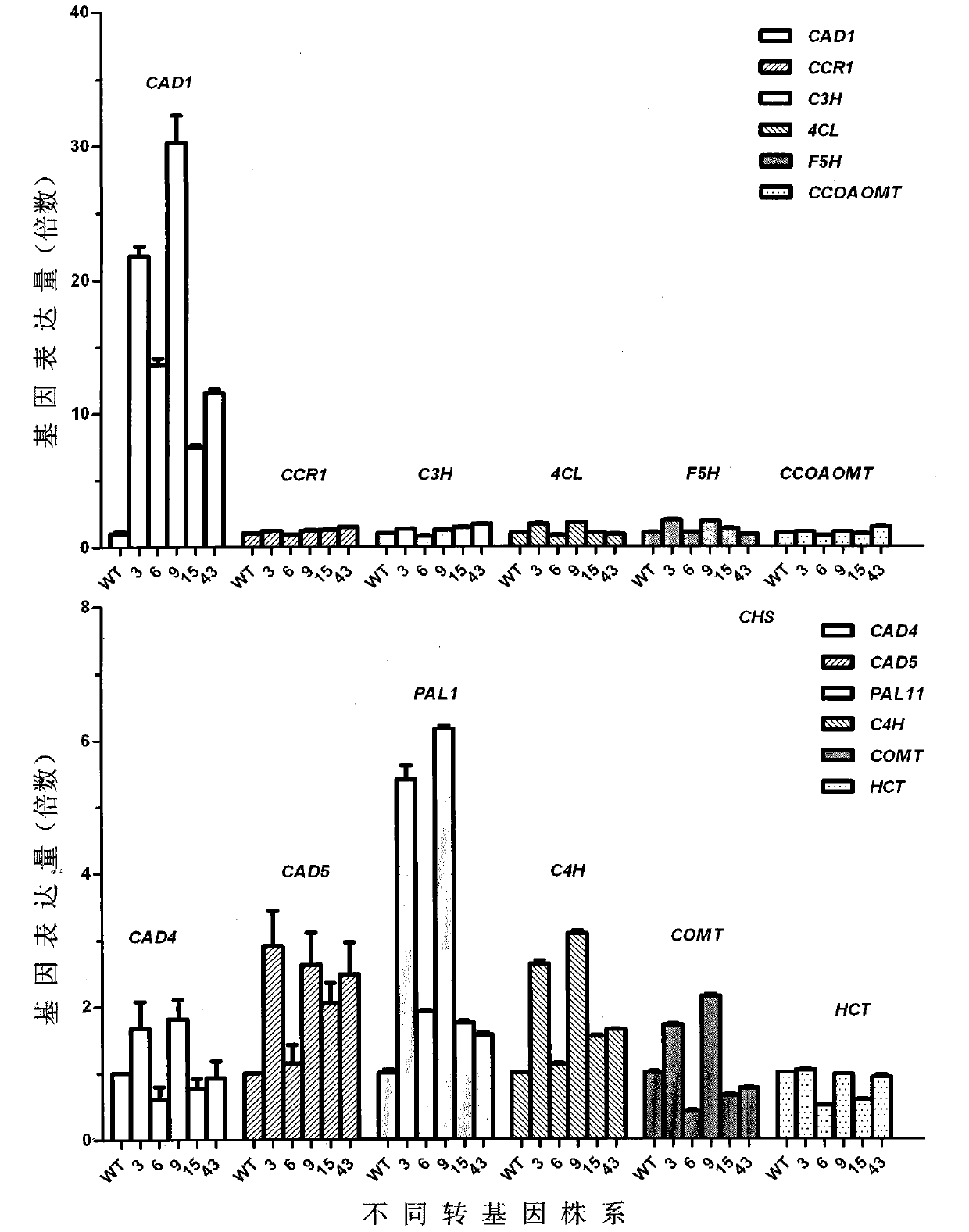
Caragana korshinskii Kom. transcription factor CkMYB4 and its gene
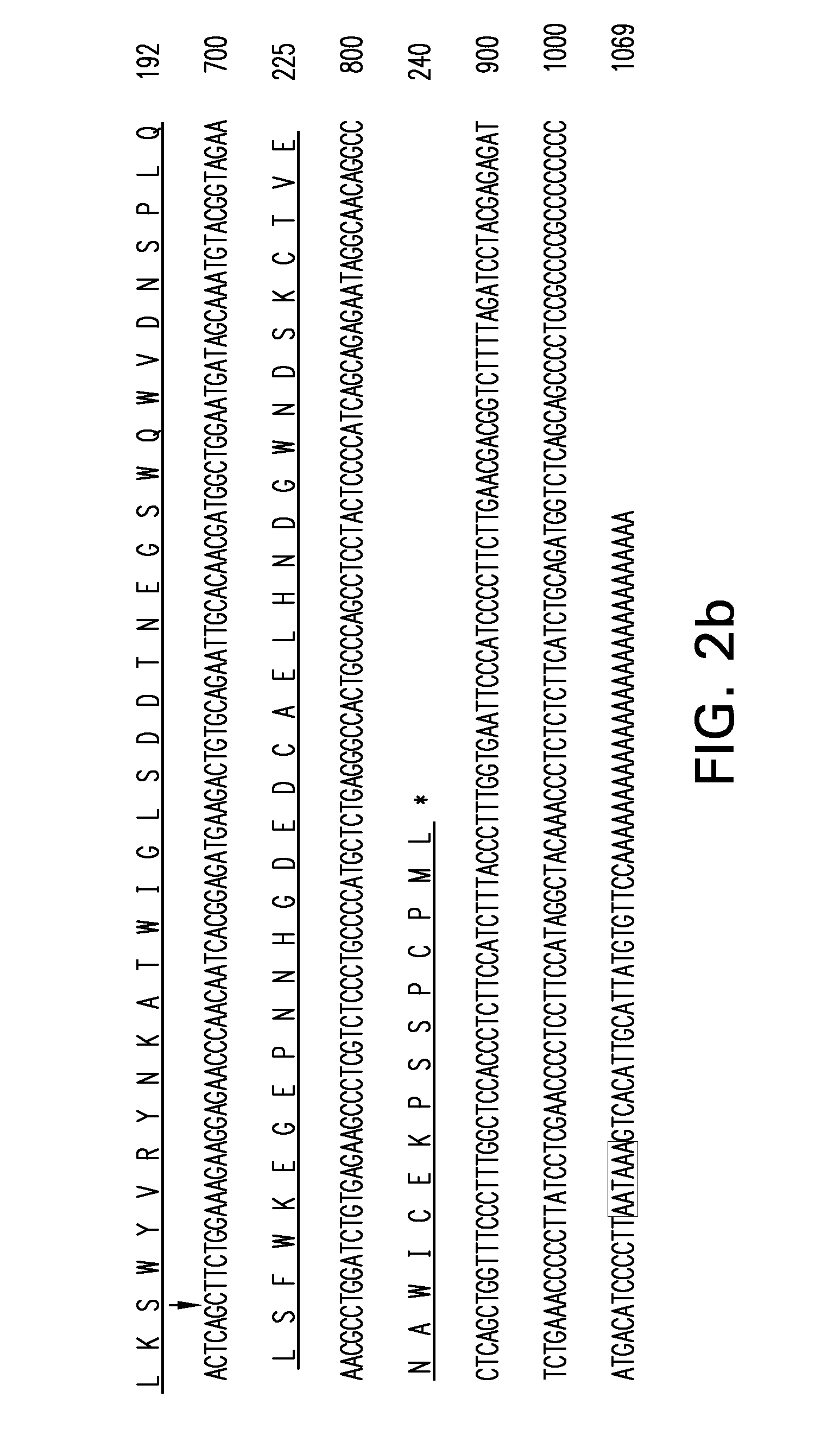
The invention provides a new CkMYB4 transcription factor and its gene. The gene originates from Leguminosae Caragana shrubs Caragana korshinskii Kom., and codes R2R3 MYB transcription factors. The full length cDNA sequence of the gene comprises 1106bps, comprises a 798bp open reading frame, and codes 266 amino acids. The cDNA sequence obtained after cloning is used for constructing a plant expression vector, and the vector is transformed into wild Arabidopis thaliana to obtain a transgenic plant. Transgenic Arabidopis thaliana lignin synthetase gene and lignin content detection results show that the gene provided by the invention can change the expression of the plant lignin synthesis gene and the lignin content.
Owner:INNER MONGOLIA AGRICULTURAL UNIVERSITY
Human diaphanous-3 gene and methods of use therefor
InactiveUS20050054826A1Cell receptors/surface-antigens/surface-determinantsSugar derivativesFull length cdnaMitotic cell
The present invention is directed to the full-length cDNA sequence encoding human diaphanous-3 (DIAPH3), to DIAPH3 encoded thereby, and to fragments of DIAPH3 and the cDNA. The present invention also provides for the use of the cDNA, and of DIAPH3, as a marker of poor prognosis of breast cancer. Because DIAPH3 appears essential for proper spindle pole formation during mitosis, DIAPH3 is a useful target for screening assays designed to identify inhibitors or modulators of DIAPH3 activity, which are useful for the treatment of cancer, particularly breast cancer. Thus, the invention further provides methods of using DIAPH3, or fragments thereof, in assays to identify such compounds.
Owner:ROSETTA INPHARMATICS LLC
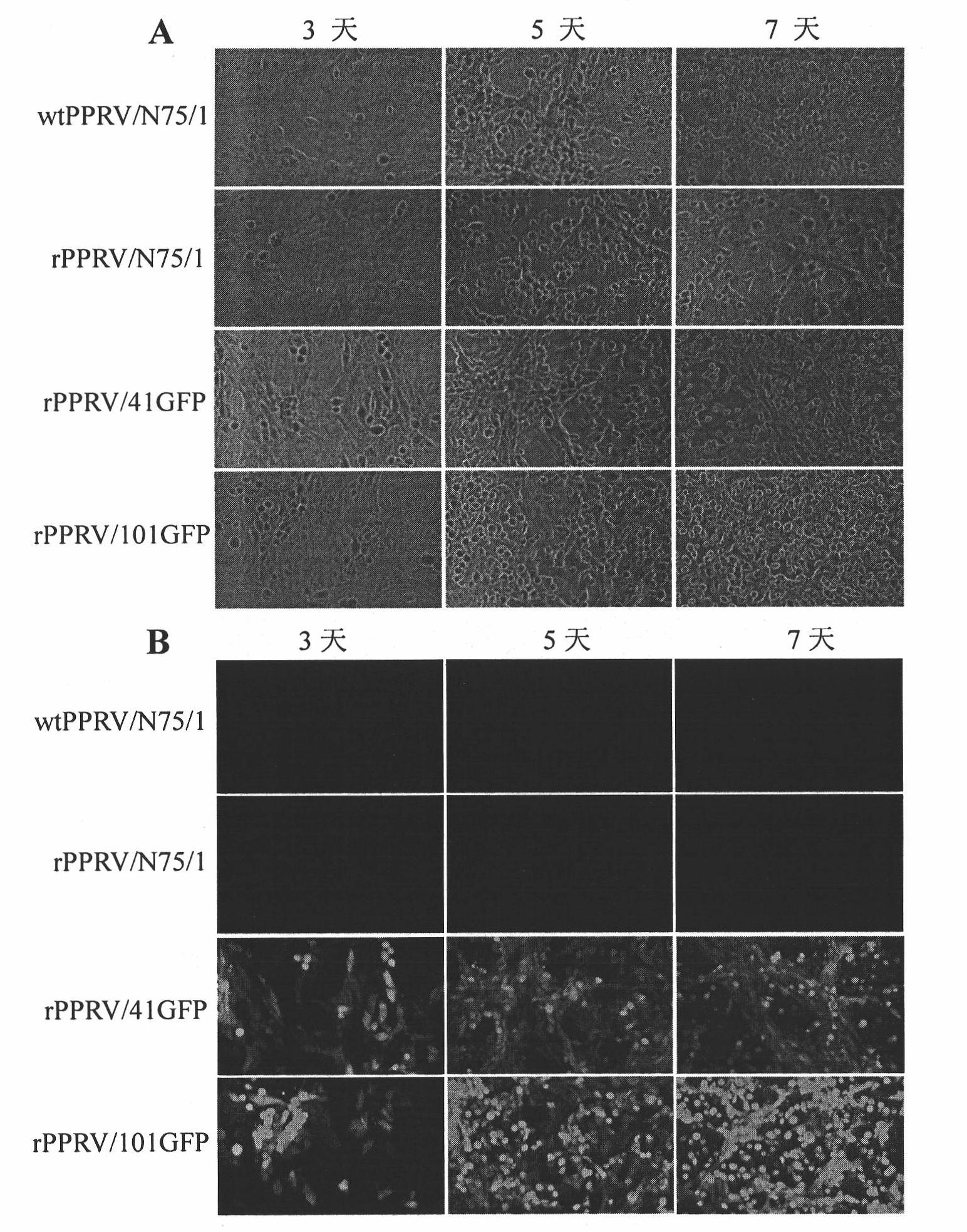
Peste des petits ruminants virus (PPRV) reverse genetic operating system and application thereof
ActiveCN102071218ALow costReduce labor costsInactivation/attenuationVector-based foreign material introductionComplementary deoxyribonucleic acidFull length cdna
The invention relates to a peste des petits ruminants virus (PPRV) reverse genetic operating system and an application thereof. The PPRV reverse genetic operating system comprises a transcription plasmid and one or more helper plasmids, wherein the transcription plasmid can express the genome full-length cDNA (complementary deoxyribonucleic acid) sequence of the PPRV; and the helper plasmid(s) can express nucleoprotein (N), phosphoprotein (P) and polymerase large protein (L) of the PPRV, and virus replication-permitting host cells of the PPRV. By using the PPRV reverse genetic operating system, the recombined PPRV is successfully saved. The establishment of the PPRV reverse genetic operating technical platform provides an excellent technical platform for the development of PPRV live vector vaccines and the fundamental research related to PPRV.
Owner:HARBIN VETERINARY RES INST CHINESE ACADEMY OF AGRI SCI
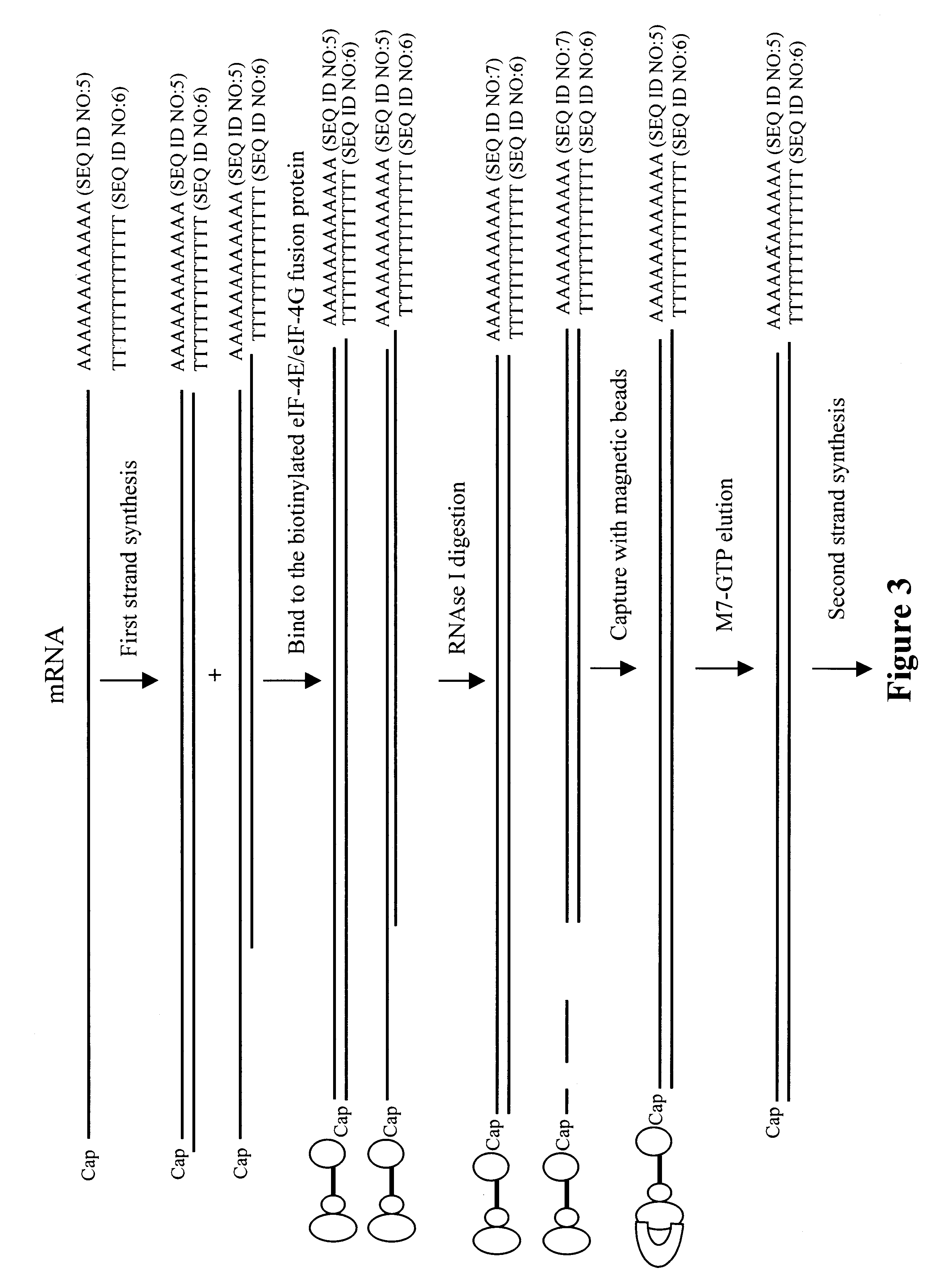
Methods and compositions for producing full length cDNA libraries
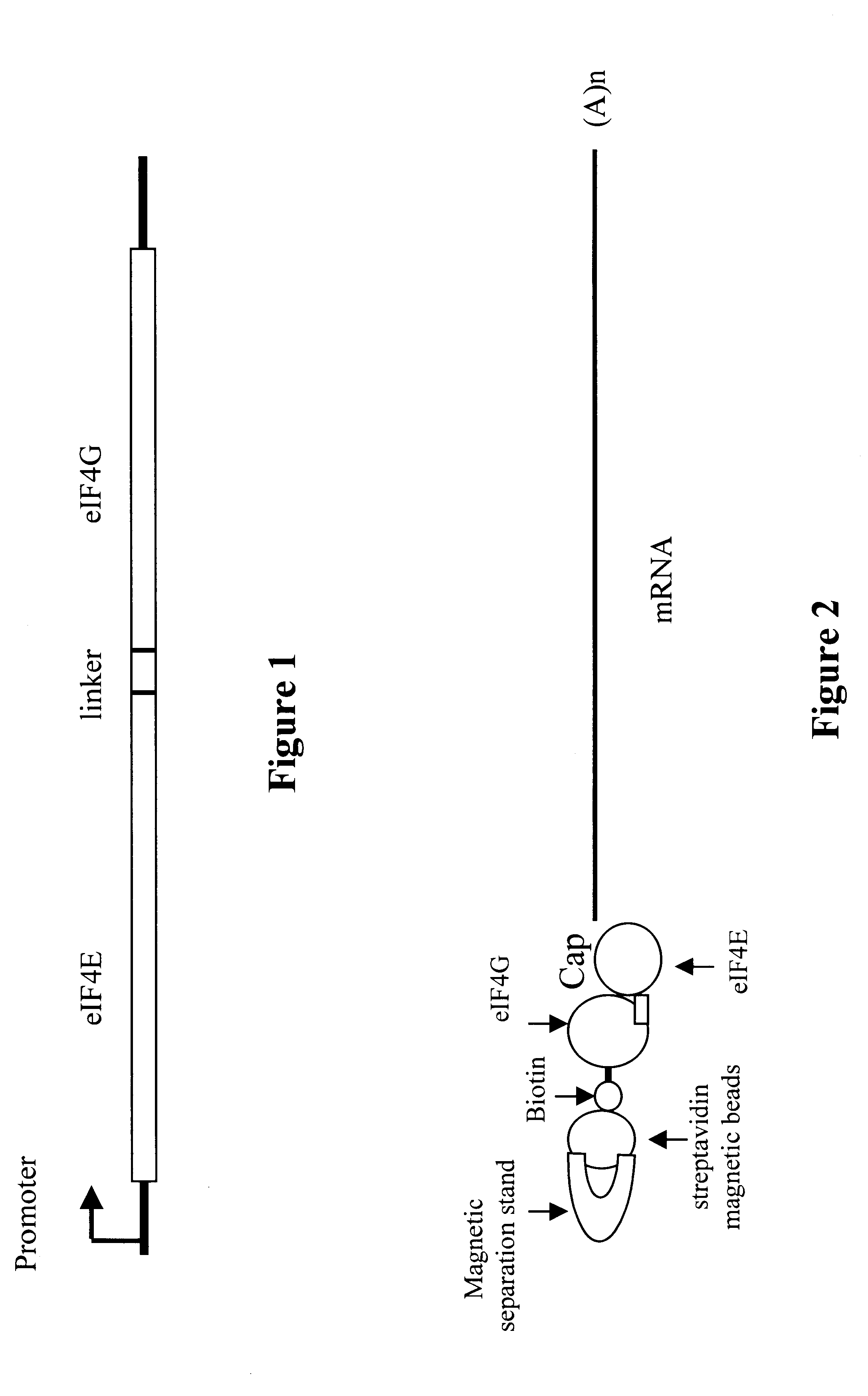
Methods and compositions are provided for producing full-length cDNA libraries. In the subject methods, full length first strand cDNAs are isolated using a fusion protein of an eIF-4E domain and an eIF-4G domain separated by a flexible linker. Also provided is the novel fusion protein employed in the subject methods, as well as nucleic acids encoding, and host cells capable of expressing, the same. Finally, kits for use in practicing the subject methods are provided. The subject invention finds use in a variety of applications in which full-length CDNA libraries are employed.
Owner:INCYTE PHARMA INC
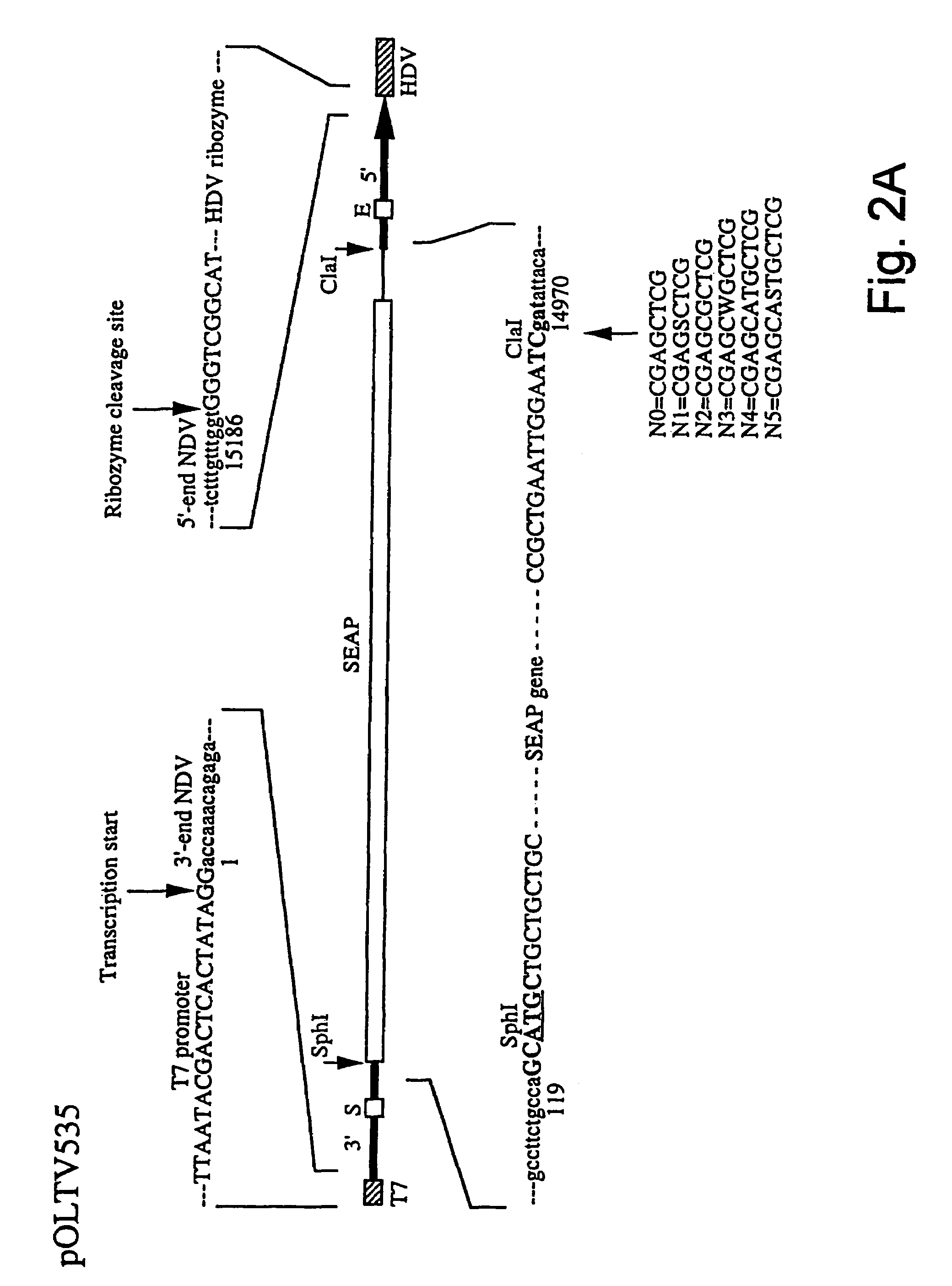
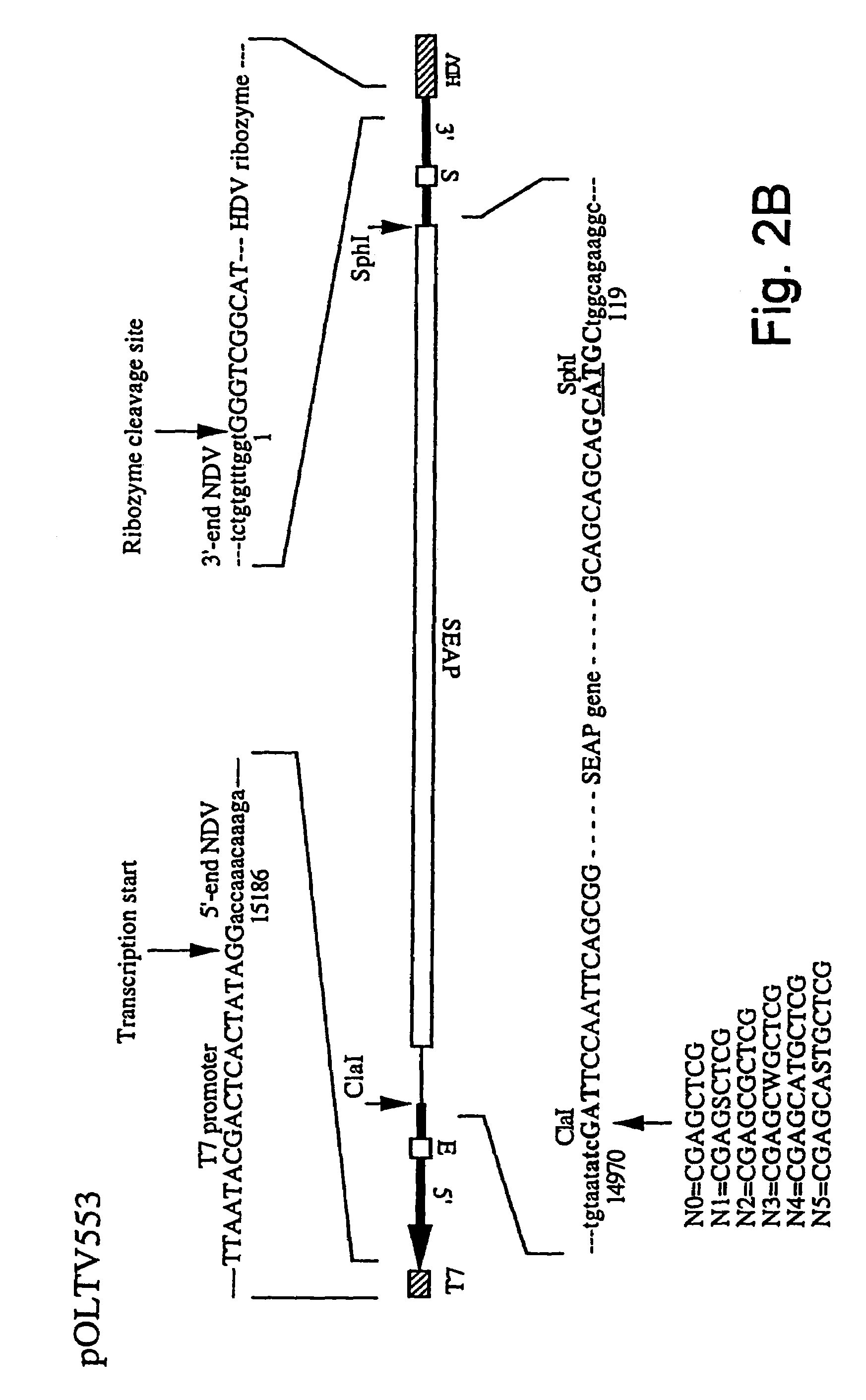
Newcastle disease virus infectious clones, vaccines and new diagnostic assays
InactiveUS7332169B2Immune responseImprove propertiesSsRNA viruses negative-senseGenetic material ingredientsAntigenVirulent characteristics
The invention relates to a process for generating infectious Newcastle disease virus (NDV) entirely from cloned full-length cDNA and to the use of vaccines and diagnostic assays generated with and derived from the process. The process offers the possibility to modify the NDV genome by means of genetic modification and allows for the introduction of mutations, deletions and / or insertions. The process can be used to modify the virulence of NDV, thus generating new attenuated live vaccines with enhanced properties. The process can be used to modify the antigenic make-up of NDV, to allow the generation of live NDV marker vaccines that can be serologically distinguished from NDV field strains.
Owner:STICHTING DIENST LANBOUWKUNDIG ONDERZOEK
Method of preparing normalized and/or subtracted cDNA
InactiveUS7049065B2Avoid disadvantagesImproving and subtractionSugar derivativesMicrobiological testing/measurementSingle-Stranded RNAFull length cdna
Owner:YOSHIHIDE HAYASHIZAKI +2
Method for cultivating transgenic plant with improved insect resistance by using RNA interference technology and special DNA fragment thereof
ActiveCN101851622ANormal growth and reproductionImprove insect resistanceClimate change adaptationVector-based foreign material introductionNicotiana tabacumWild type
The invention discloses a method for cultivating a transgenic plant with improved insect resistance by using RNA interference technology and a special DNA fragment thereof. The DNA fragment is a gene fragment shown as a formula I, wherein a forward sequence (SEQ) is any one fragment which at least comprises 19bp in full-length cDNA fragments of an ATP synthase E subunit gene; a reverse sequence (SEQ) and the forward sequence (SEQ) are mutually reversely complementary; X is a spacer sequence between the forward sequence (SEQ) and the reverse sequence (SEQ), and the X and the forward sequence (SEQ) or the X and the reverse sequence (SEQ) are both not complementary mutually; and the full-length cDNA of the ATP synthase E subunit gene is shown as a sequence 1 in a sequence table. The invention also aims to provide application of the ATP synthase E subunit gene serving as an RNA interference target gene in cultivating the transgenic plant with improved insect resistance. Experiments prove that aphids inoculated to wild-type tobacco grow and reproduce normally; and aphids inoculated to transgenic tobacco starts to die on the fourth day, dead aphid bodies is blackened after five days and no live aphids can be seen after seven days. The formula (I) is forward sequence (SEQ)-X- reverse sequence (SEQ).
Owner:INST OF MICROBIOLOGY - CHINESE ACAD OF SCI
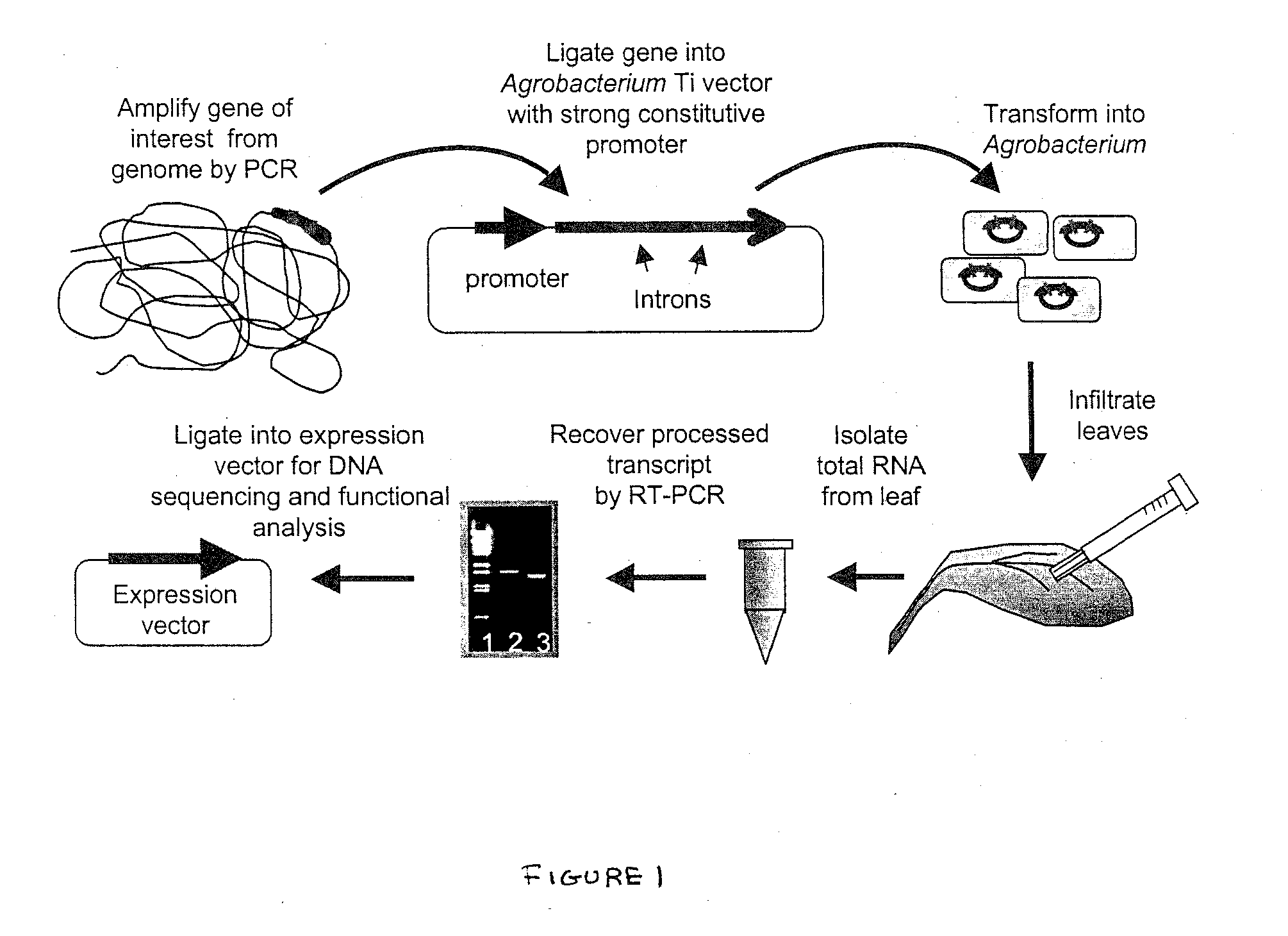
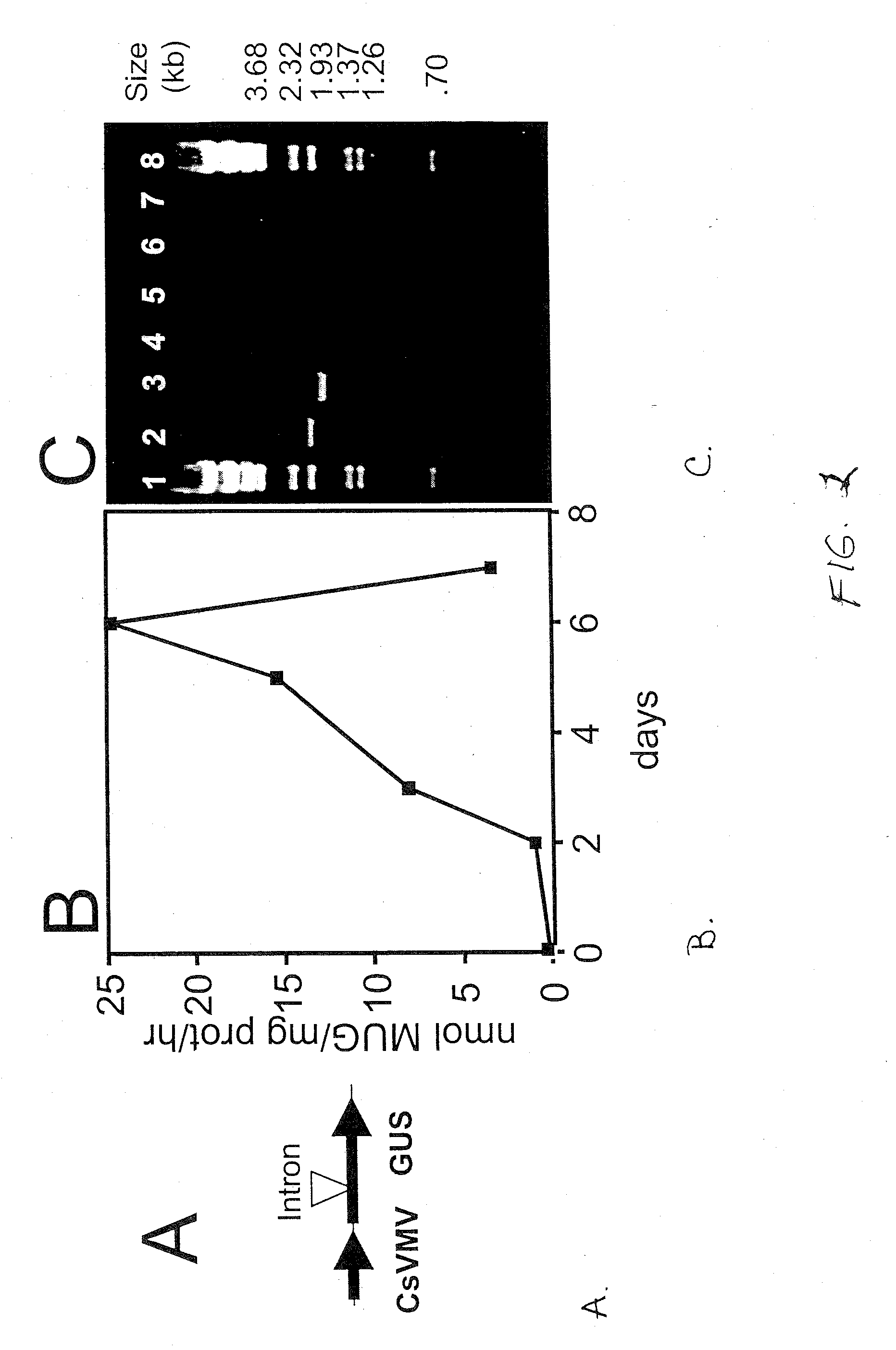
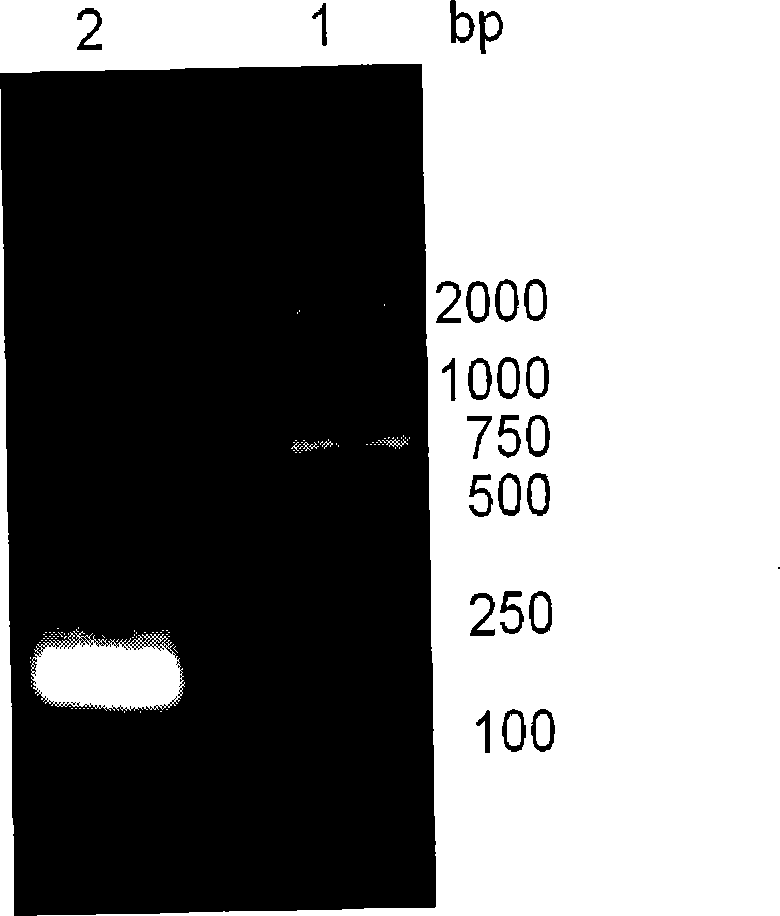
Methods for Splicing Plant Genes
InactiveUS20070089198A1Other foreign material introduction processesFermentationPcr assayFunctional testing
A method for the recovery of full-length cDNAs for plant genes containing introns is described. The method utilizes Agrobacterium-mediated transient expression coupled with an RT-PCR assay to facilitate expression cloning of processed transcripts. Subsequent expression of intron-less cDNAs in a suitable prokaryotic or eucaryotic host provides for direction functional testing of the encoded gene product.
Owner:UNIV OF KENTUCKY RES FOUND
Recombinant thymosin beta 4 two repeat protein and preparation thereof
InactiveCN101434651ALow immunogenicityOvercoming the defect of unstable expressionThymopoietinsPeptide preparation methodsEscherichia coliMouse Lymphocyte
The invention relates to recombinant thymosin Beta 4 two-repeat protein and a preparation method thereof. Human thymosin Beta 4 two-repeat gene expression vector pET-22b(+)-T Beta (2) is constructed through recombinant human thymosin Beta 4 full length cDNA combined with PCR technology, human thymosin Beta 4 that can not be expressed directly in colibacillus is highly actively expressed in colibacillus in the form of two-repeat and purified human thymosin Beta 4 two-repeat protein has biologic activity, can promote multiplication of lymphocyte of mice, has low immunogenicity and lays a foundation for the further research and wide application of human thymosin Beta 4.
Owner:FOURTH MILITARY MEDICAL UNIVERSITY
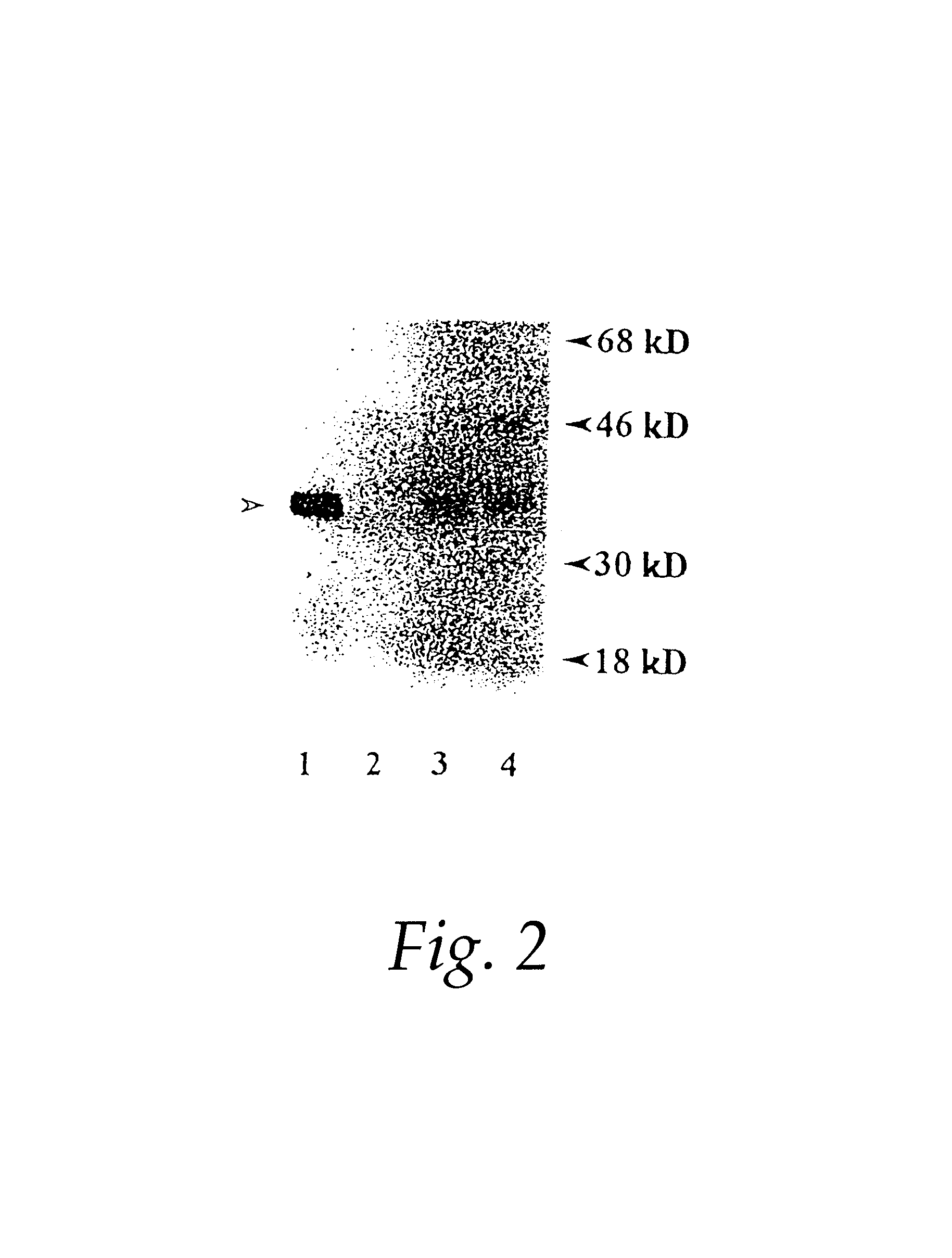
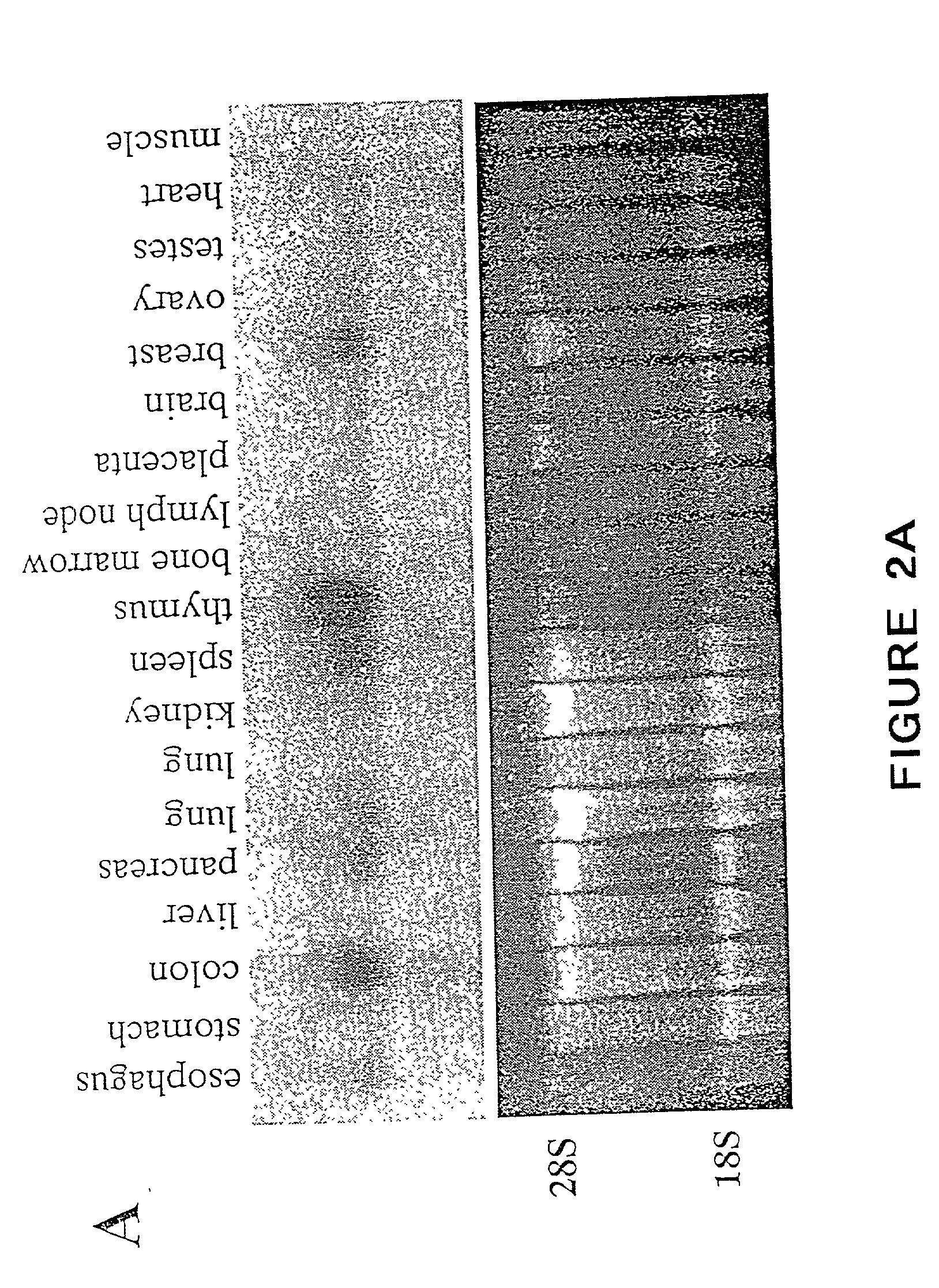
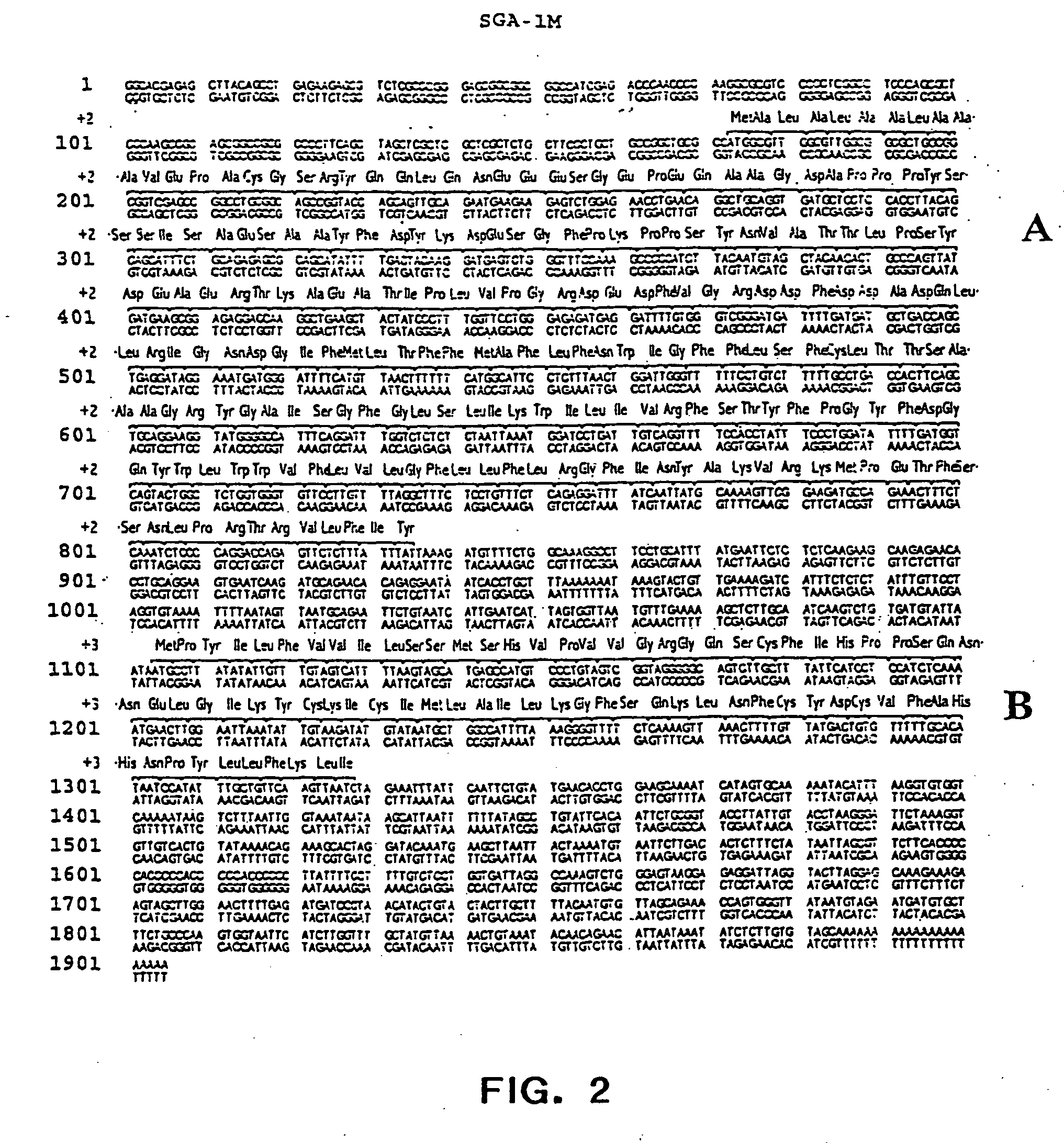
Sga-1m, a cancer associated antigen, and uses thereof
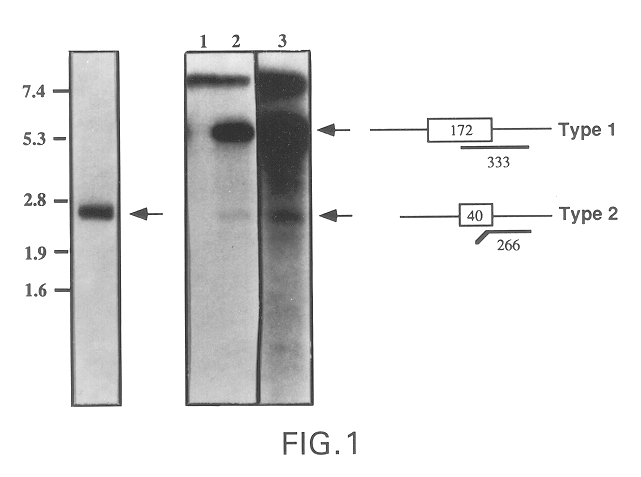
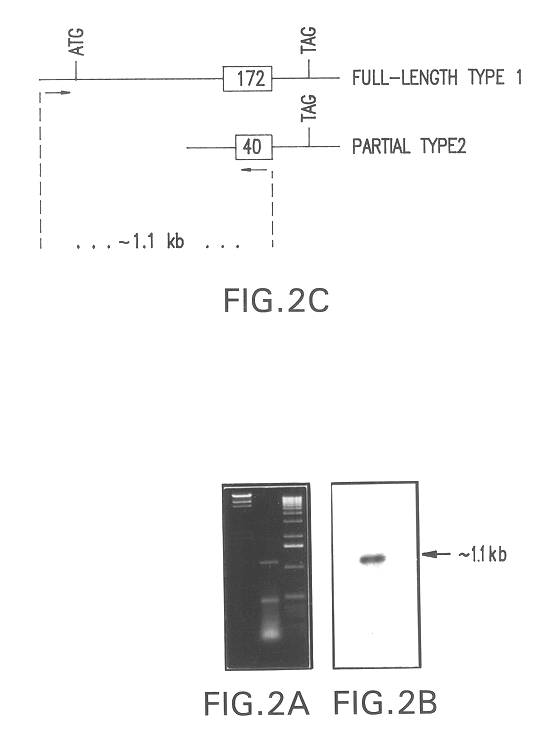
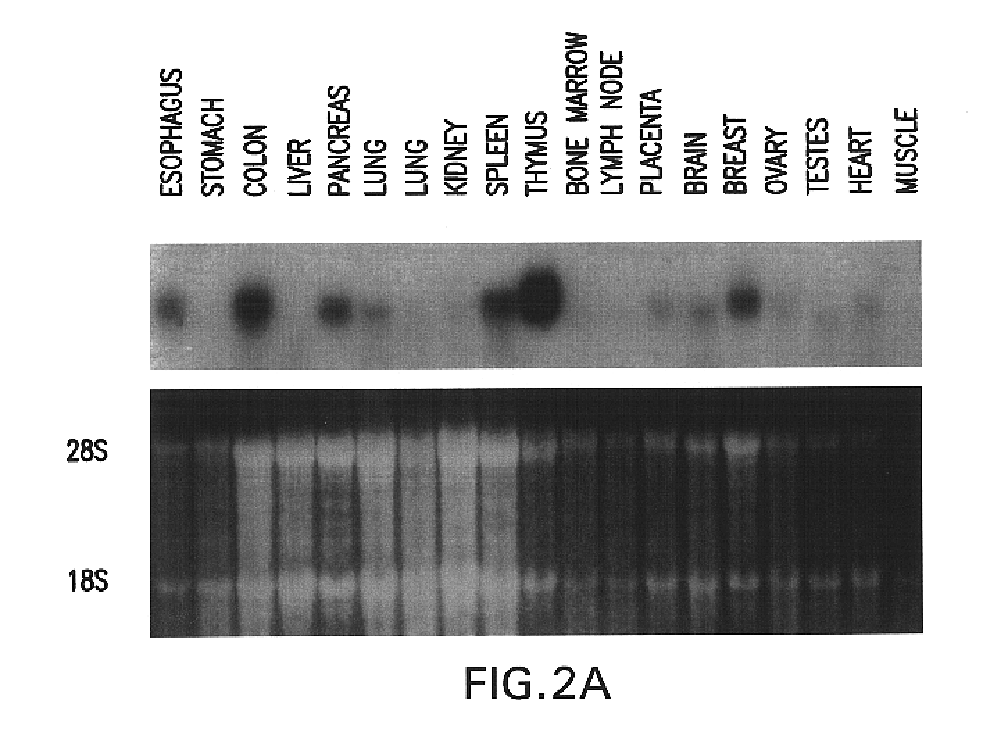
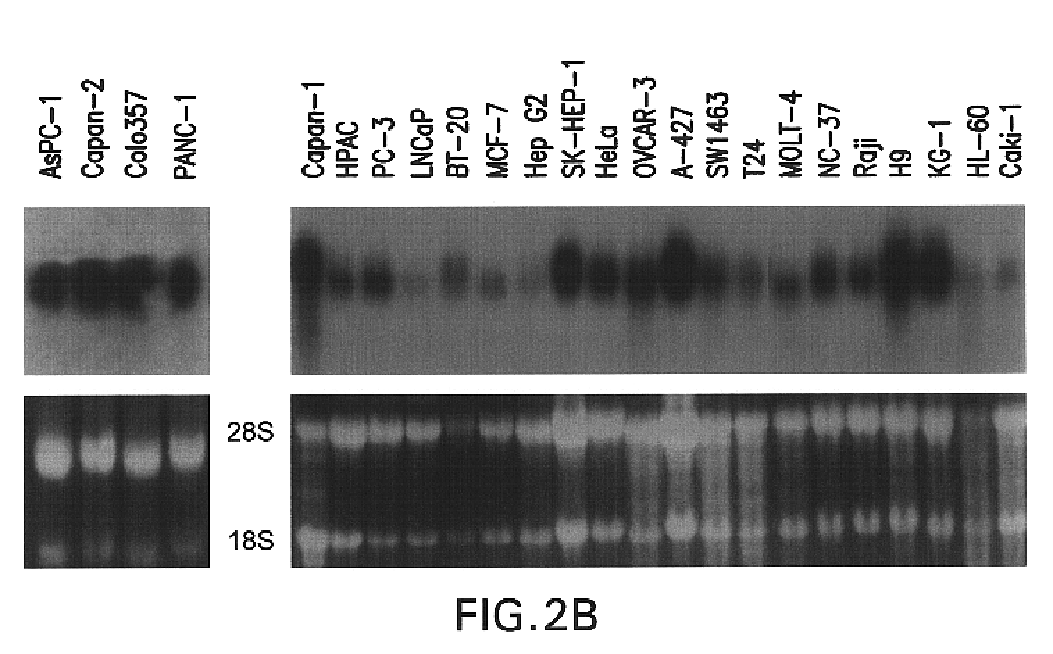
The present invention relates to a gene and gene product, SGA-1M that is differentially expressed in cancer tissues and cell lines. Suppression Subtractive Hybridization and microarray screening were used to screen for differential expression of SGA-1M in cancer tissues and cell lines. Expression analysis has demonstrated overexpression of SGA-1M in breast cancer tissue and breast cancer derived cell lines, in ovarian cancer, skin cancer, a cancer of the lymphoid system, thyroid cancer, pancreatic cancer, stomach cancer, and lung cancer. The gene is expressed as a 1.95 kb mRNA. The full length cDNA comprises two open reading frames encoding polypeptides of 221 and 75 amino acids, respectively. Monitoring expression levels of SGA-1M is useful for the diagnosis and prognosis of cancer as well as for evaluating the risk of developing certain types of cancers and the risk of metastasis of cancer. Reagents that target SGA-1M are useful for the treatment of cancer.
Owner:SEATTLE GENETICS INC
Human cDNA clones comprising polynucleotides encoding polypeptides and methods of their use
InactiveUS20060084799A1Diagnostic and prophylactic and therapeutic effectModulating biological activityPeptide/protein ingredientsGenetic material ingredientsFull length cdnaPolynucleotide
The invention provides novel human full-length cDNA clones, novel polynucleotides, related polypeptides, related nucleic acid and polypeptide compositions, and related modulators, such as antibodies and small molecule modulators. The invention also provides methods to make and use these cDNA clones, polynucleotides, polypeptides, related compositions, and modulators. These methods include diagnostic, prophylactic and therapeutic applications. The compositions and methods of the invention are useful in treating proliferative disorders, e.g., cancers, and inflammatory, immune, bacterial, and viral disorders.
Owner:FIVE PRIME THERAPEUTICS
Preparation method and application of molecular marker of rape male sterile restoring gene
InactiveCN102220316AHigh speedImprove identification efficiencyMicrobiological testing/measurementDNA preparationSequence analysisRapid identification
The invention discloses a preparation method and an application of a molecular marker of a rape male sterility restoring gene. The preparation method comprises the following steps of A, designing degenerate primers according to PPR gene conservative amino acid sites, B, extracting a lamina DNA of a rape cytoplasm male sterility restoring line through a cetyltrimethyl ammonium bromide (CTAB) method and magnifying the lamina DNA through a touchdown polymerase chain reaction (PCR) process to obtain a specific restoring gene candidate segment, C, carrying out recovery, conversion, sequencing and sequence analysis processes on PCR products, D, carrying out 5 ' RACE and 3 ' RACE processes and splicing a sequencing result to obtain a full-length cDNA sequence, and E, designing specific primers in 5 ' UTR and 3 ' UTR zones of the full-length cDNA sequence according to the full-length cDNA sequence to obtain a molecular marker. The invention also discloses an application of a restoring gene molecular marker in hybrid rape parent breeding. The preparation method has the advantages of improvements of a speed of restorer seeding and an efficiency of verification, simple and easy method, good operability, rapid identification speed of restoring genes, great reduction of necessary workload of test cross and progeny planting observation in a restorative verification process, and improvement of an efficiency of hybrid seeding.
Owner:INST OF OIL CROPS RES CHINESE ACAD OF AGRI SCI
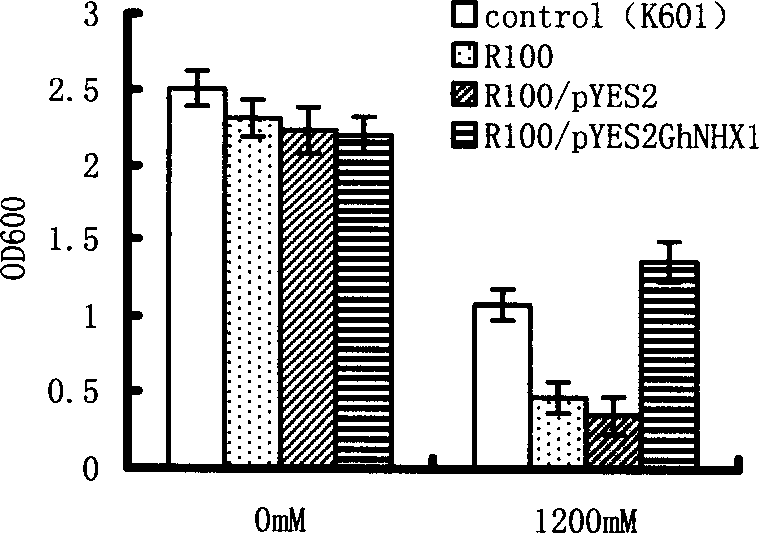
Cotton Na+/H+ reverse transport protein gene and its cloning method and use
The present invention relates to the cloning, recombination and salt tolerance analysis of cotton Na+ / H+ reverse transport protein GhNHX1 gene, and belongs to the field of molecular biology and biological technology. Total RNA is extracted from cotton leaf treated with 0.4 mol / L NaCl solution and inversely transcribed into cDNA. Facultative primer is used for conventional PCR to obtain intermediate segment, and whole length cDNA is obtained through fast amplification in 3' and 5' ends. The gene is transformed to salt-sensitive yeast mutant to restore its salt tolerant capacity to some degree; and the gene is further transformed to tobacco to obtain transgenic plant capabl eof growing normally in salt concentration of 200 mmol / L. Therefore, the gene is one important salt tolerant gene and may be used in raising the salt tolerant capacity of plant for plantation in saline land.
Owner:SHANDONG AGRICULTURAL UNIVERSITY
Popular searches
Features
- R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
Why Patsnap Eureka
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Social media
Patsnap Eureka Blog
Learn More Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com