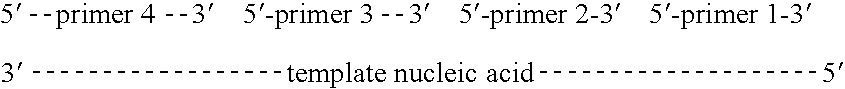Patents
Literature
Hiro is an intelligent assistant for R&D personnel, combined with Patent DNA, to facilitate innovative research.
3252results about "Nucleotide libraries" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
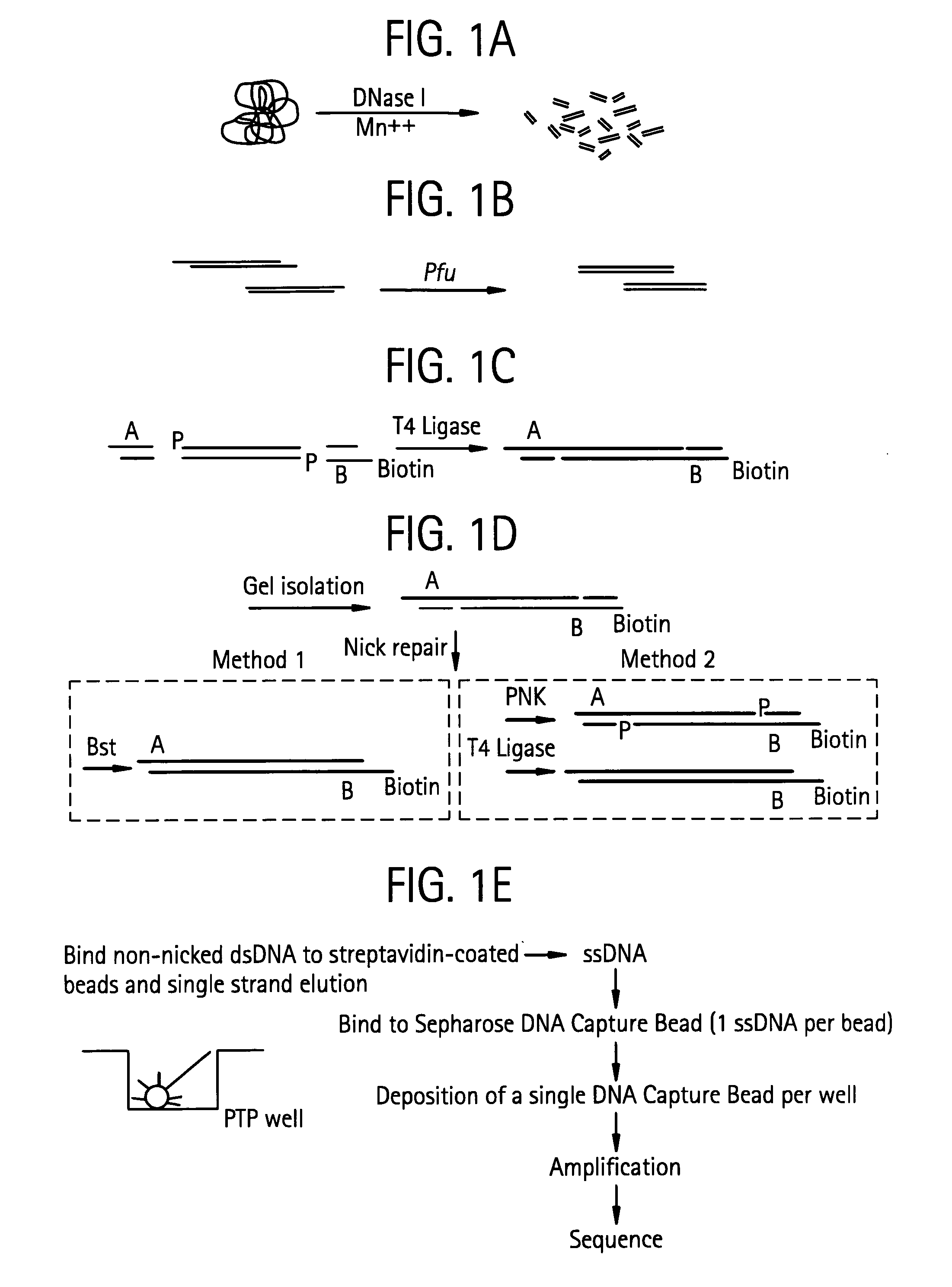
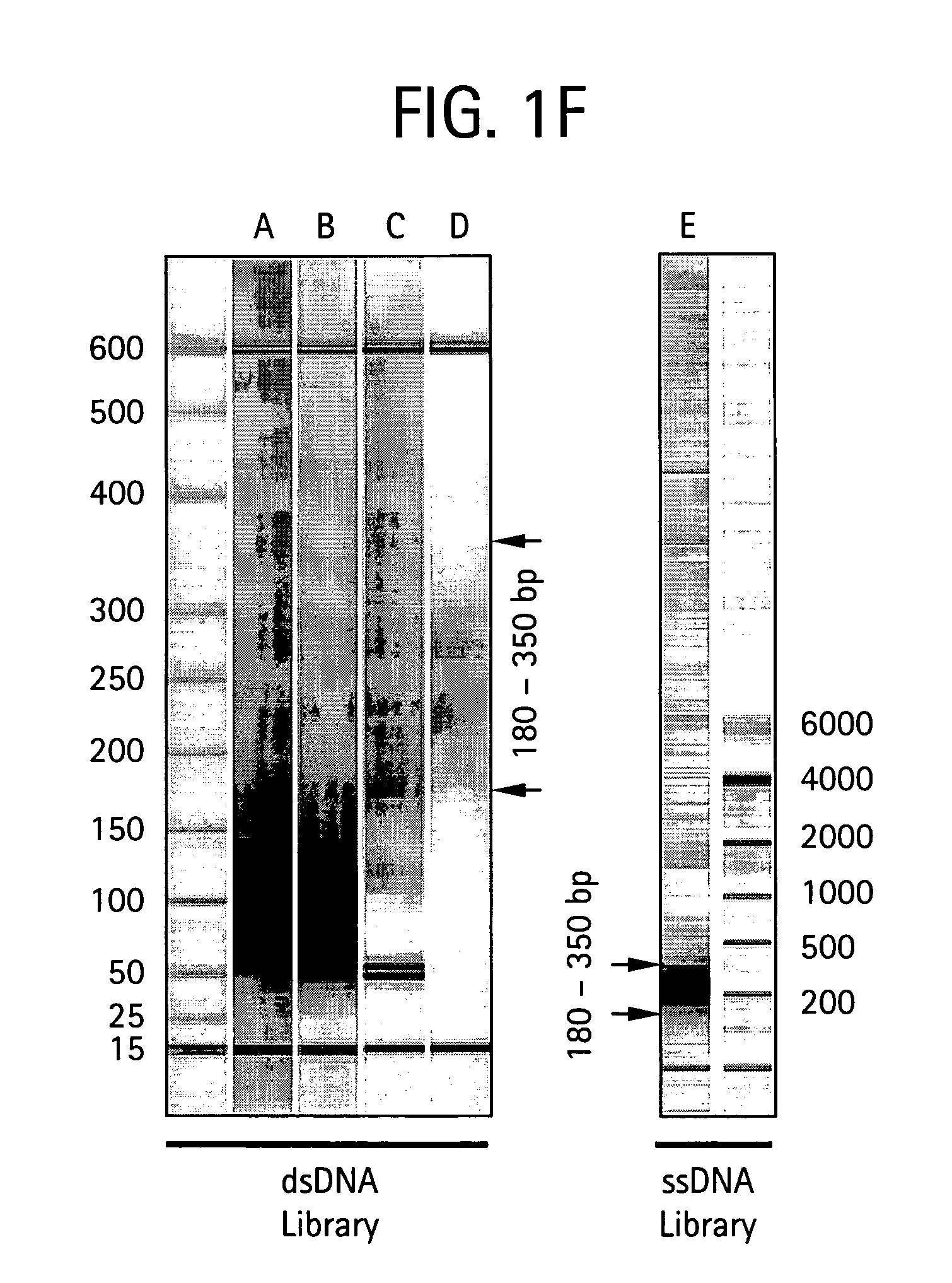
Methods of amplifying and sequencing nucleic acids
An apparatus and method for performing rapid DNA sequencing, such as genomic sequencing, is provided herein. The method includes the steps of preparing a sample DNA for genomic sequencing, amplifying the prepared DNA in a representative manner, and performing multiple sequencing reaction on the amplified DNA with only one primer hybridization step.
Owner:454 LIFE SCIENCES CORP
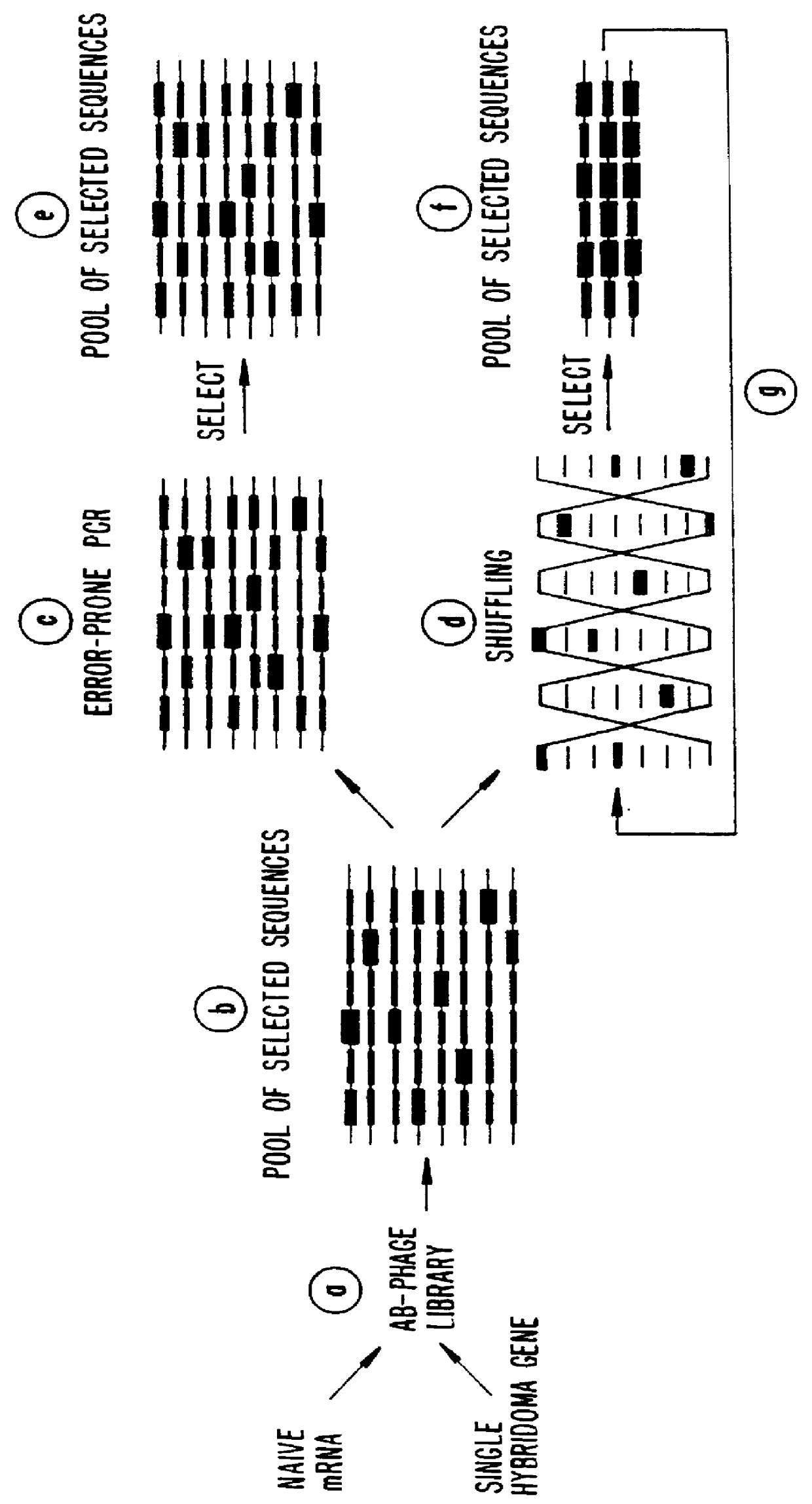
Methods for generating polynucleotides having desired characteristics by iterative selection and recombination
InactiveUS6117679ALess immunogenicLibrary screeningDirected macromolecular evolutionMutated proteinNucleic acid sequencing
A method for DNA reassembly after random fragmentation, and its application to mutagenesis of nucleic acid sequences by in vitro or in vivo recombination is described. In particular, a method for the production of nucleic acid fragments or polynucleotides encoding mutant proteins is described. The present invention also relates to a method of repeated cycles of mutagenesis, shuffling and selection which allow for the directed molecular evolution in vitro or in vivo of proteins.
Owner:CODEXIS MAYFLOWER HLDG LLC
Analying polynucleotide sequences
InactiveUS6054270AStable duplexReduce impactSequential/parallel process reactionsSugar derivativesHybridization reactionSequence determination
This invention provides an apparatus and method for analyzing a polynucleotide sequence; either an unknown sequence or a known sequence. A support, e.g. a glass plate, carries an array of the whole or a chosen part of a complete set of oligonucleotides which are capable of taking part in hybridization reactions. The array may comprise one or more pair of oligonucleotides of chosen lengths. The polynucleotide sequence, or fragments thereof, are labelled and applied to the array under hybridizing conditions. Applications include analyses of known point mutations, genomic fingerprinting, linkage analysis, characterization of mRNAs, mRNA populations, and sequence determination.
Owner:OXFORD GENE TECH
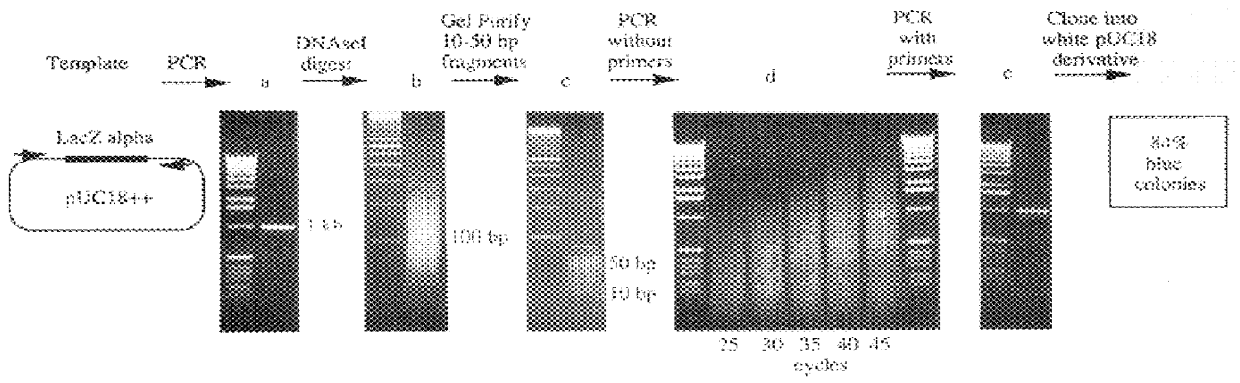
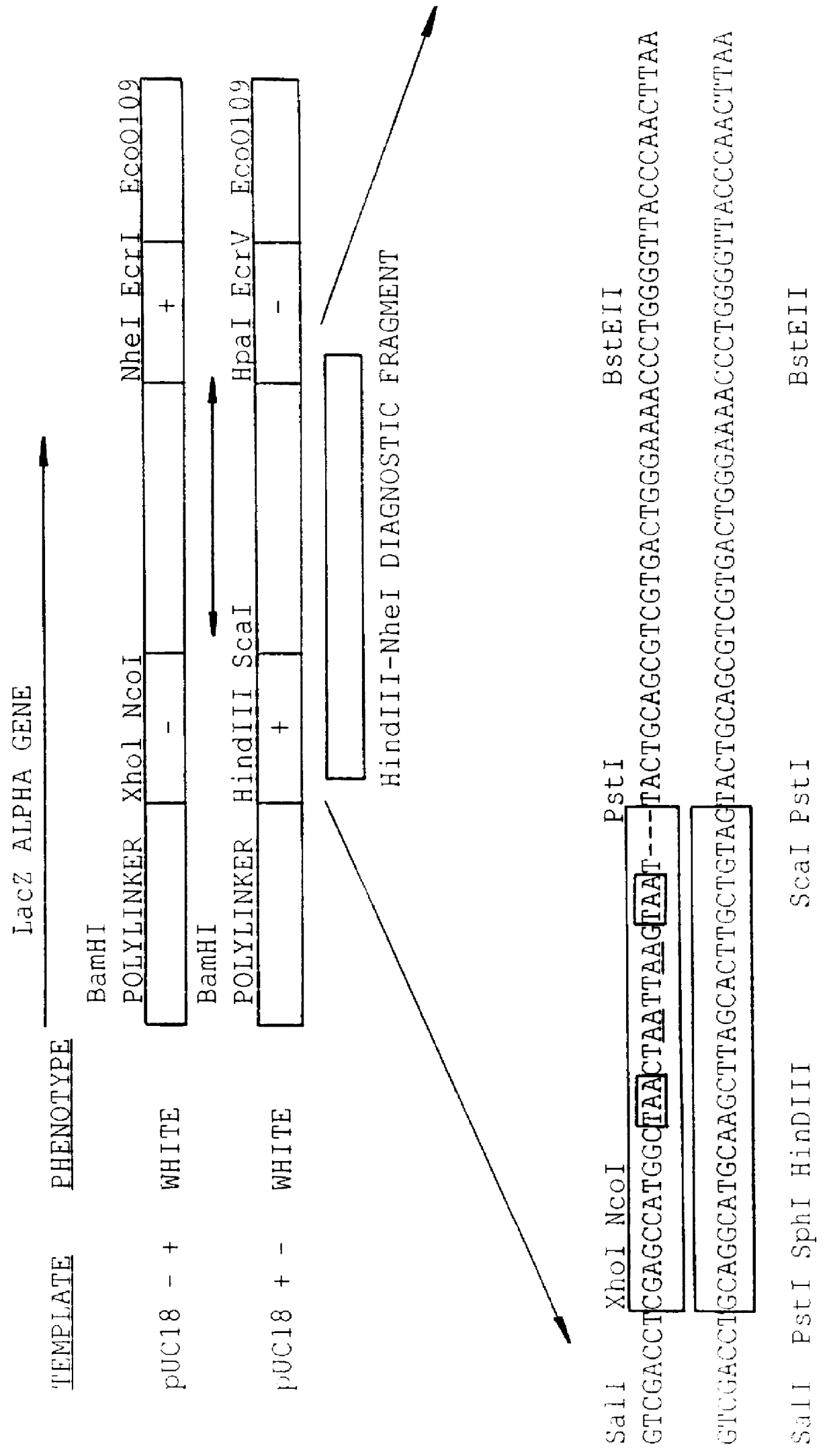
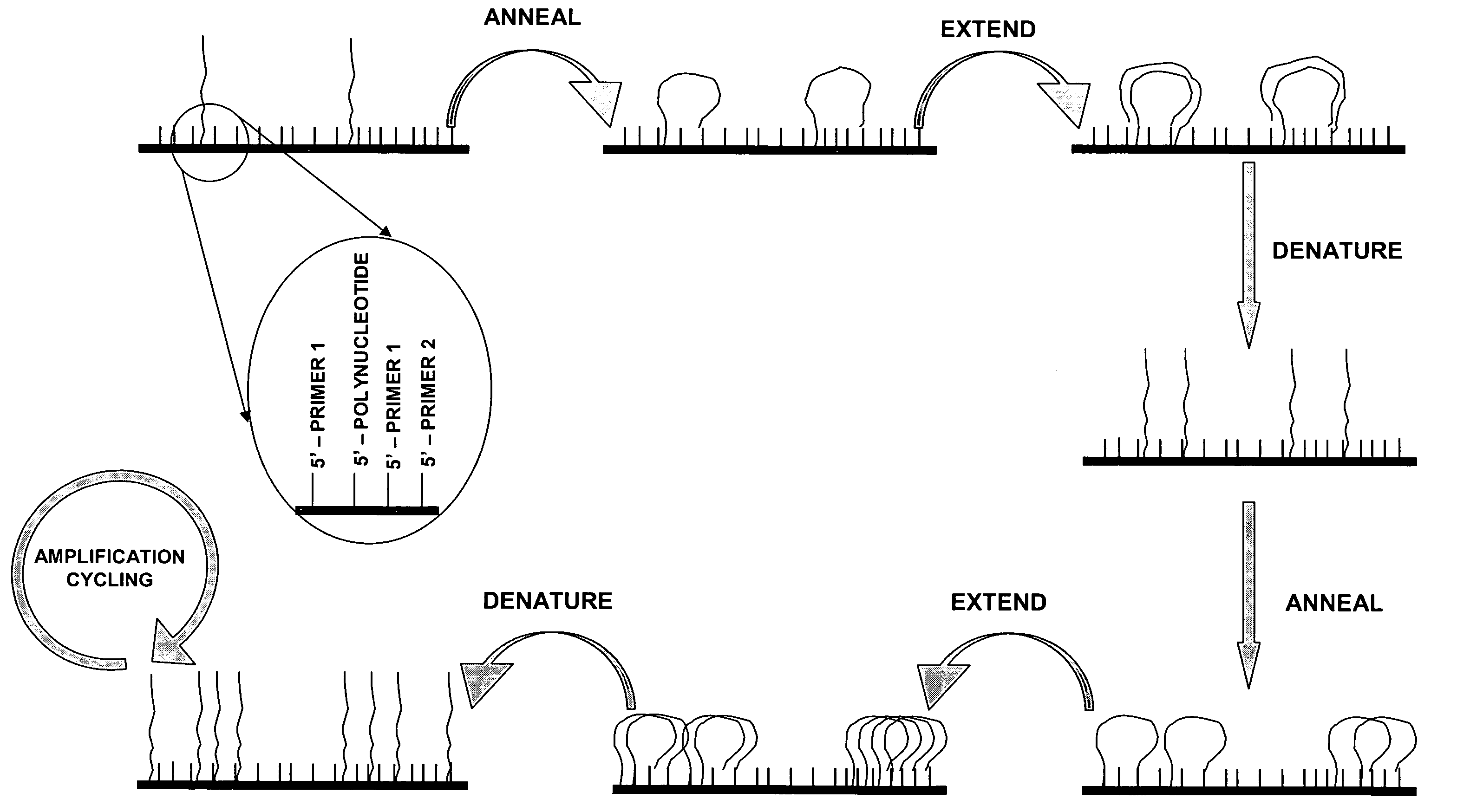
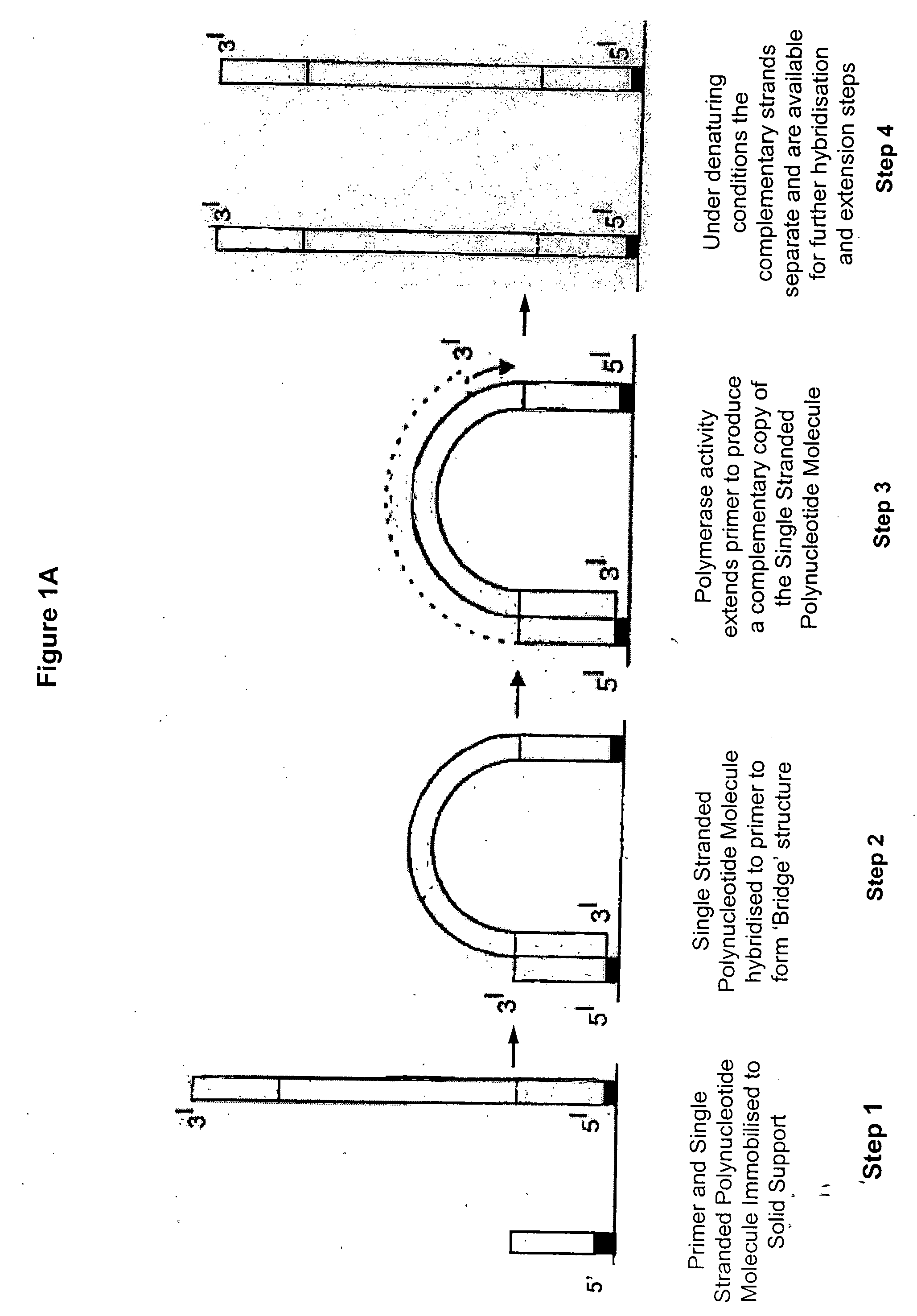
Isothermal methods for creating clonal single molecule arrays
InactiveUS20080009420A1Efficient amplificationIncrease diversityNucleotide librariesMicrobiological testing/measurementMicrobiologyNucleic acid sequencing
The present invention is directed to a method for isothermal amplification of a plurality of different target nucleic acids, wherein the different target nucleic acids are amplified using universal primers and colonies produced thereby can be distinguished from each other. The method, therefore, generates distinct colonies of amplified nucleic acid sequences that can be analyzed by various means to yield information particular to each distinct colony.
Owner:SOLEXA
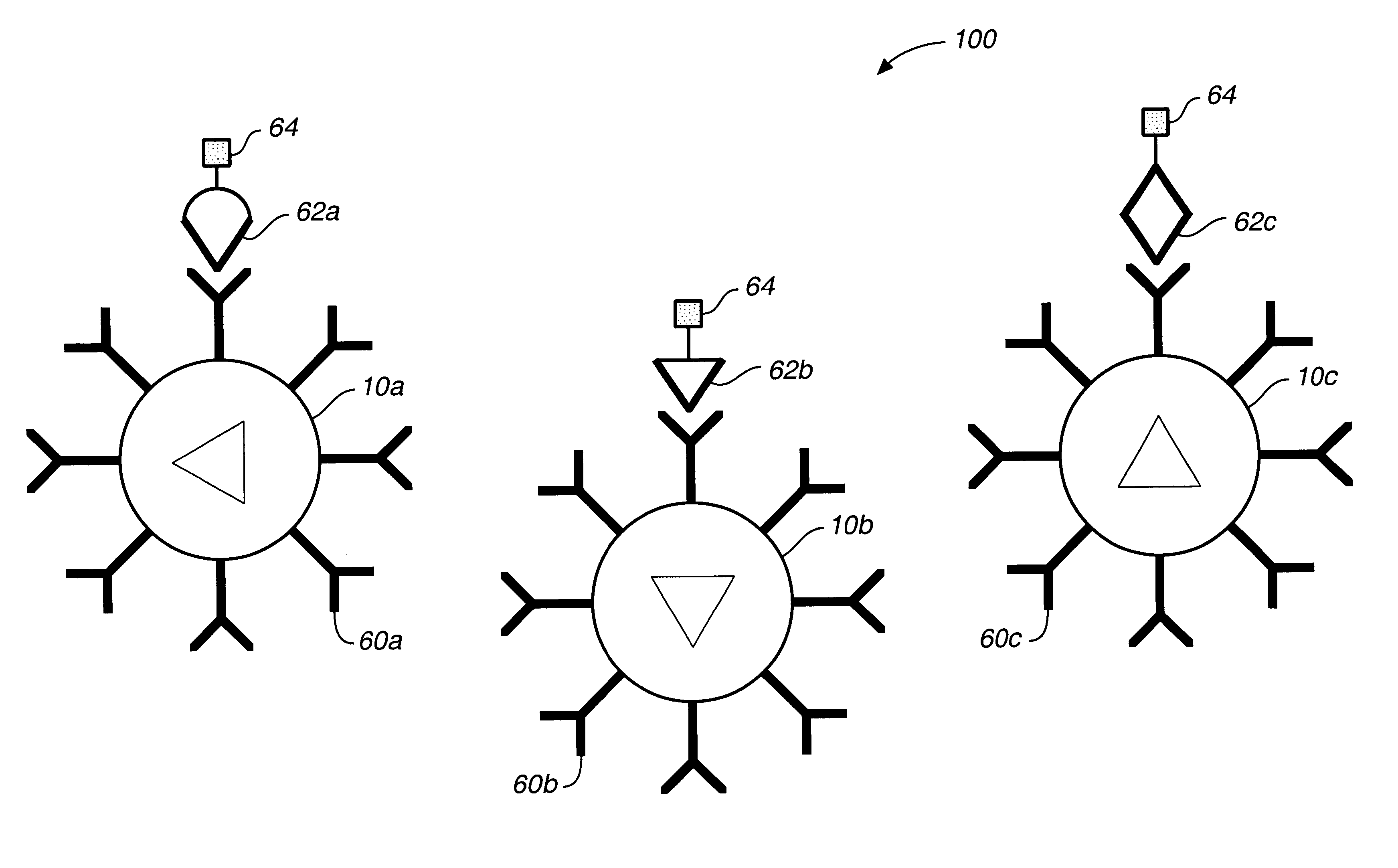
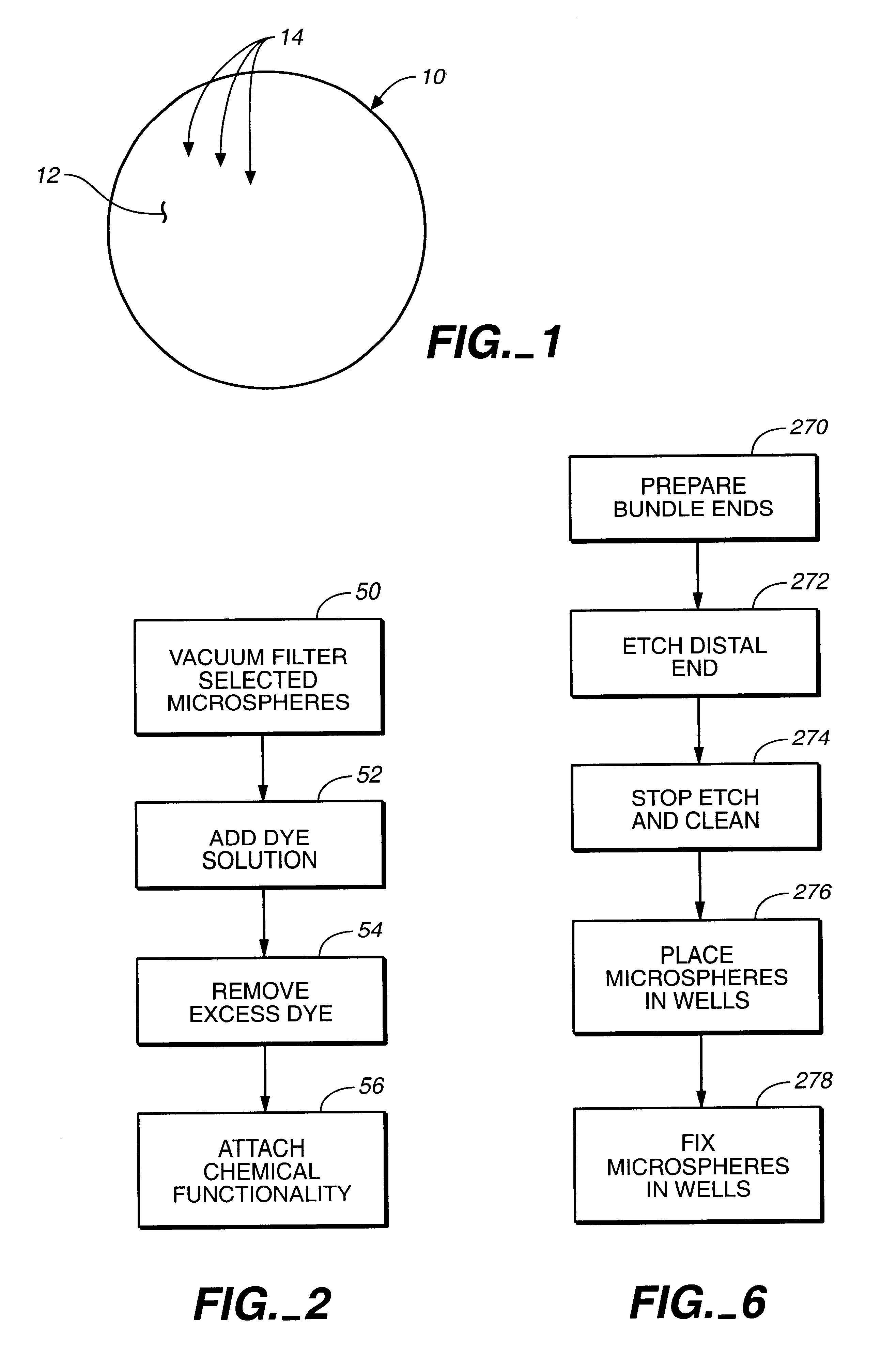
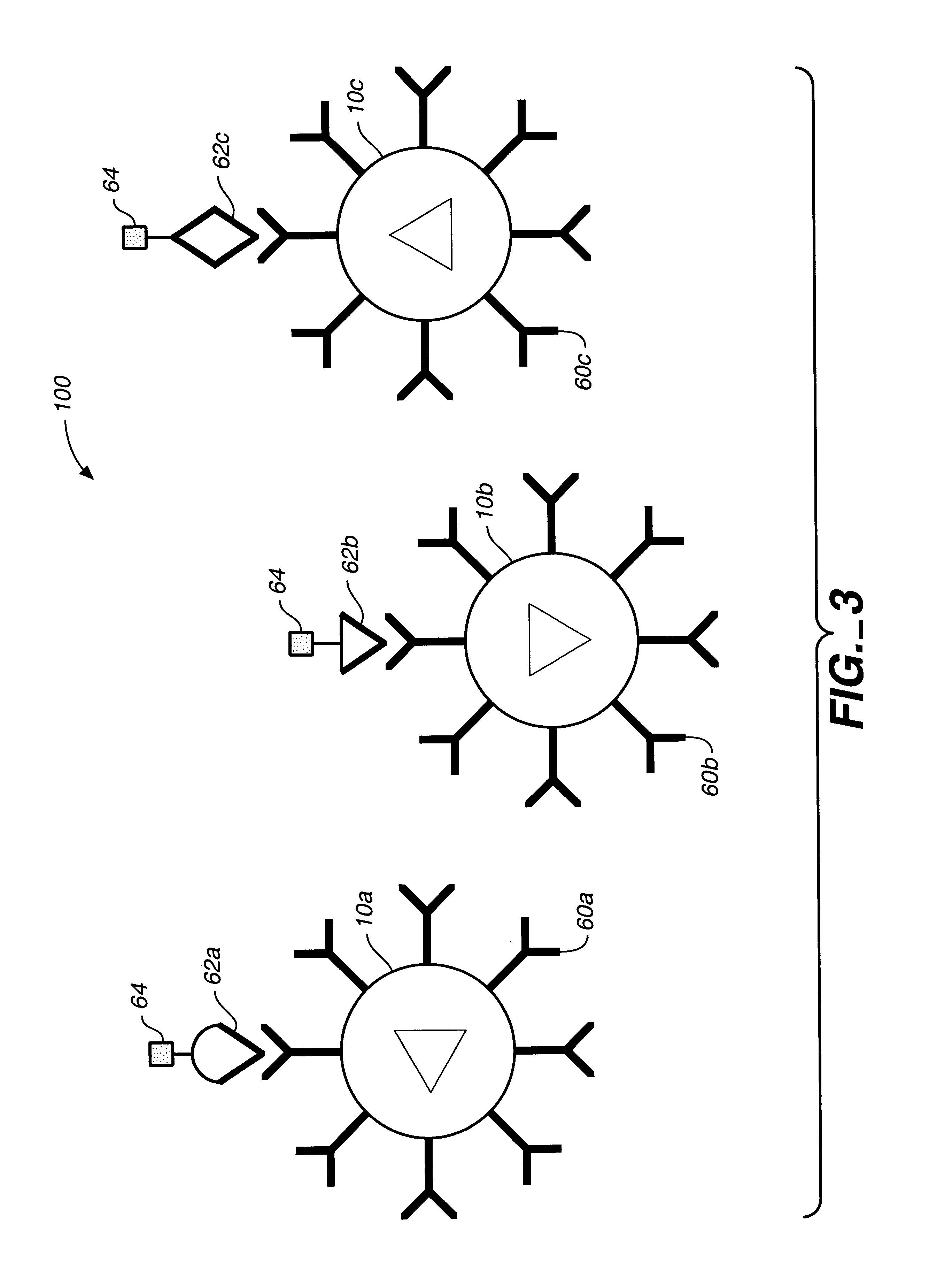
Target analyte sensors utilizing microspheres
A microsphere-based analytic chemistry system and method for making the same is disclosed in which microspheres or particles carrying bioactive agents may be combined randomly or in ordered fashion and dispersed on a substrate to form an array while maintaining the ability to identify the location of bioactive agents and particles within the array using an optically interrogatable, optical signature encoding scheme. A wide variety of modified substrates may be employed which provide either discrete or non-discrete sites for accommodating the microspheres in either random or patterned distributions. The substrates may be constructed from a variety of materials to form either two-dimensional or three-dimensional configurations. In a preferred embodiment, a modified fiber optic bundle or array is employed as a substrate to produce a high density array. The disclosed system and method have utility for detecting target analytes and screening large libraries of bioactive agents.
Owner:TRUSTEES OF TUFTS COLLEGETHE
Digital Counting of Individual Molecules by Stochastic Attachment of Diverse Labels
ActiveUS20110160078A1Highly precise relative and absolute counting statisticRaise countNucleotide librariesMicrobiological testing/measurementBiologyMultiple methods
Owner:BECTON DICKINSON & CO
Method for the complete chemical synthesis and assembly of genes and genomes
InactiveUS6521427B1BiocideSequential/parallel process reactionsChemical synthesisHuman genome database
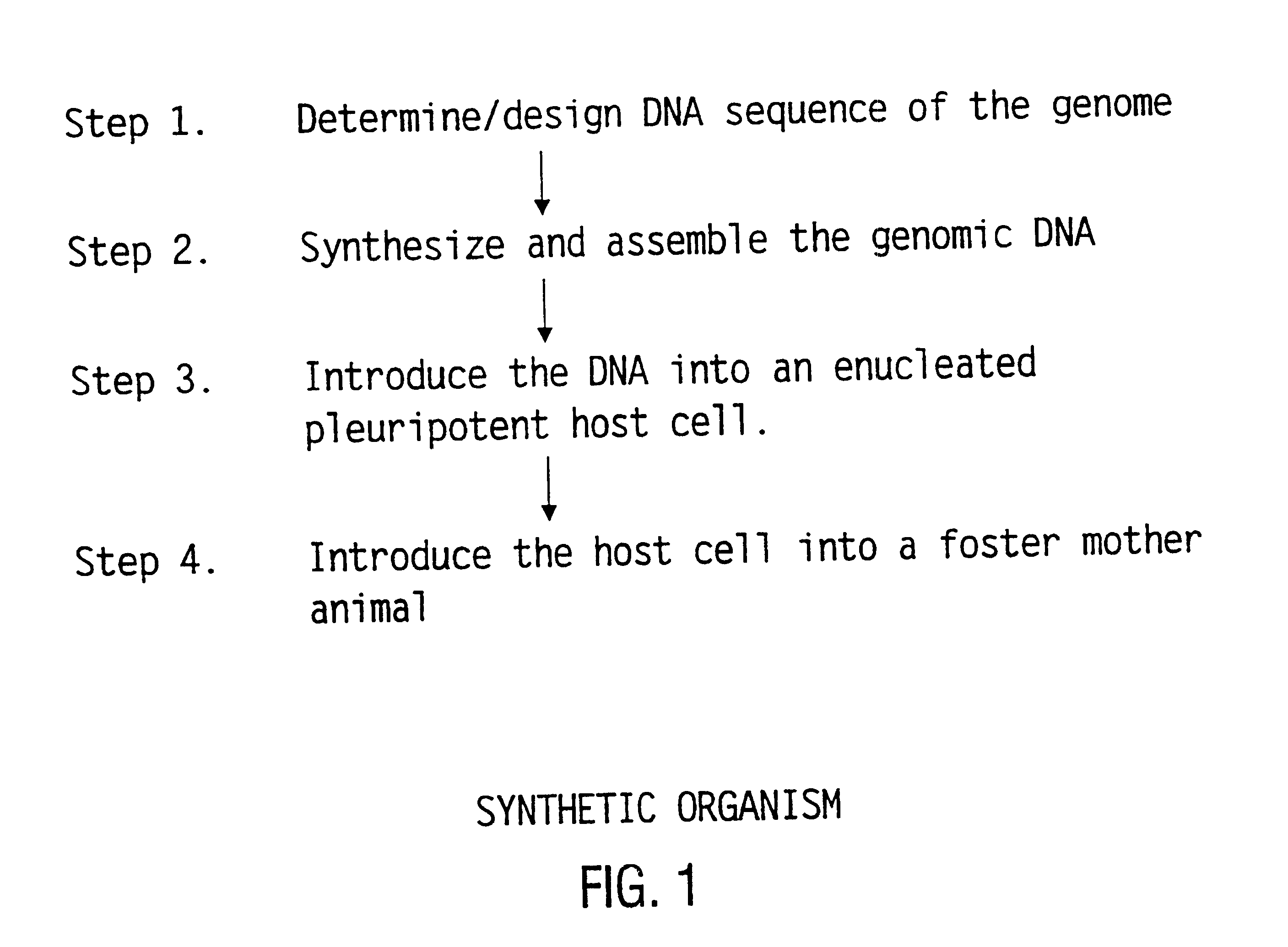
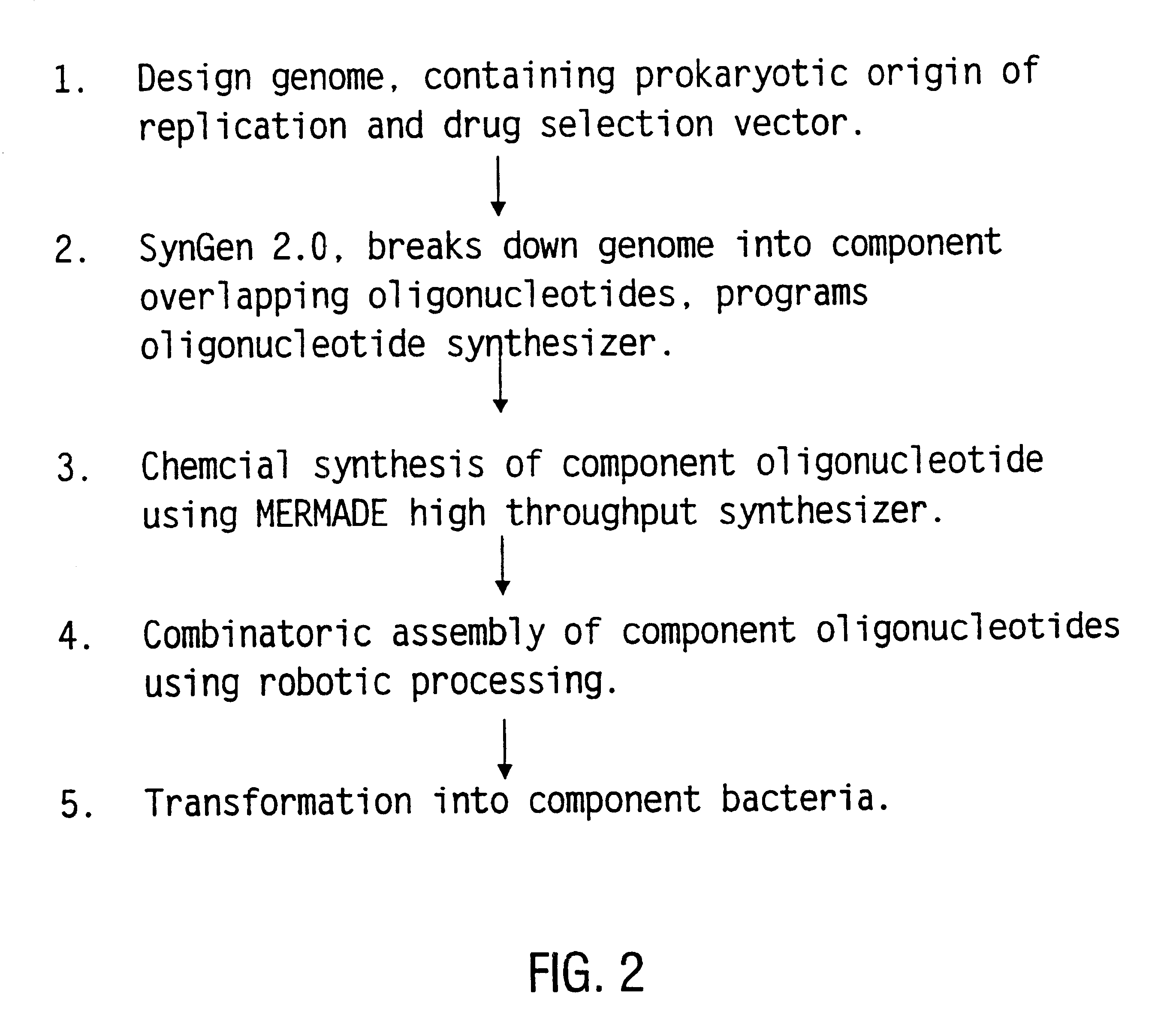
The present invention relates generally to the fields of oligonucleotide synthesis. More particularly, it concerns the assembly of genes and genomes of completely synthetic artificial organisms. Thus, the present invention outlines a novel approach to utilizing the results of genomic sequence information by computer directed gene synthesis based on computing on the human genome database. Specifically, the present invention contemplates and describes the chemical synthesis and resynthesis of genes defined by the genome sequence in a host vector and transfer and expression of these sequences into suitable hosts.
Owner:JOHNSON & JOHNSON INC (US) +3
Modified Molecular Arrays
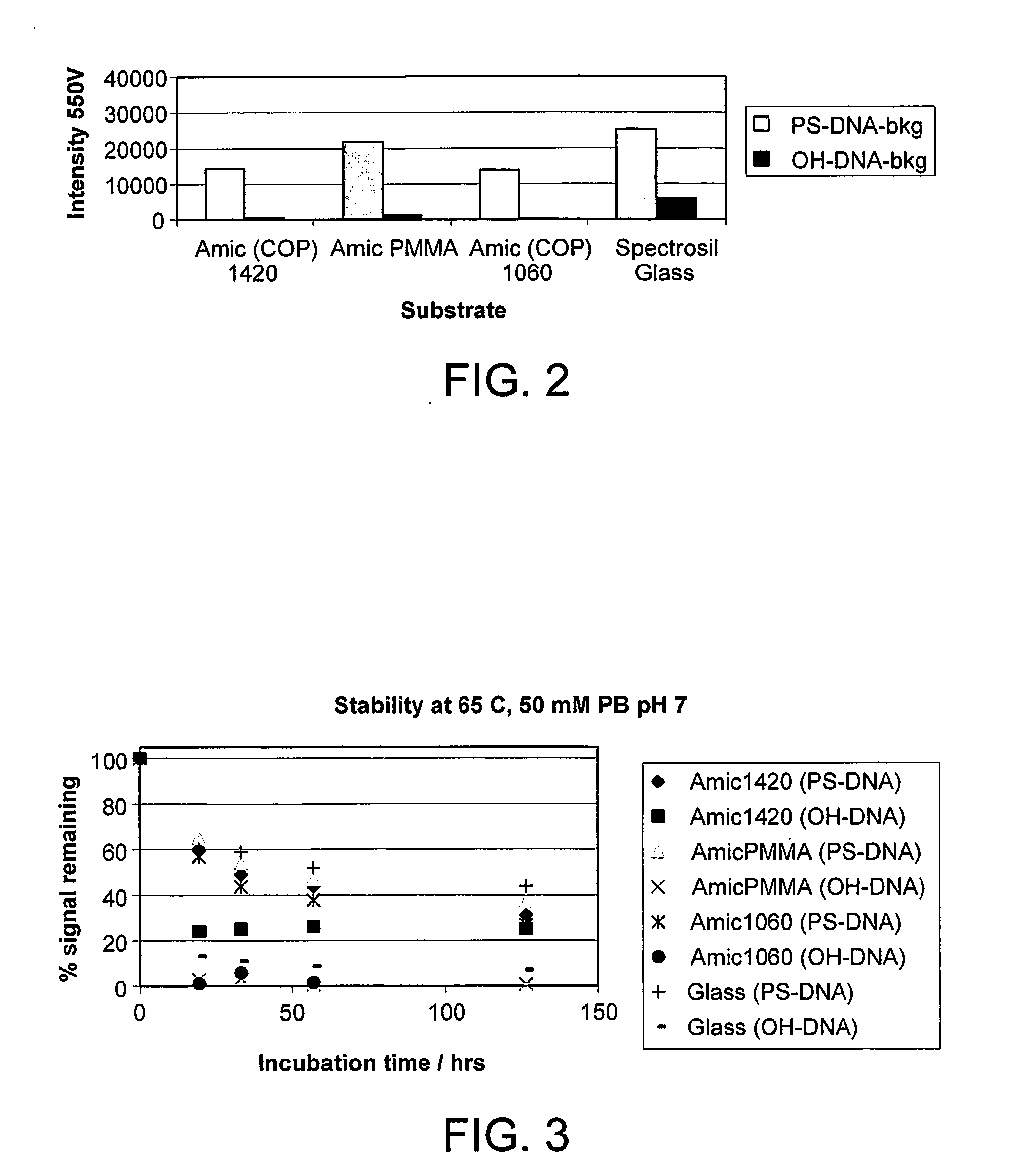
InactiveUS20110059865A1Less reactiveAccelerated programSequential/parallel process reactionsNucleotide librariesMolecular array(Hydroxyethyl)methacrylate
The invention relates to the preparation of a hydrogel surface useful in the formation and manipulation of arrays of molecules, particularly polynucleotides and to the chemical modification of these and other arrays. In particular, the invention relates to a method of preparing a hydrogel immobilised to a solid support comprising polymerising on the support a mixture of a first comonomer which is acrylamide, methacrylamide, hydroxyethyl methacrylate or N-vinyl pyrrolidinone and a second comonomer which is a functionalised acrylamide or acrylate.
Owner:ILLUMINA CAMBRIDGE LTD
Microfluidic Devices and Methods of Use in The Formation and Control of Nanoreactors
InactiveUS20100137163A1Material nanotechnologyCompound screeningHigh-Throughput Screening AssaysEmulsion
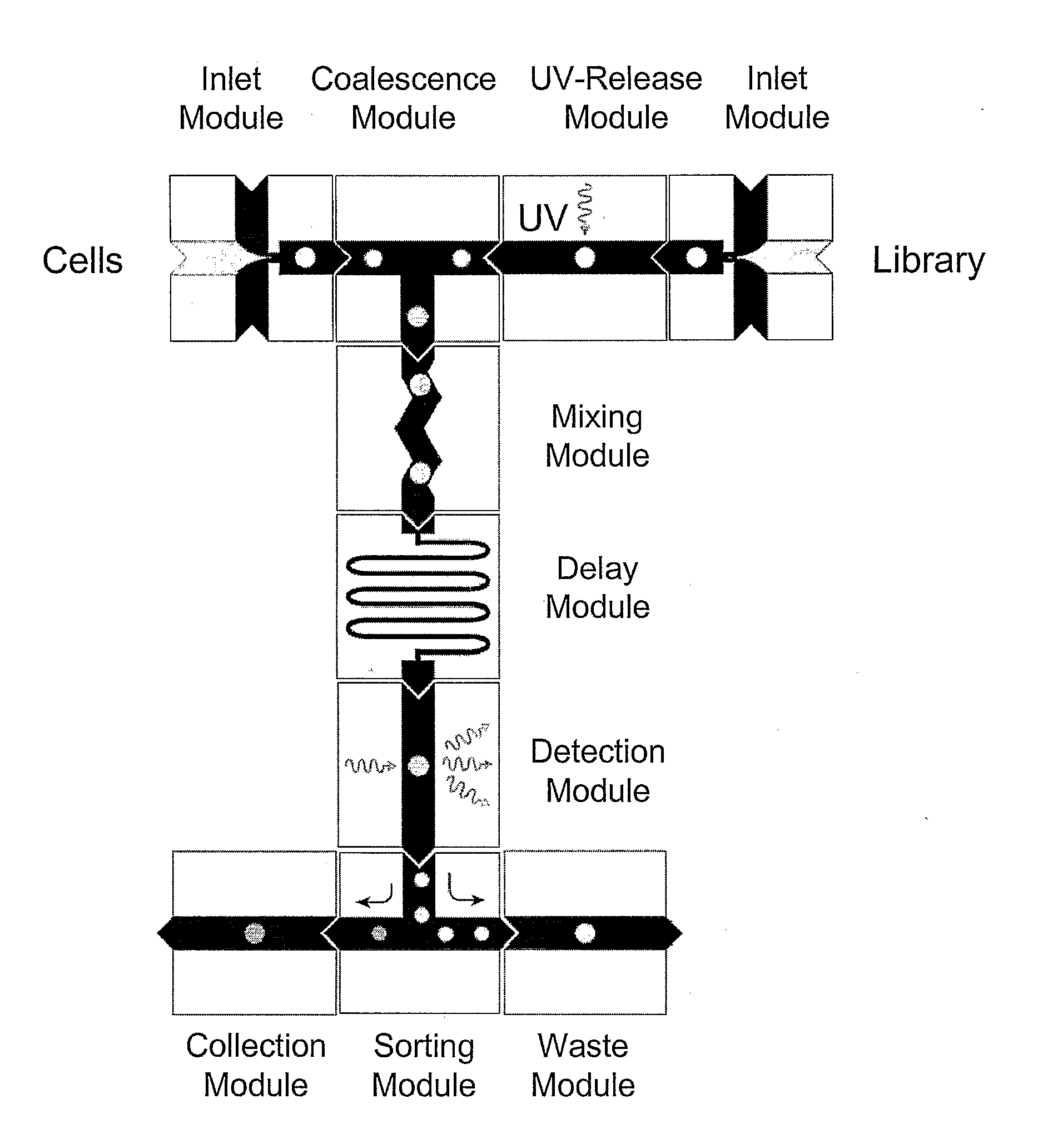
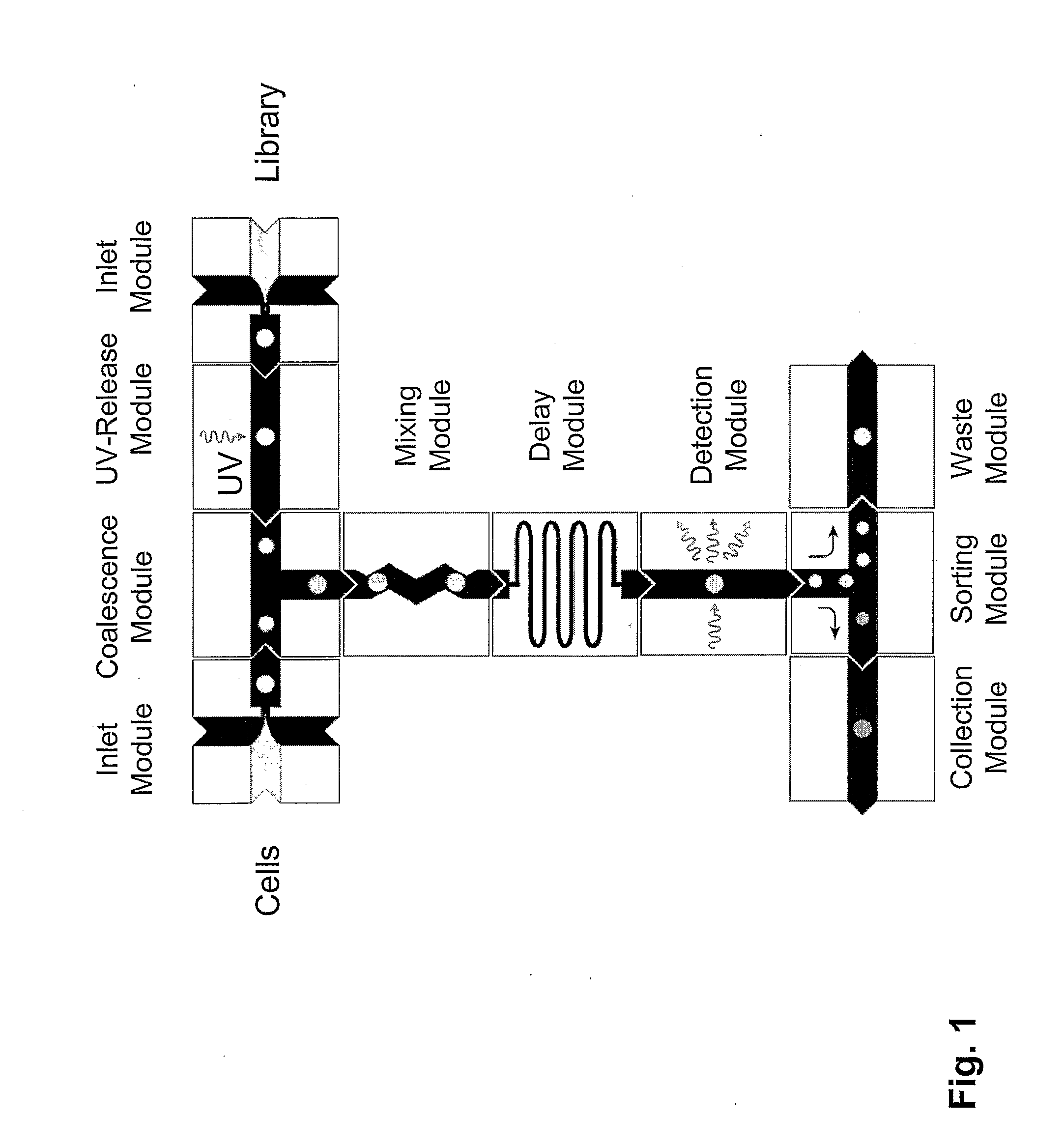
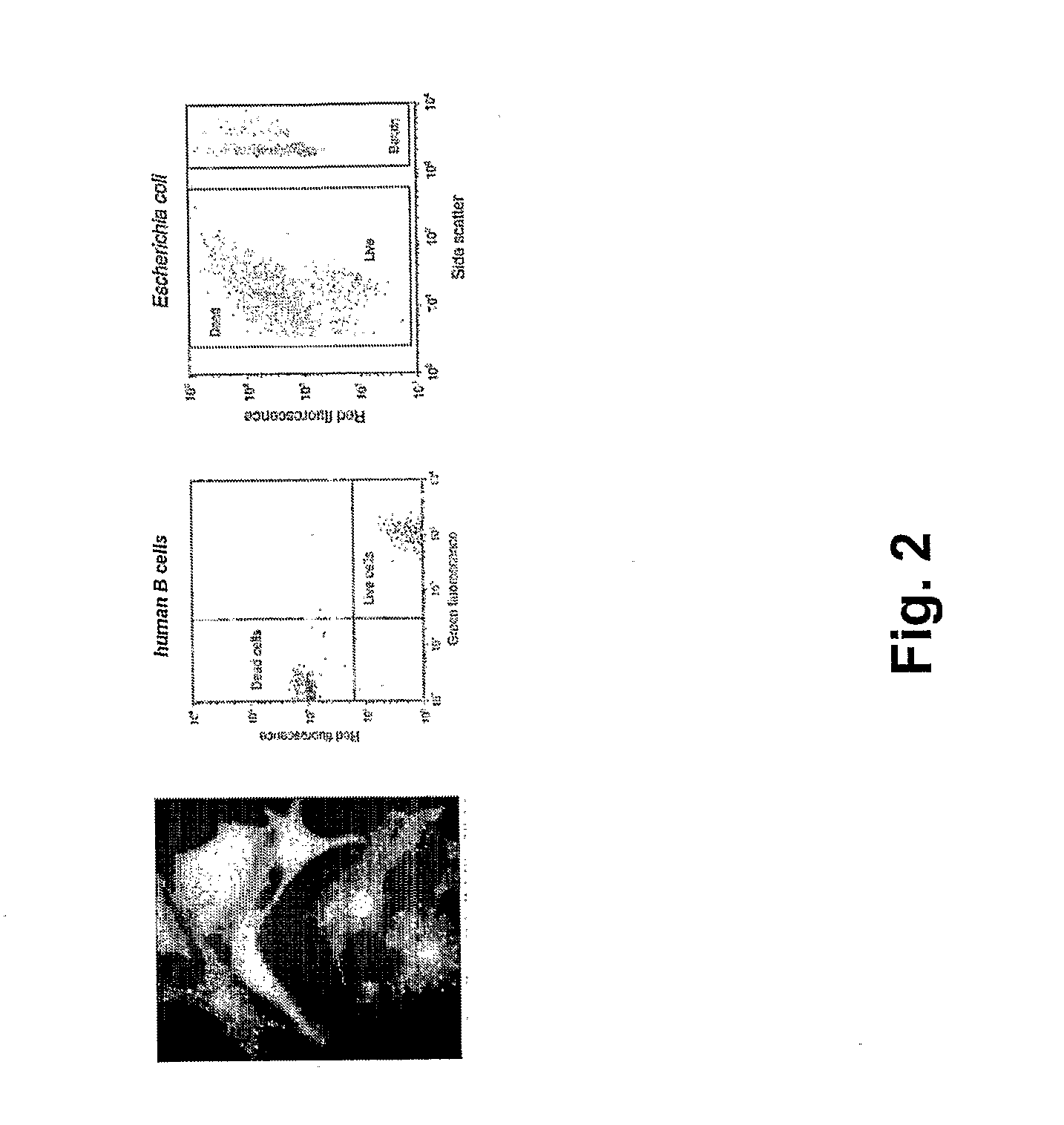
The present invention provides novel microfluidic devices and methods that are useful for performing high-throughput screening assays and combinatorial chemistry. Such methods can include labeling a library of compounds by emulsifying aqueous solutions of the compounds and aqueous solutions of unique liquid labels on a microfluidic device, which includes a plurality of electrically addressable, channel bearing fluidic modules integrally arranged on a microfabricated substrate such that a continuous channel is provided for flow of immiscible fluids, whereby each compound is labeled with a unique liquid label, pooling the labeled emulsions, coalescing the labeled emulsions with emulsions containing a specific cell or enzyme, thereby forming a nanoreactor, screening the nanoreactors for a desirable reaction between the contents of the nanoreactor, and decoding the liquid label, thereby identifying a single compound from a library of compounds.
Owner:BIO RAD LAB INC
Arrayed biomolecules and their use in sequencing
InactiveUS7232656B2Reduce interferencePermit resolutionBioreactor/fermenter combinationsSequential/parallel process reactionsGenome variationComputational biology
The invention is directed to a method for analysing genome wide variation in an individual. The method comprises randomly fragmenting the individual's genome and generating sequence reads of multiple bases on all fragments of the individual's genome, aligning the sequence reads generated with a known genomic reference sequence, and analysing variations between the sequence reads derived from the individual's genome and the known genomic reference sequence.
Owner:ILLUMINA CAMBRIDGE LTD
Nucleic acid amplification with continuous flow emulsion
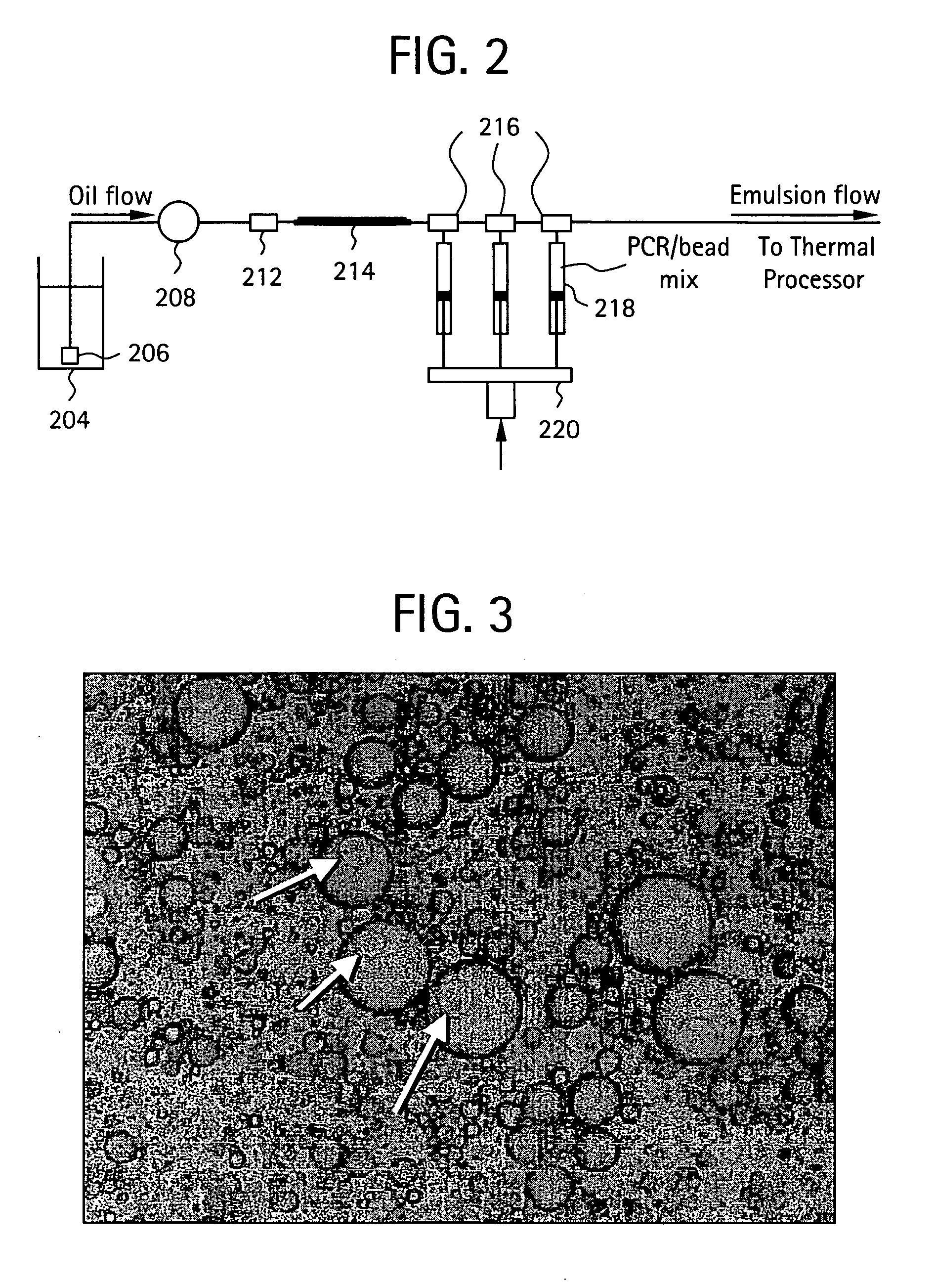
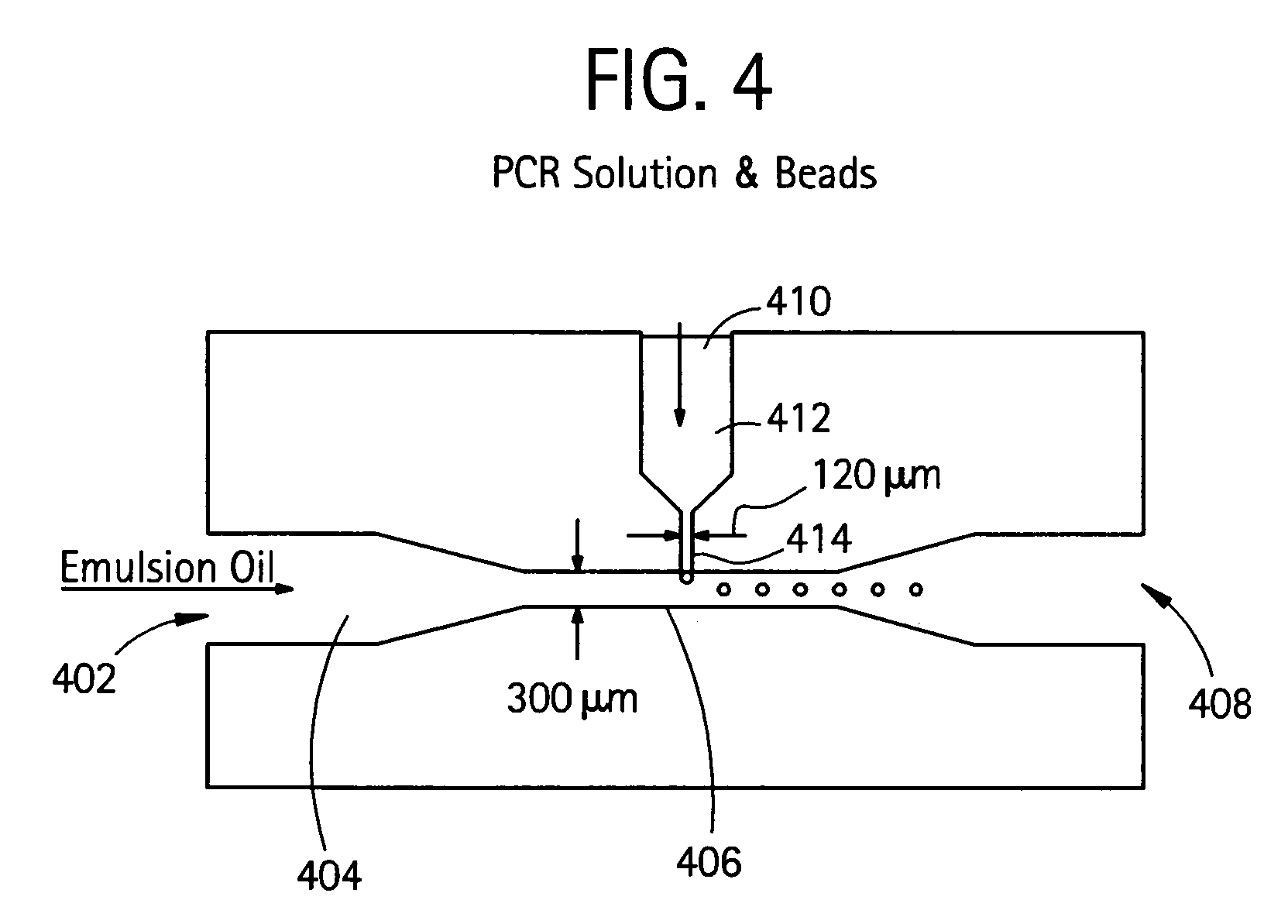
InactiveUS20050227264A1Rapid and economical mannerReduce nozzle cloggingHeating or cooling apparatusFlow mixersMicroreactorGenetic Materials
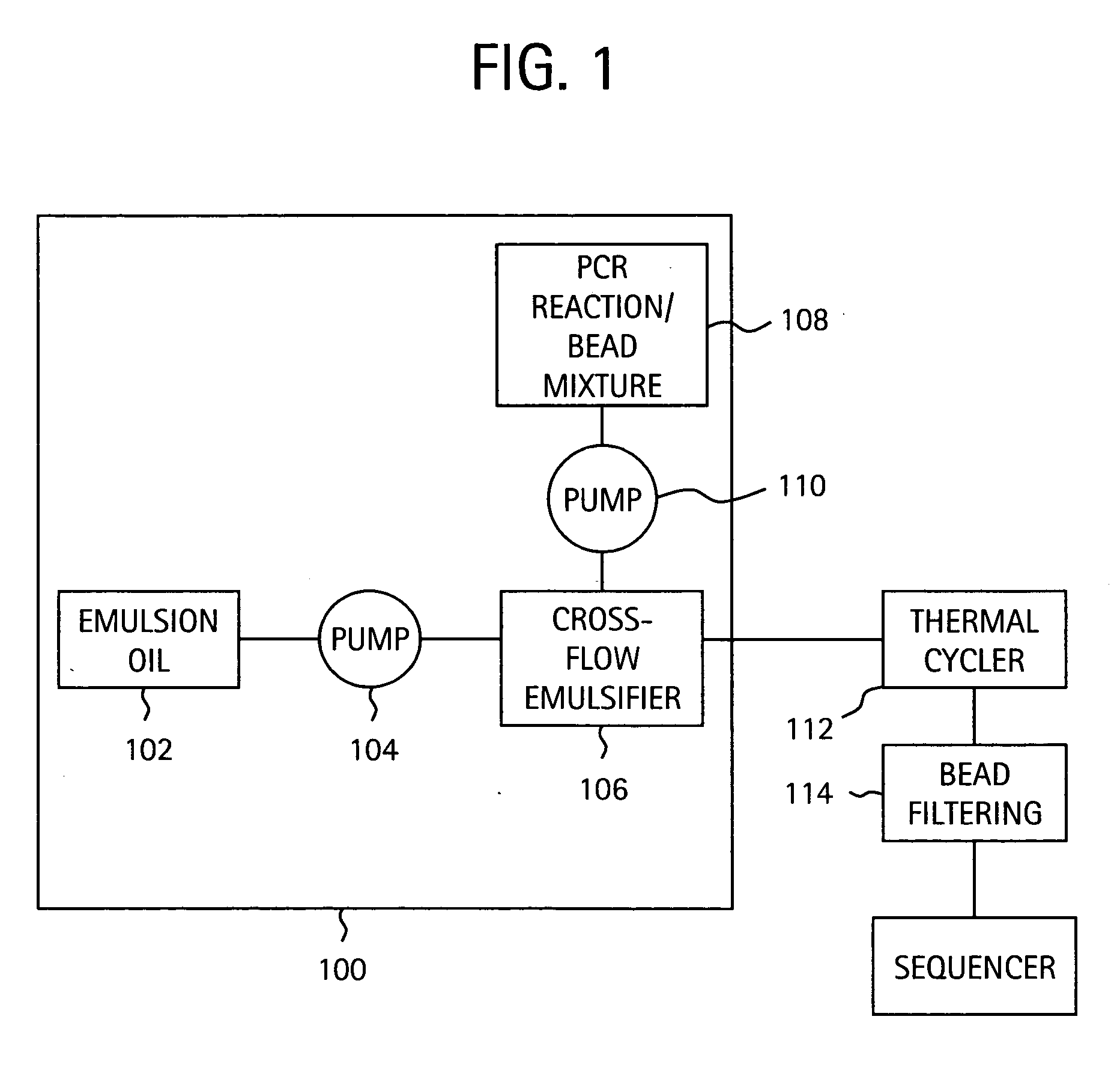
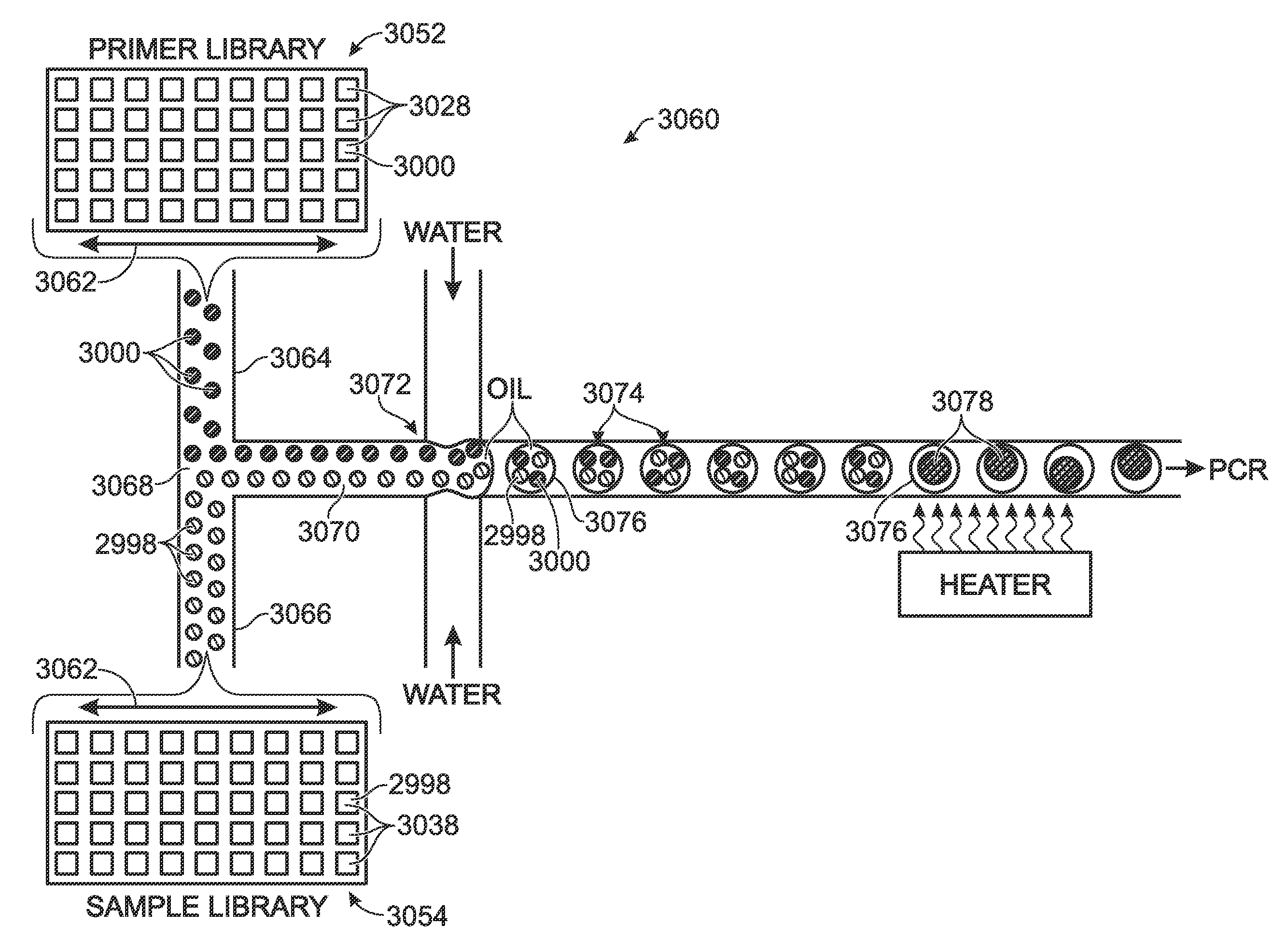
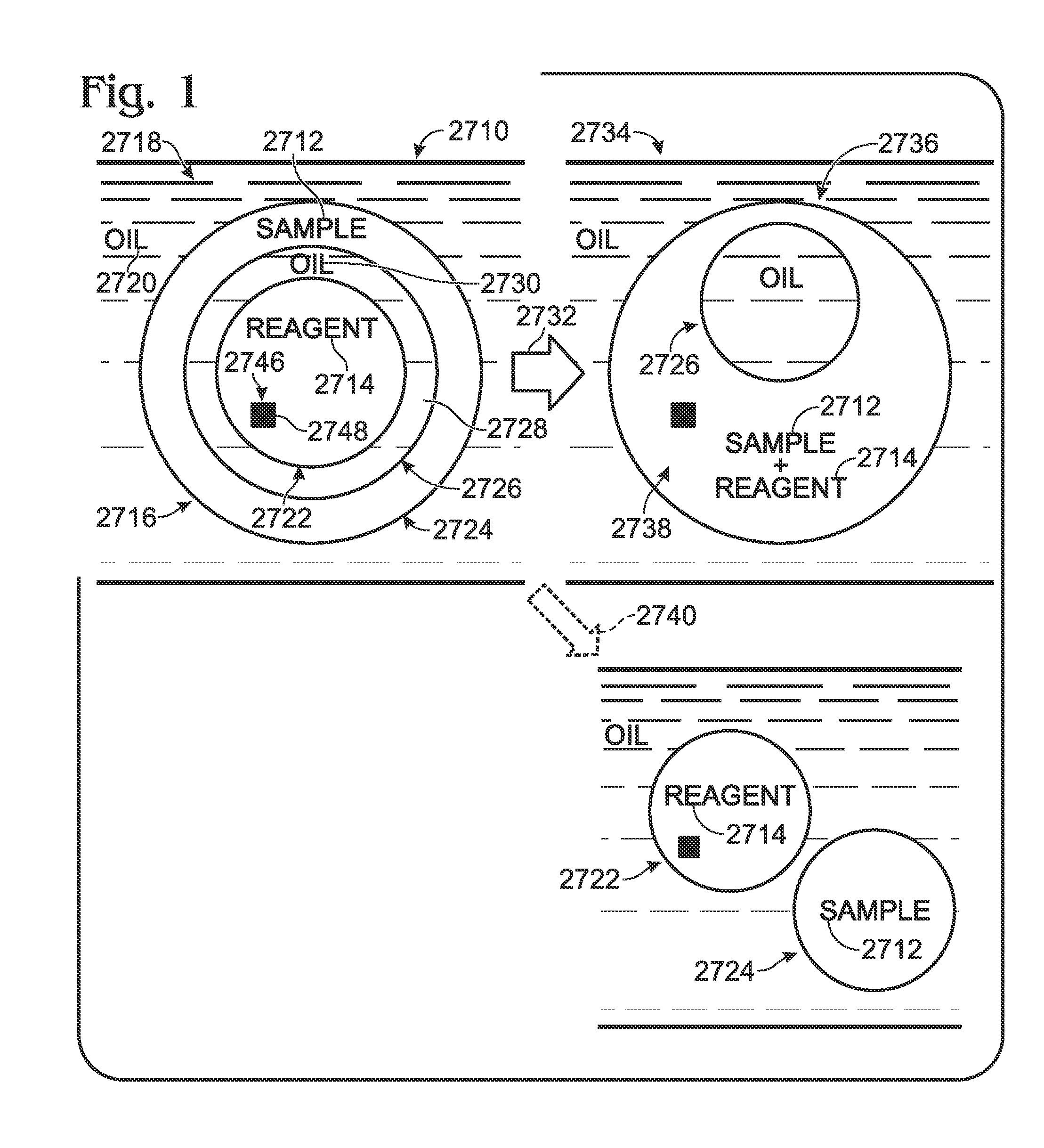
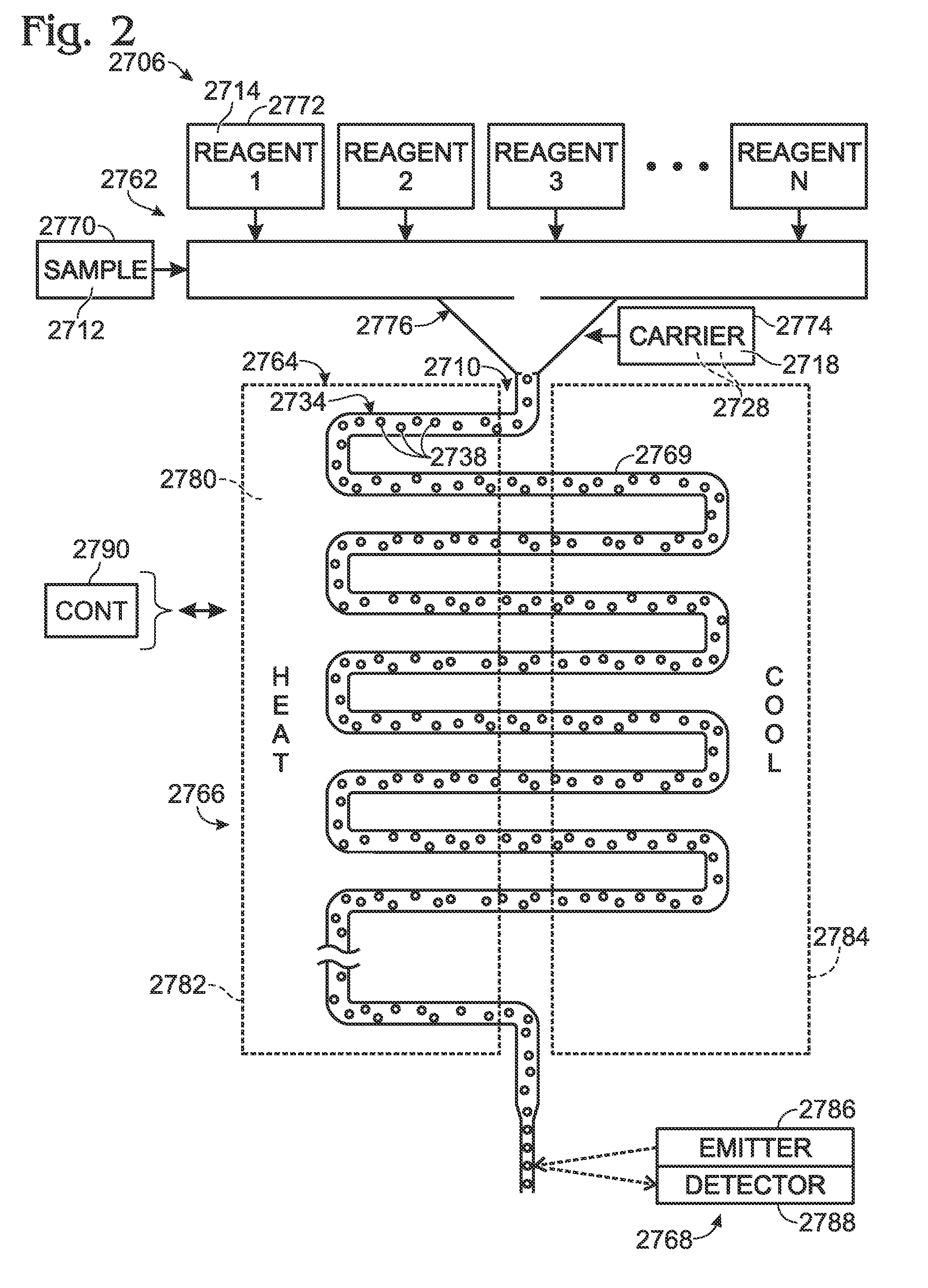
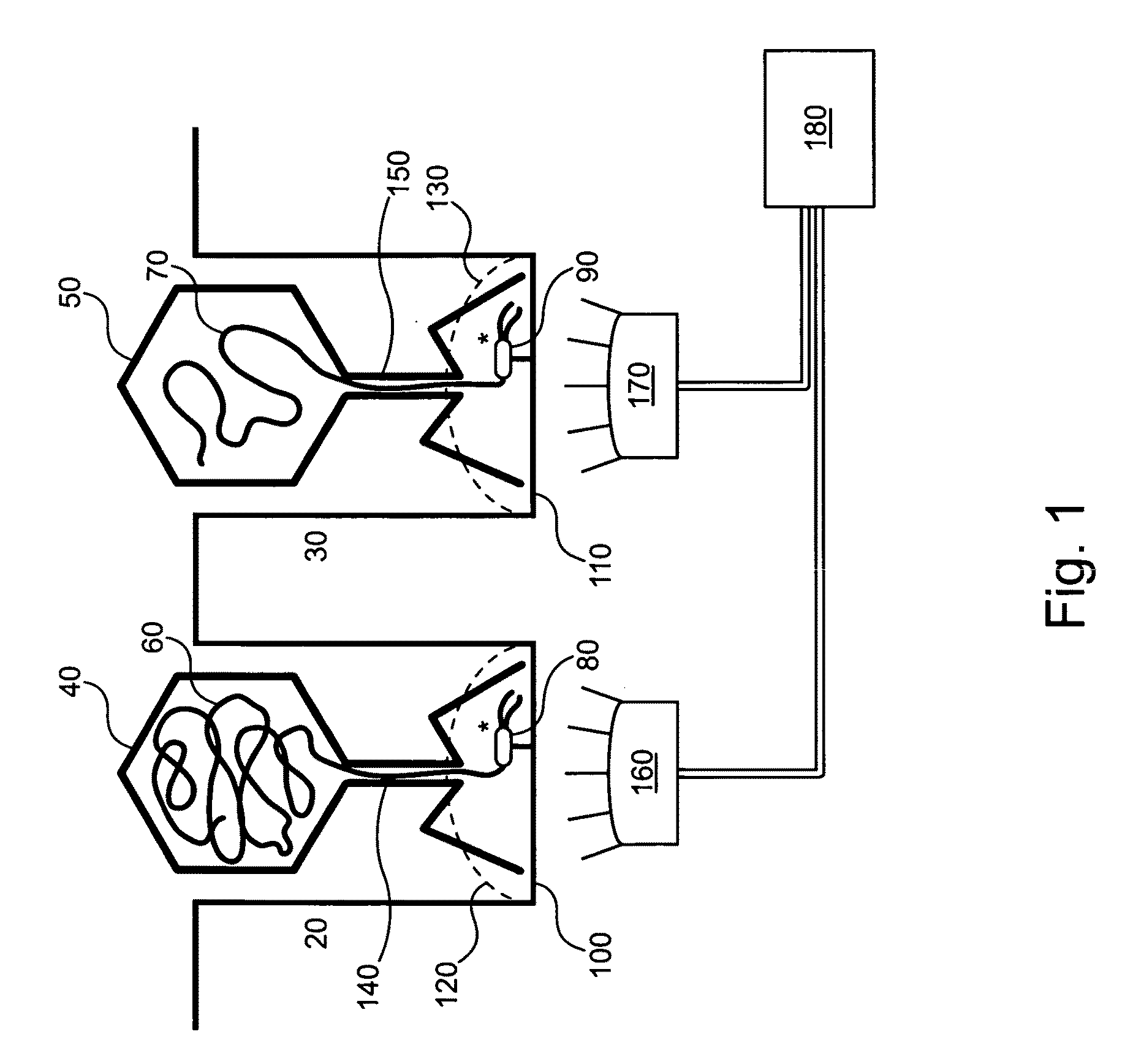
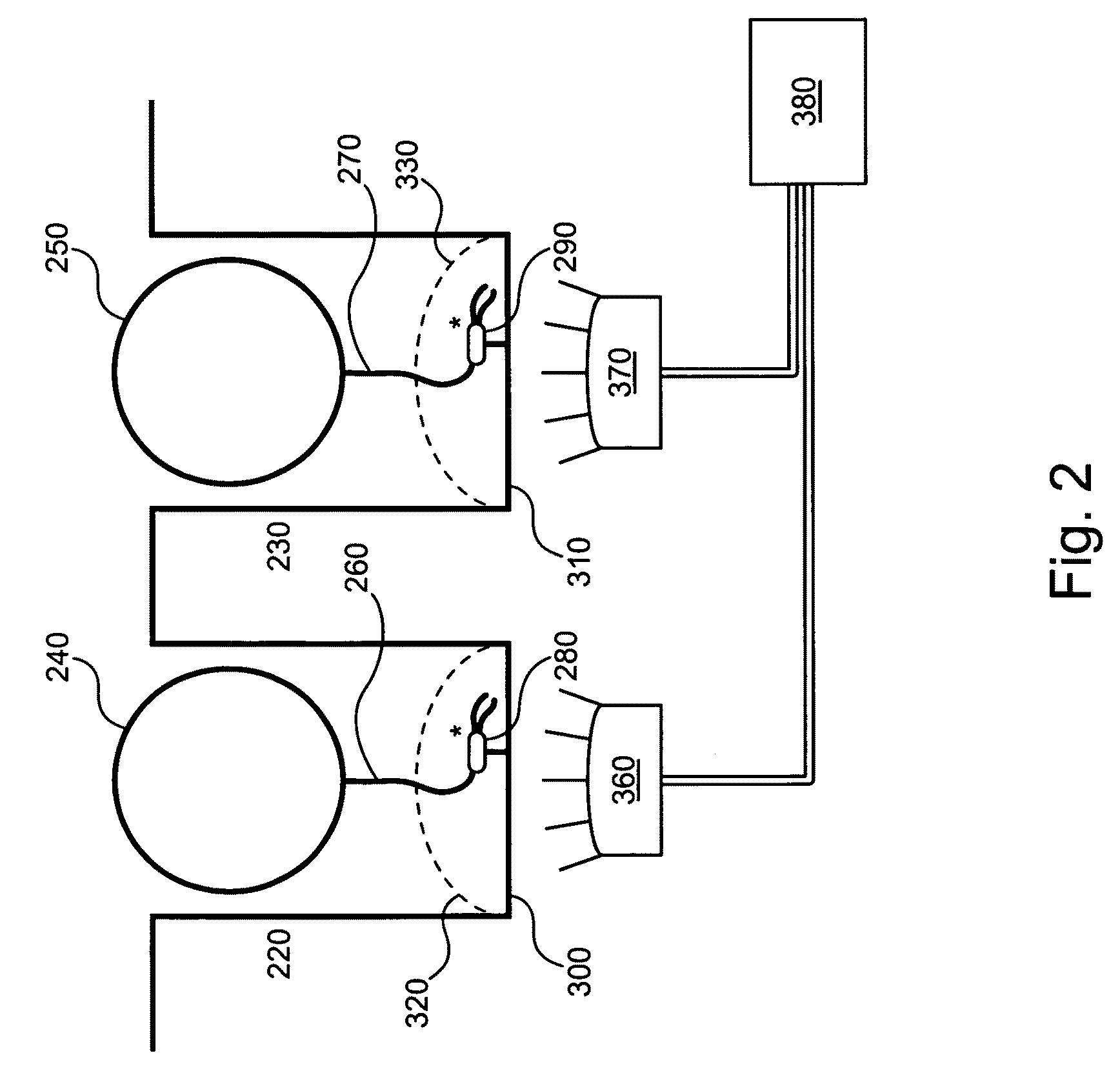
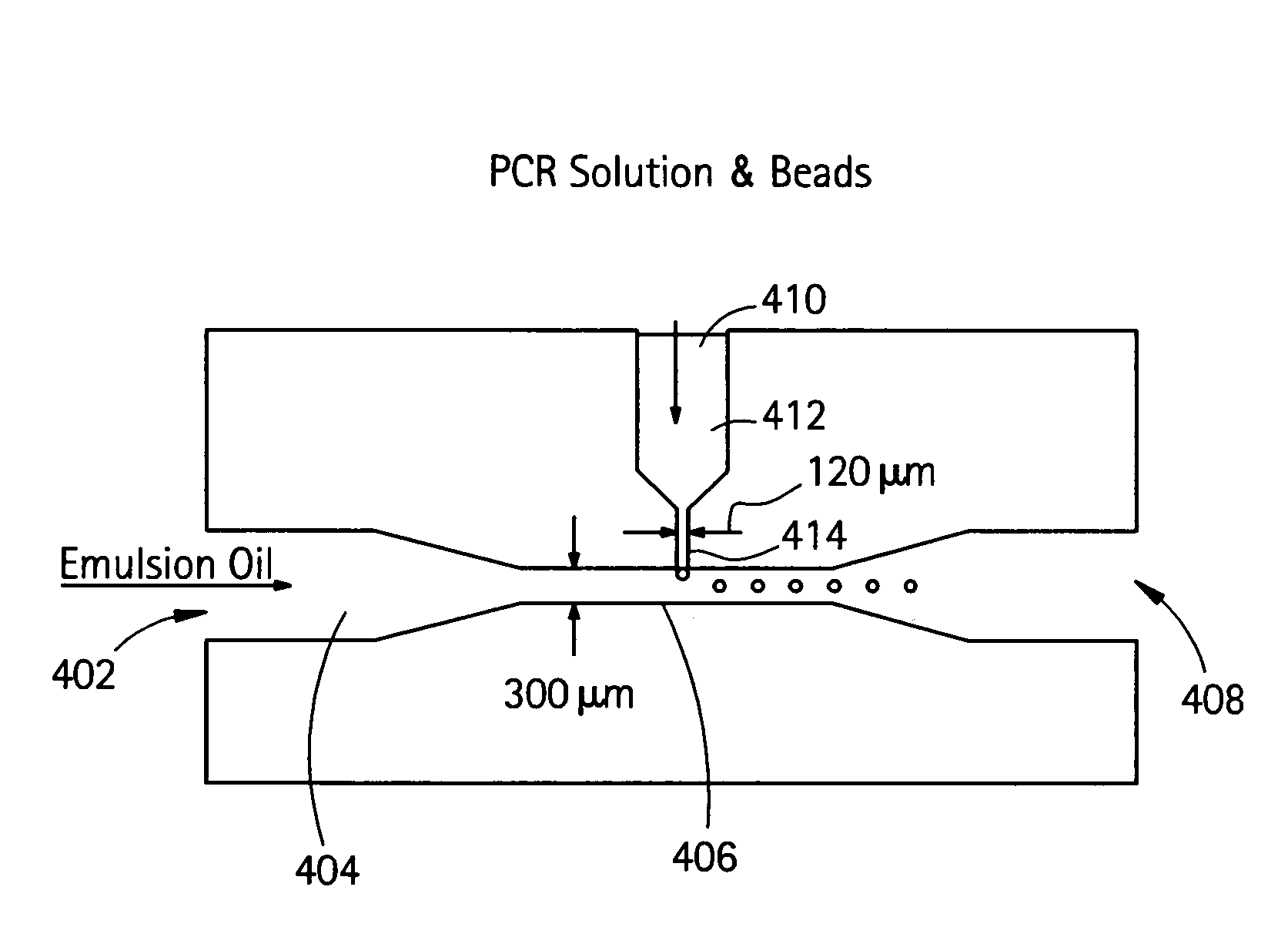
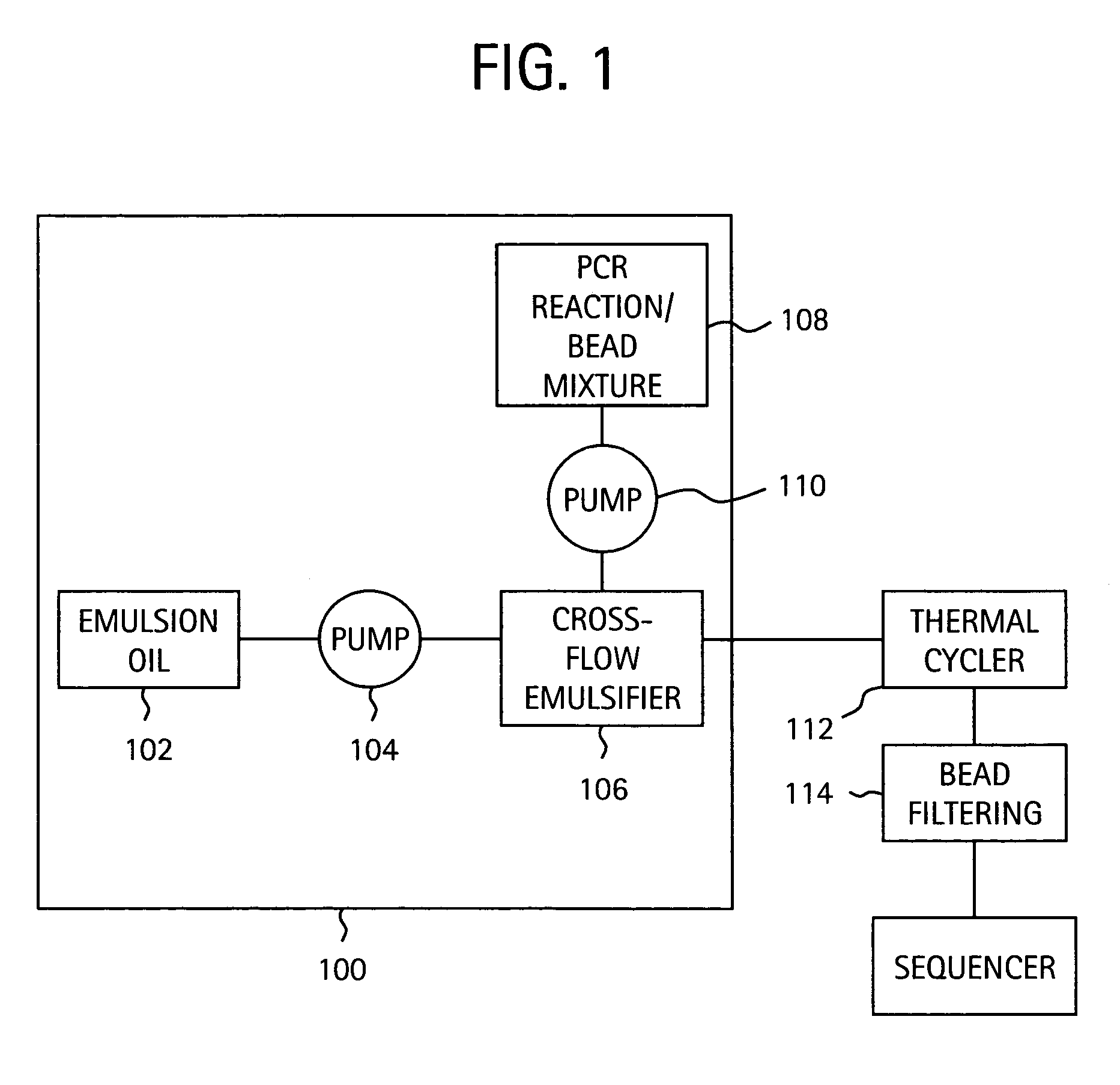
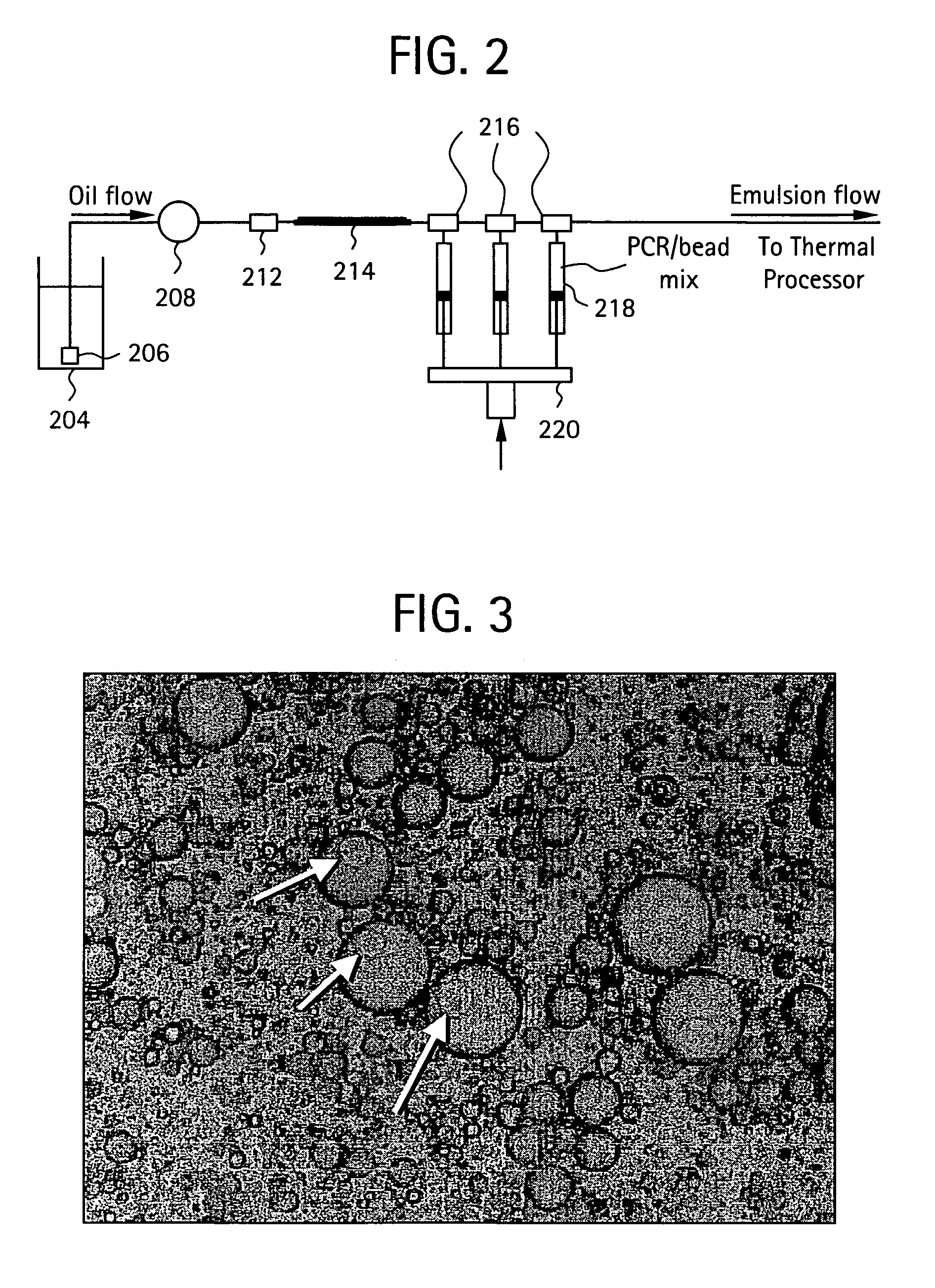
Embodiments of the present invention are directed to methods and devices / systems for amplifying genetic material and may include providing a water-in-oil emulsion in a continuous flow. The emulsion may include a plurality of water droplets comprising microreactors. Each of the plurality of microreactors may include a single bead capable of capturing a nucleic acid template, a single species nucleic acid template and sufficient reagents to amplify the copy number of the nucleic acid template. The method also includes flowing the emulsion across a first temperature zone and a second lower temperature zone to thermally process the microreactors to amplify the nucleic acid template by polymerase chain reaction.
Owner:454 LIFE SCIENCES CORP
DNA-bridged carbon nanotube arrays
InactiveUS20020172963A1High precisionHigh sensitivityBioreactor/fermenter combinationsMaterial nanotechnologyChemical ligationElectron transfer reactions
A class of biological sensing devices that include a substrate comprising an array of carbon nanotubes (CNTs) to which are chemically attached biological molecules is disclosed. The attached biological molecules are capable of electrical conductivity that is responsive to chemical changes occurring as a result of their interaction with target species. A means for means for using DNA as a material of potential in molecular electronic sensor devices, being primarily based on molecular electron-transfer reaction processes between DNA-binding donors and acceptors is also disclosed, including composition, method of manufacture and their use are described.
Owner:TRUSTEES OF BOSTON COLLEGE THE
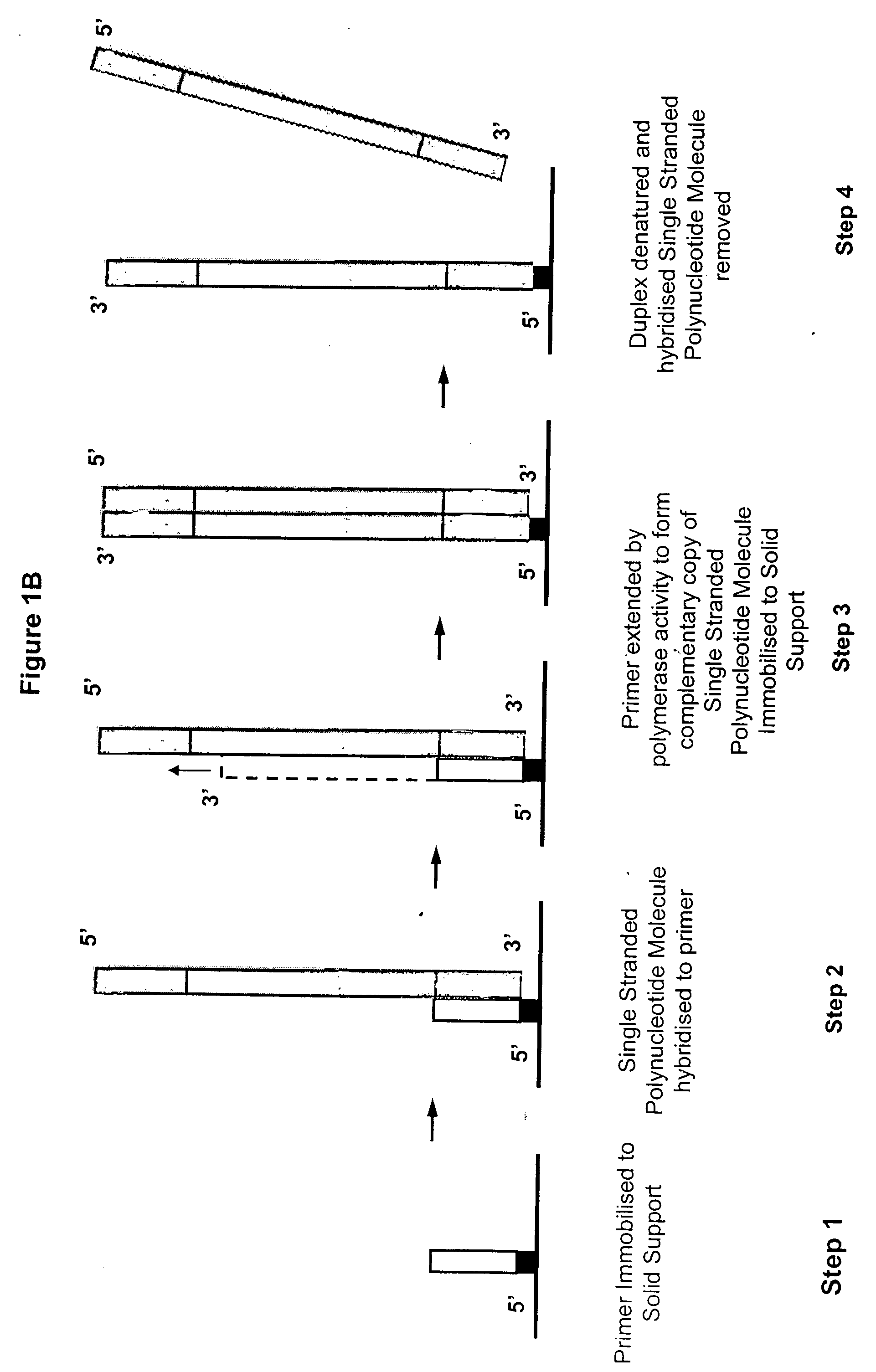
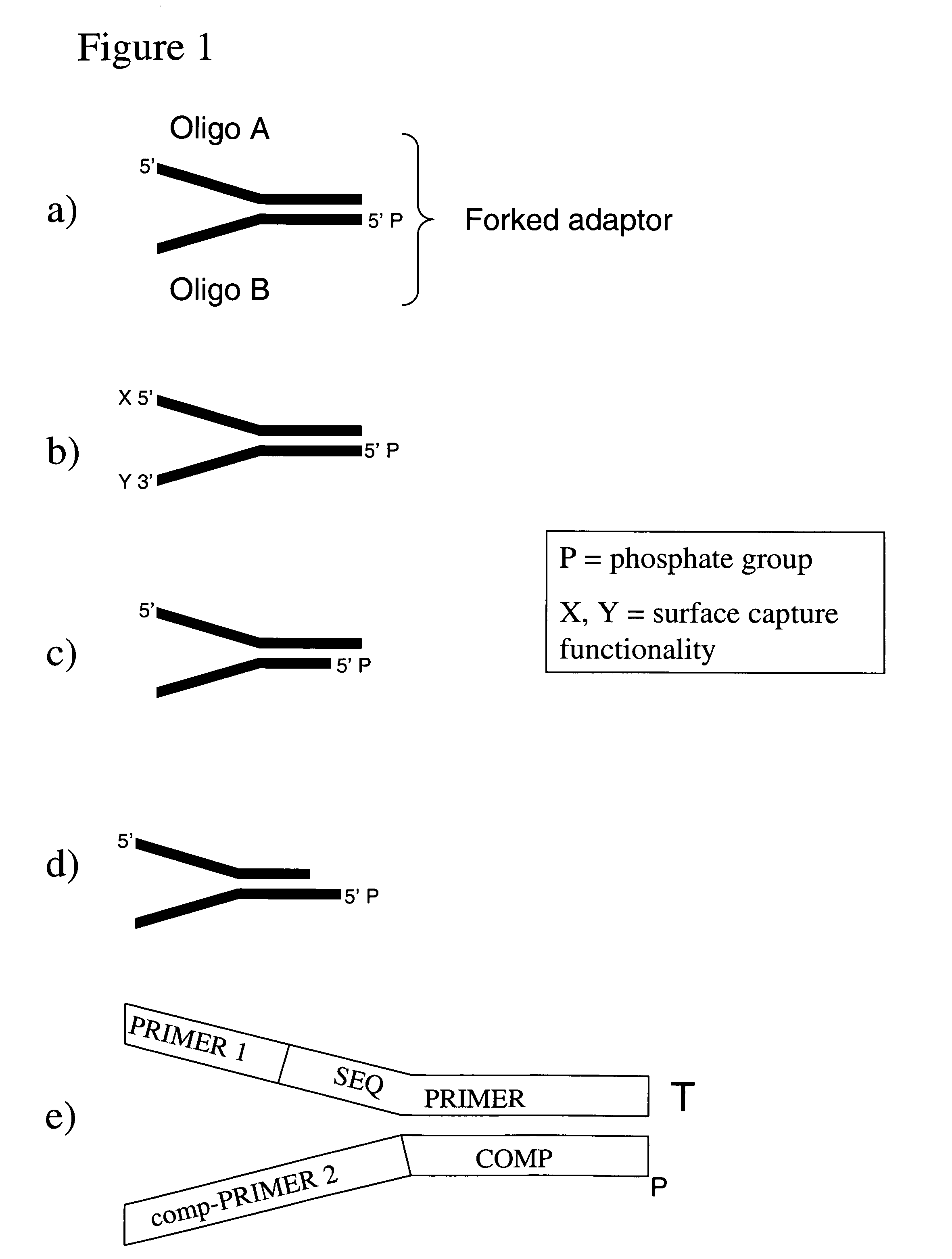
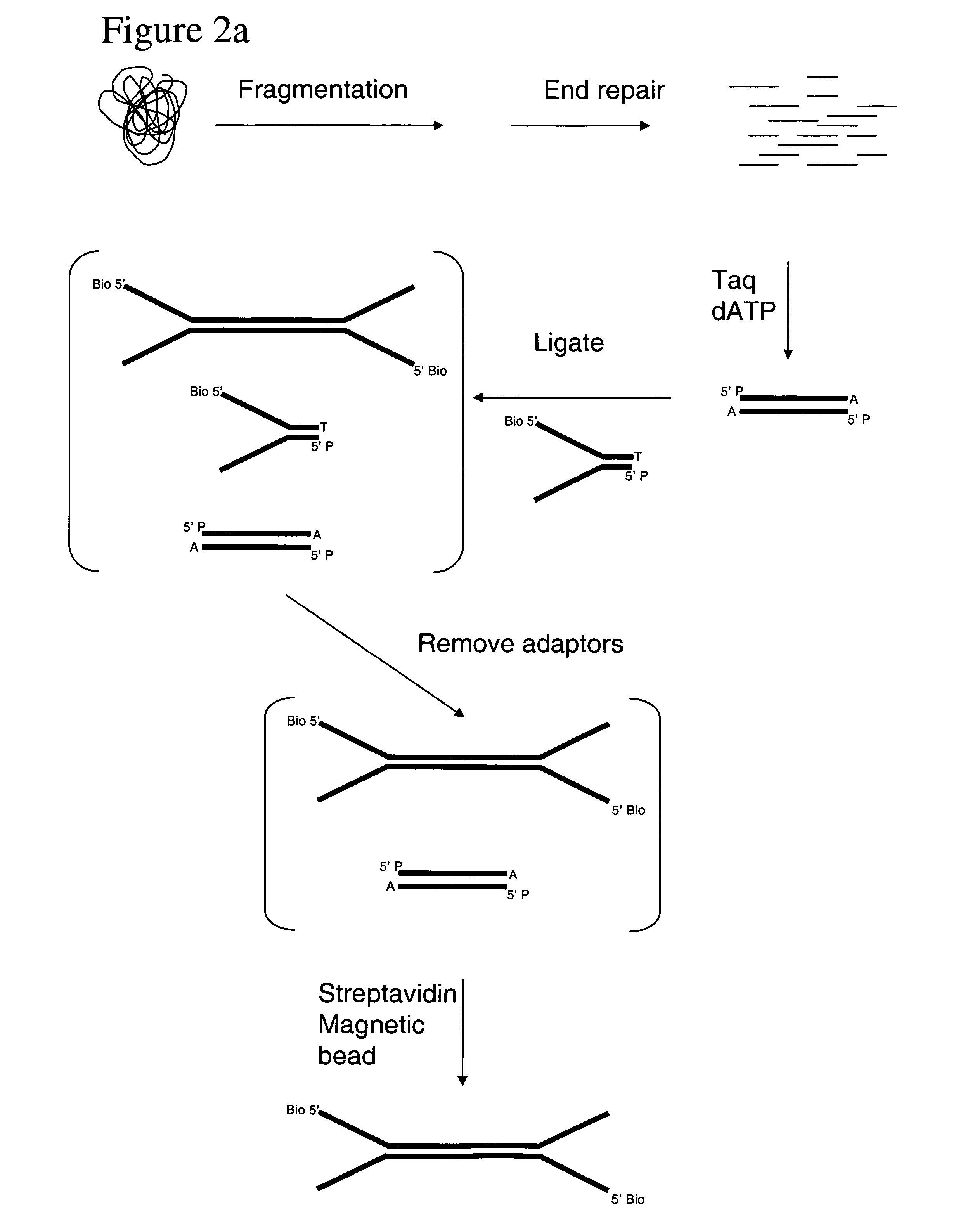
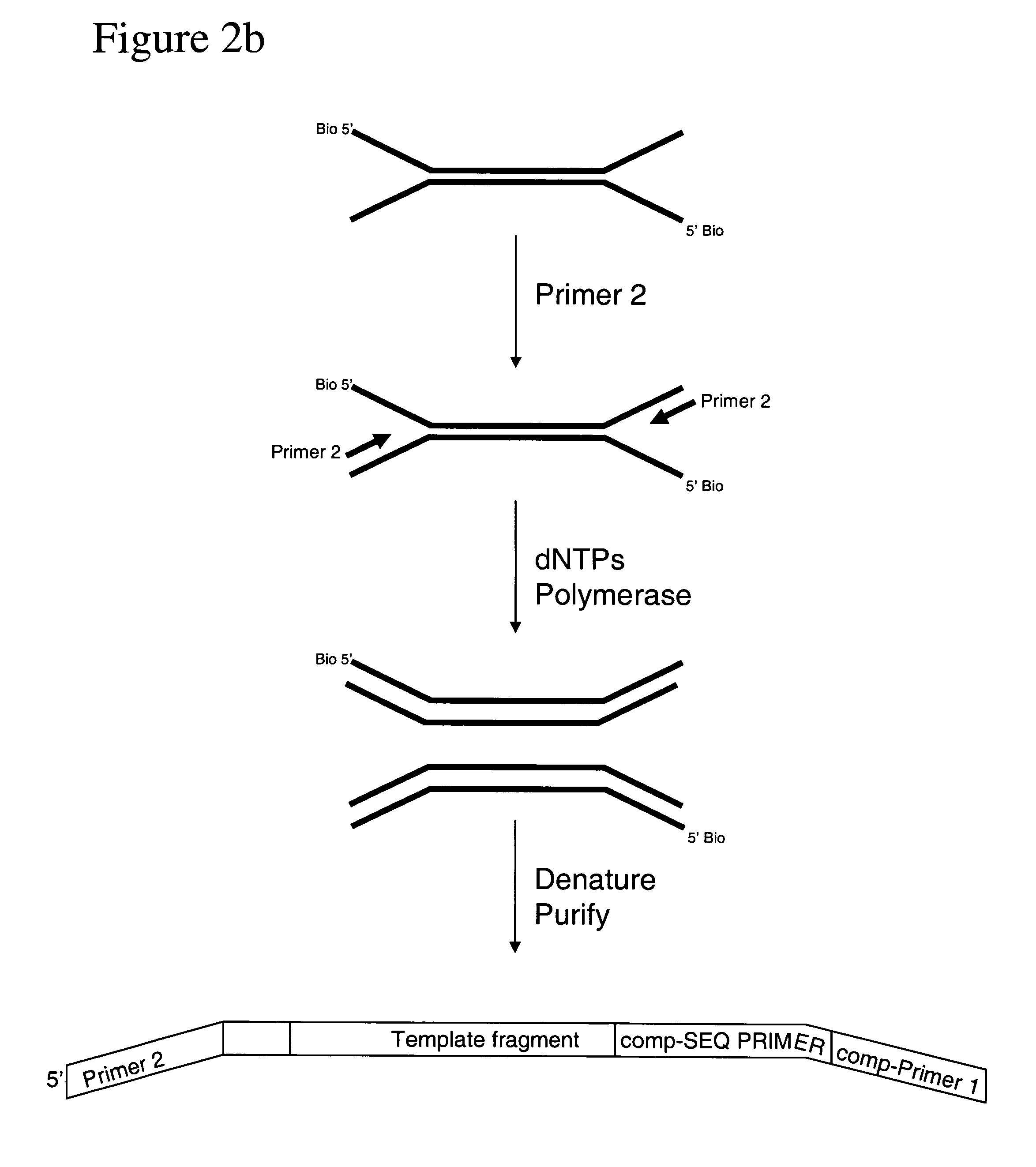
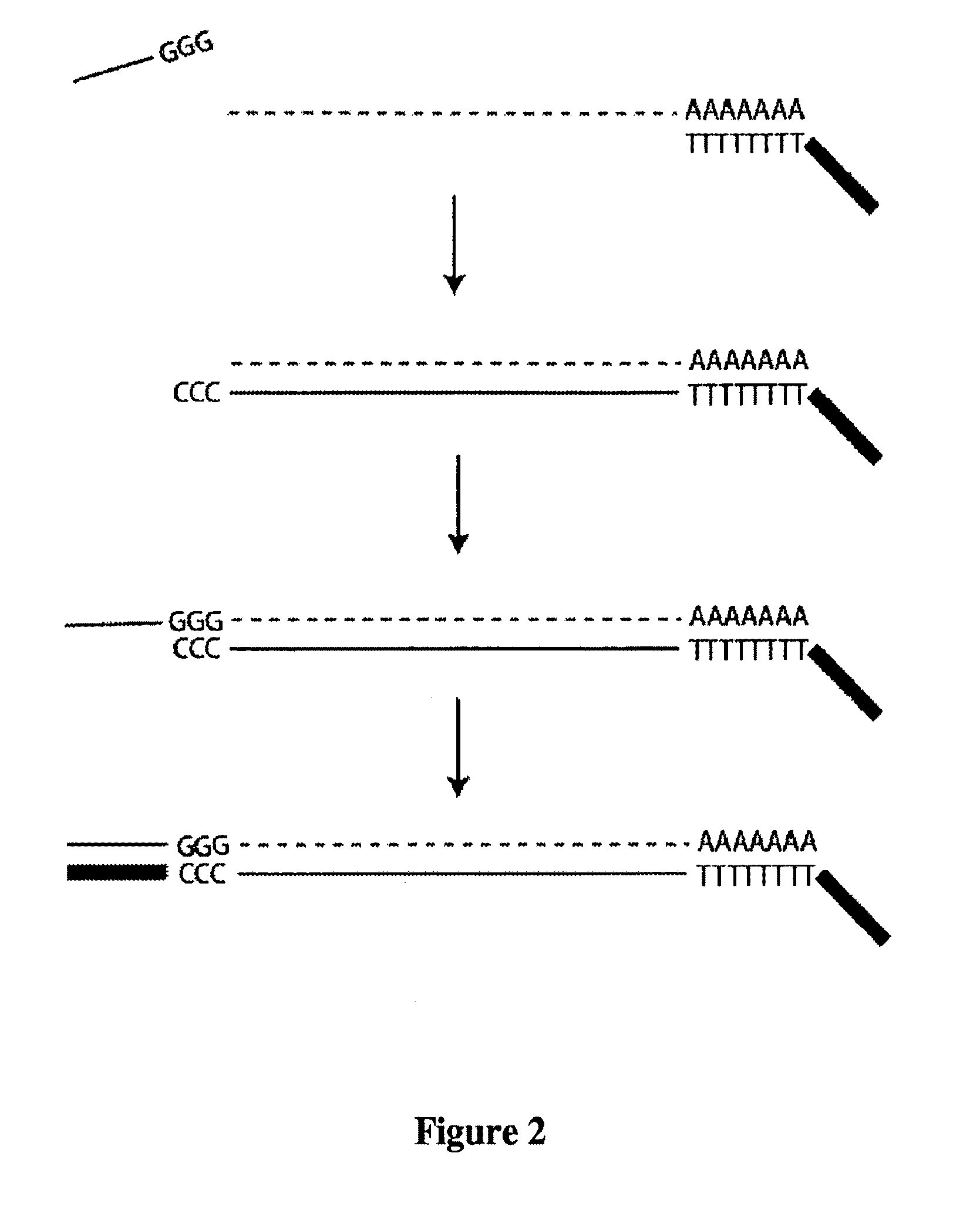
Method of preparing libraries of template polynucleotides
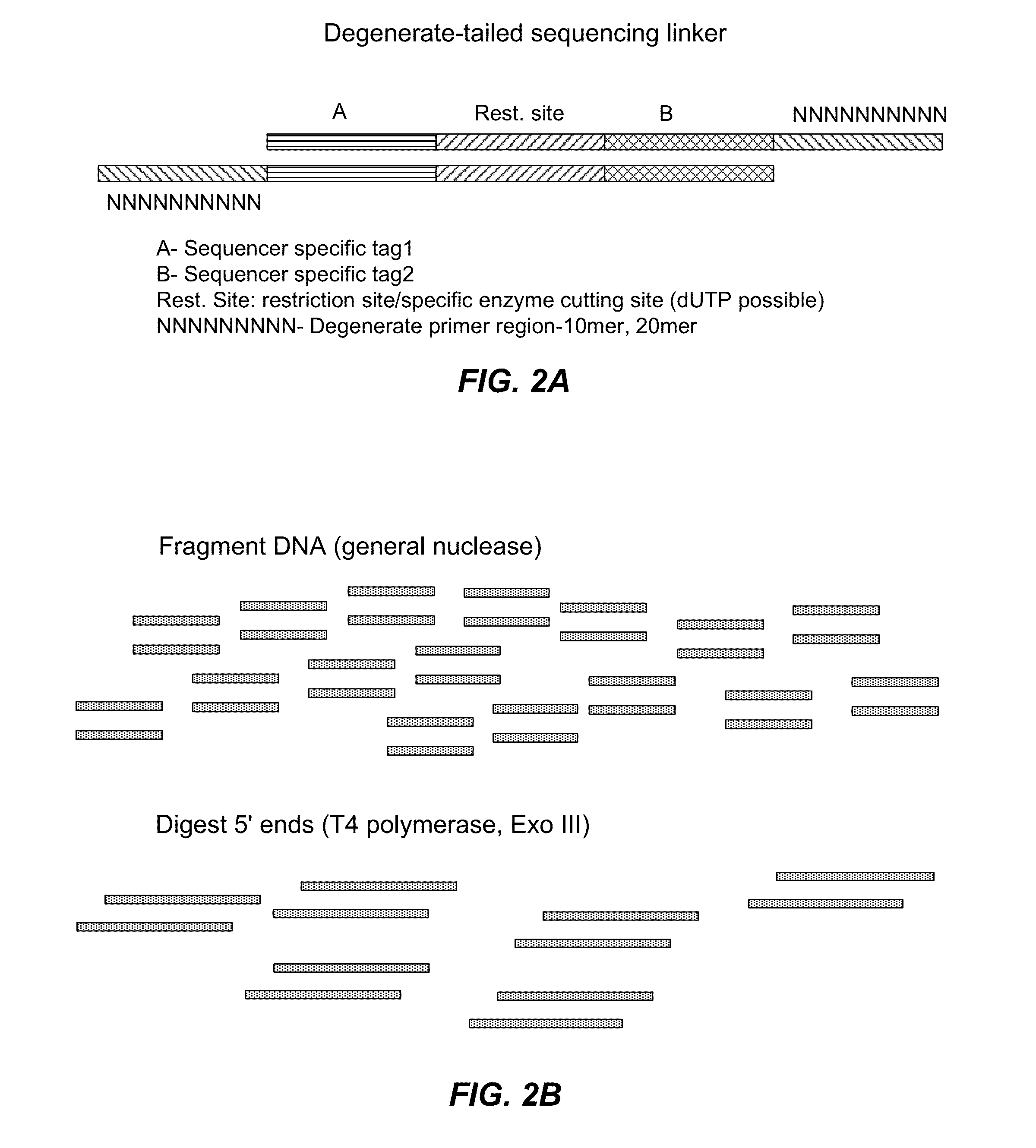
The present invention relates to a method for preparing a library of template polynucleotides and use thereof in methods of solid-phase nucleic acid amplification. More specifically, the invention relates to a method for preparing a library of template polynucleotides that have common sequences at their 5′ ends and at their 3′ ends.
Owner:ILLUMINA CAMBRIDGE LTD
Digital Counting of Individual Molecules by Stochastic Attachment of Diverse Label-Tags
ActiveUS20130116130A1Highly precise relative and absolute counting statisticRaise countNucleotide librariesMicrobiological testing/measurementBiology
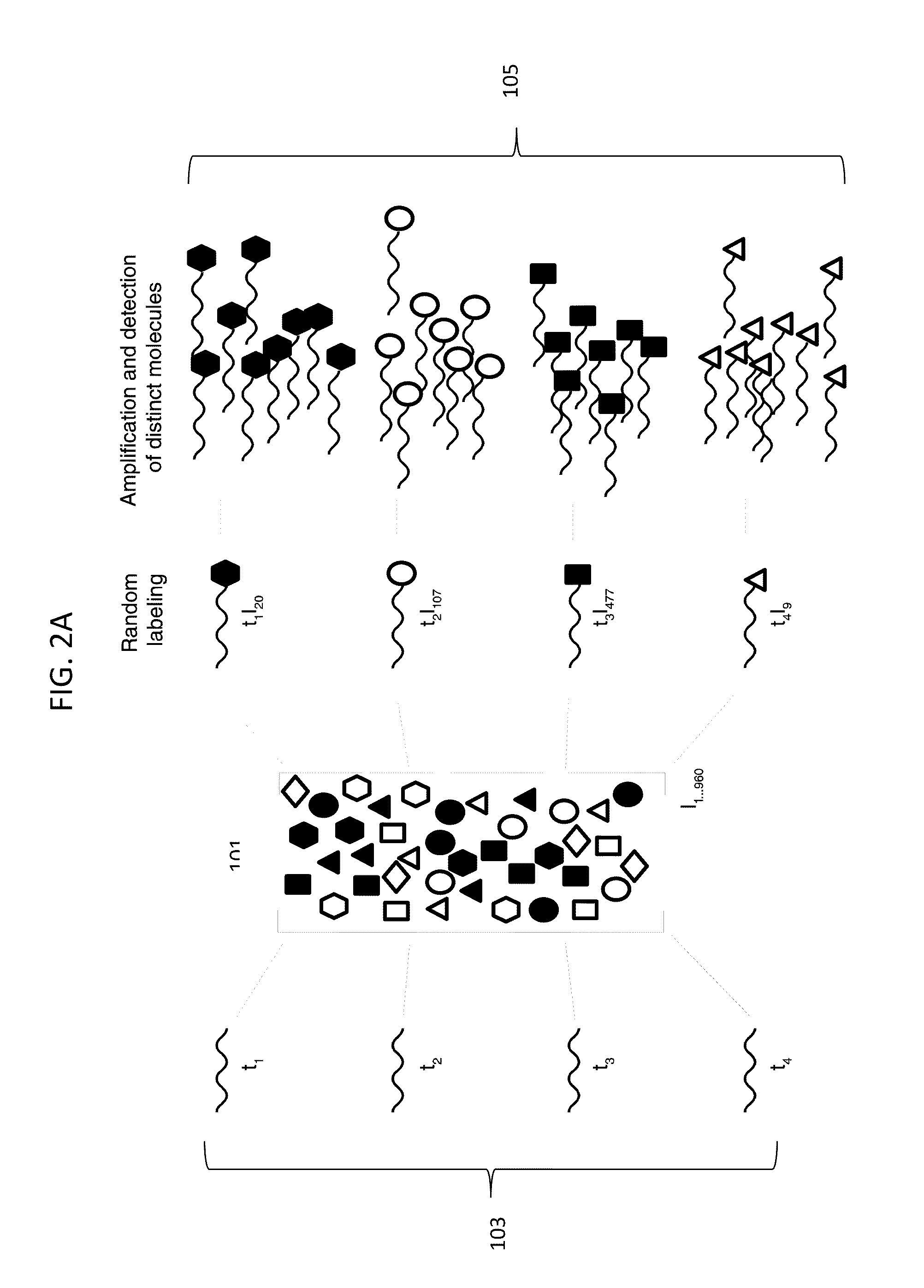
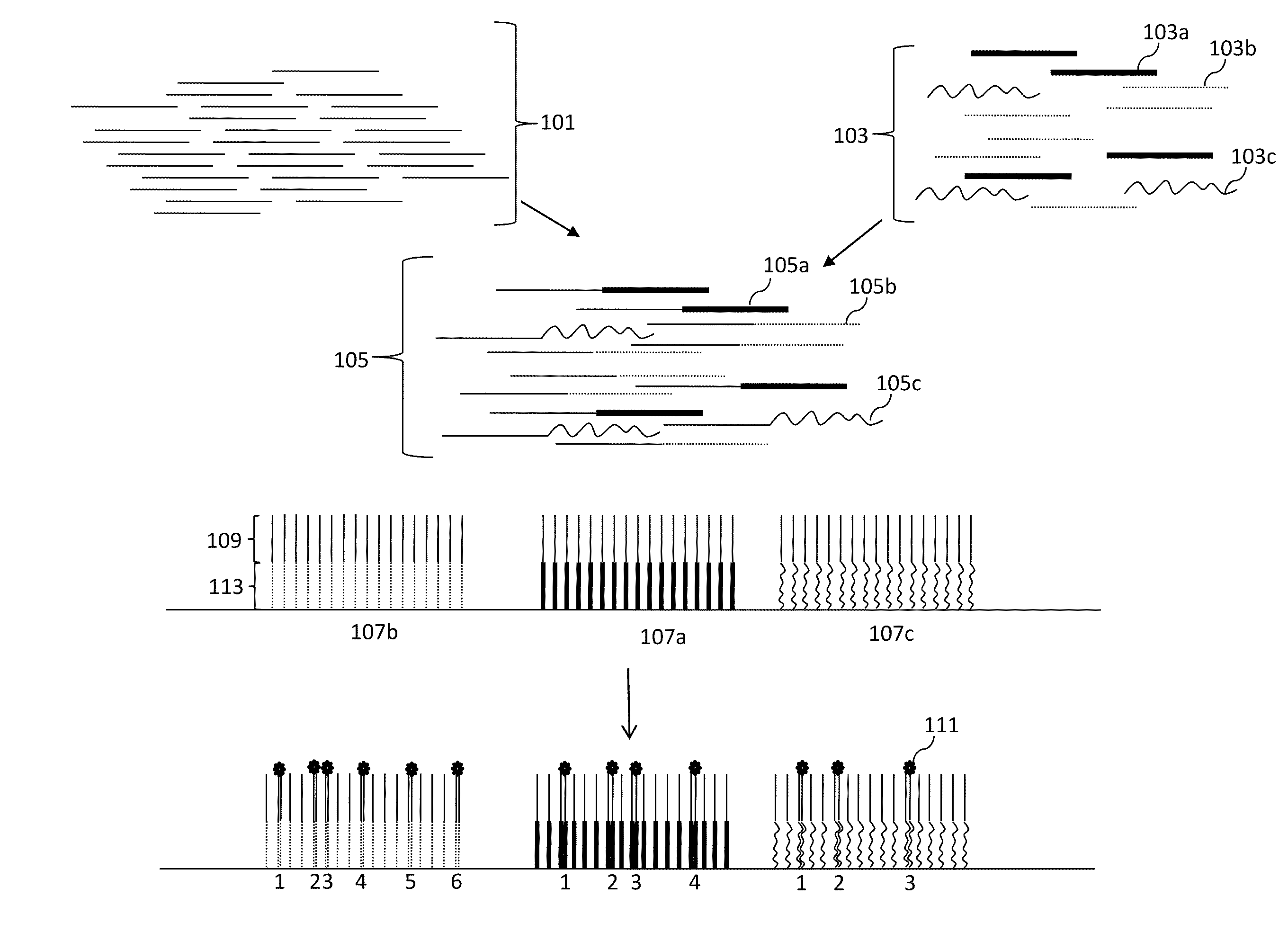
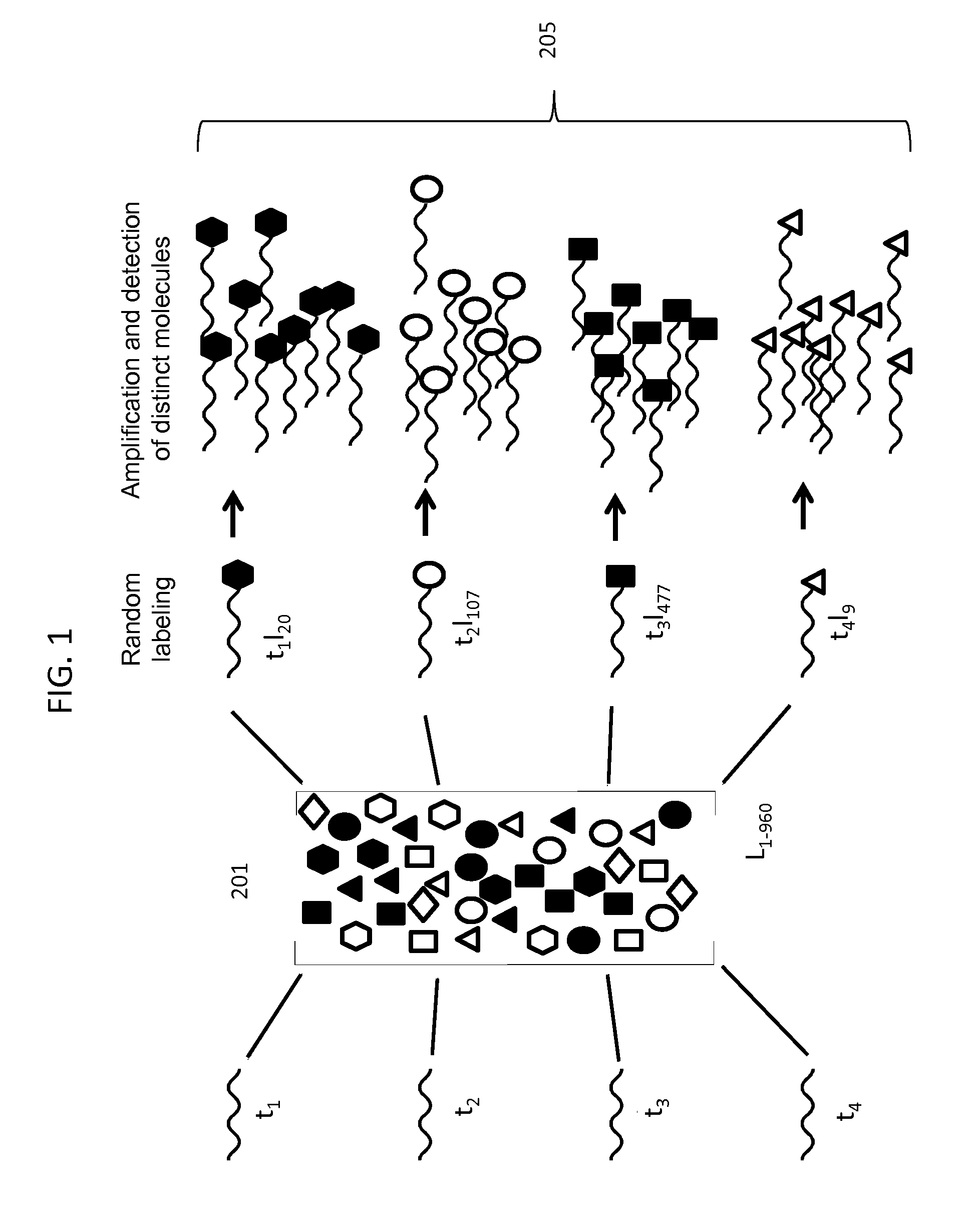
Compositions, methods and kits are disclosed for high-sensitivity counting of individual molecules by stochastic labeling of a identical molecules in mixtures of molecules by attachment of a unique label-tags from a diverse pool of label tags to confer uniqueness to otherwise identical or indistinguishable events. Individual occurrences of target molecules randomly choose from a non-depleting reservoir of diverse label-tags. Labeled molecules may be detected by hybridization or sequencing based methods. Molecules that would otherwise be identical in information content are labeled to create a separately detectable product that can be distinctly detected. The disclosed stochastic transformation methods reduce the problem of counting molecules from one of locating and identifying identical molecules to a series of binary digital questions detecting whether preprogrammed label-tags are present. The methods may be used, for example, to count a given species of molecule within a sample.
Owner:BECTON DICKINSON & CO
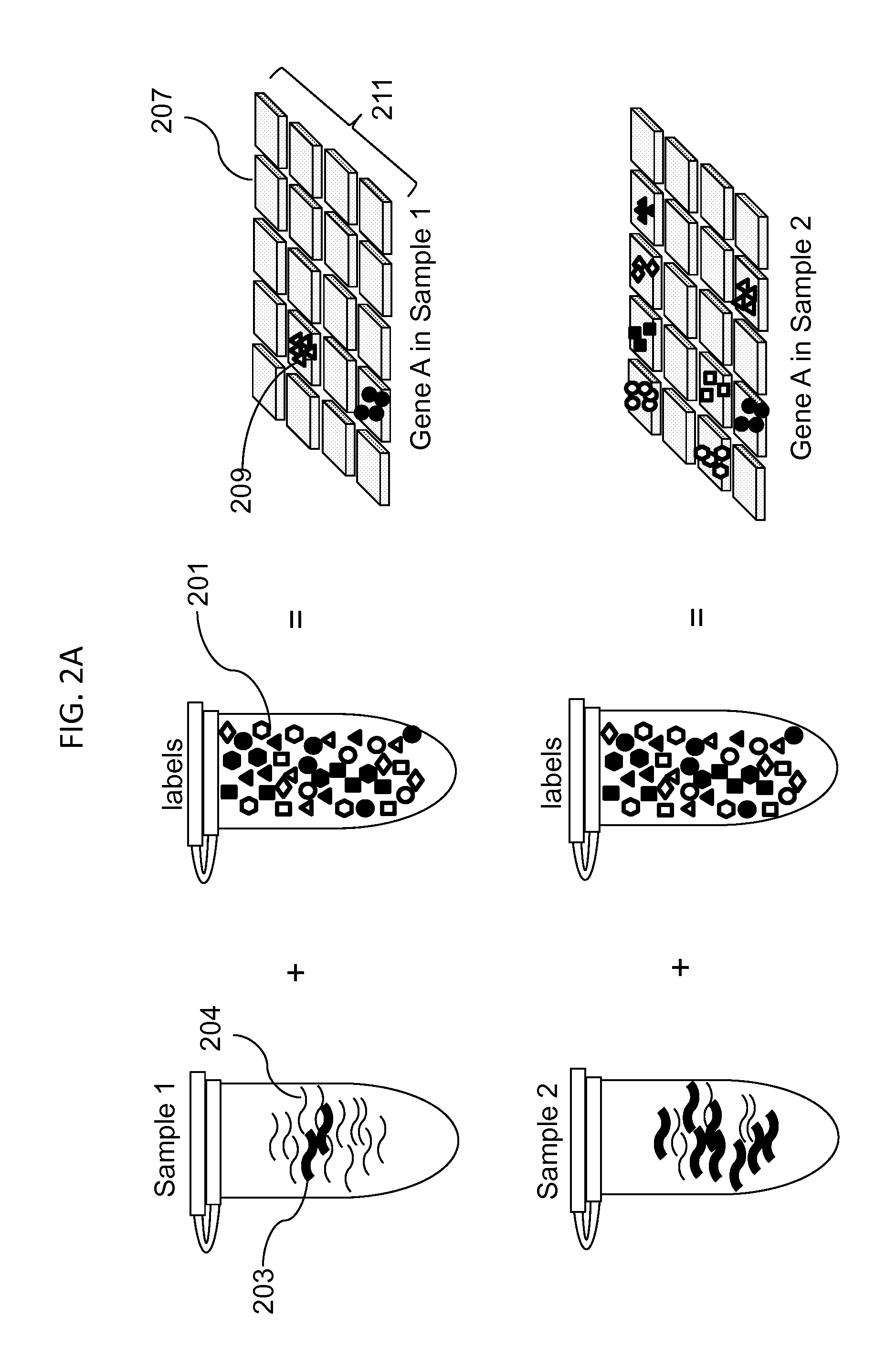
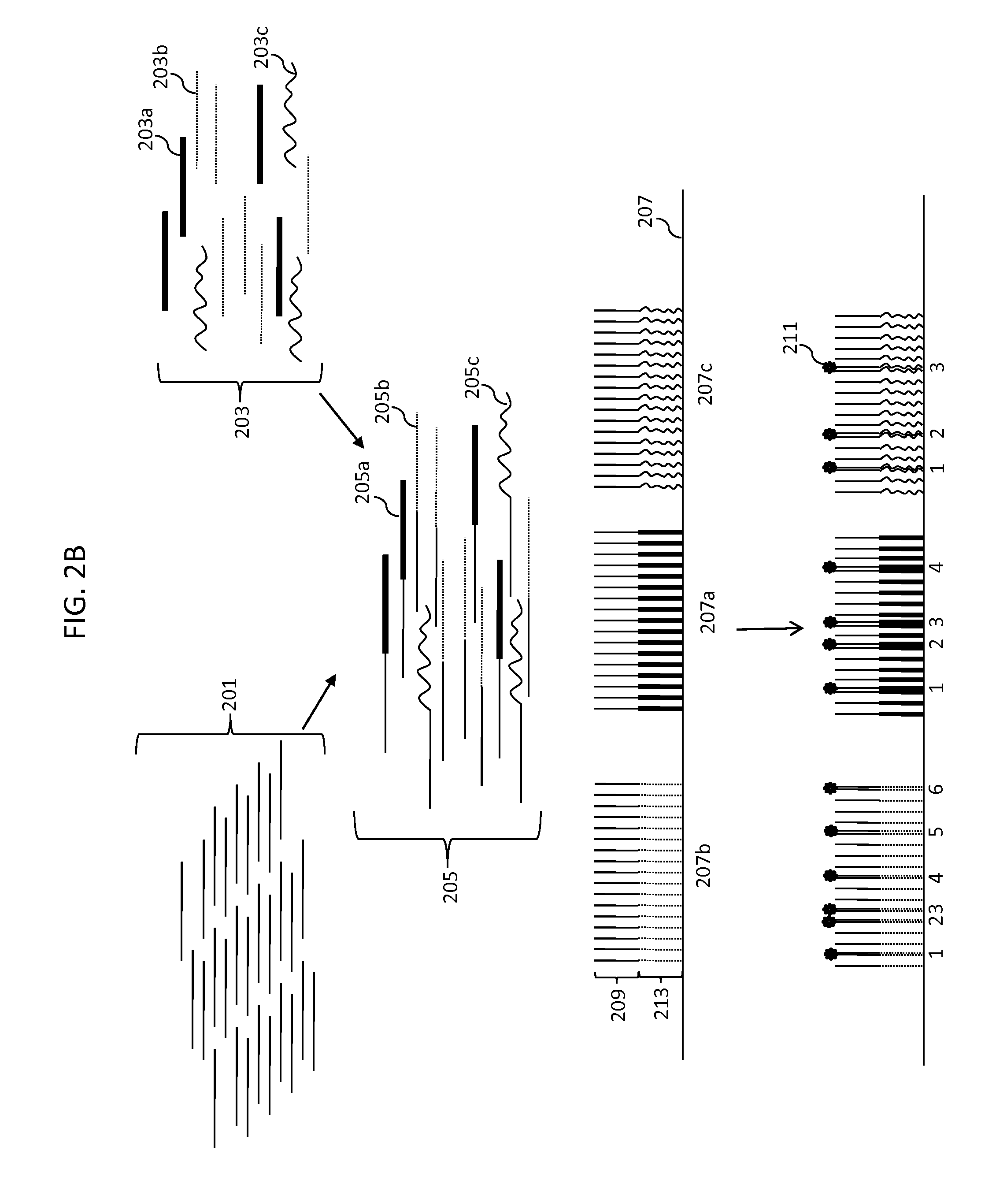
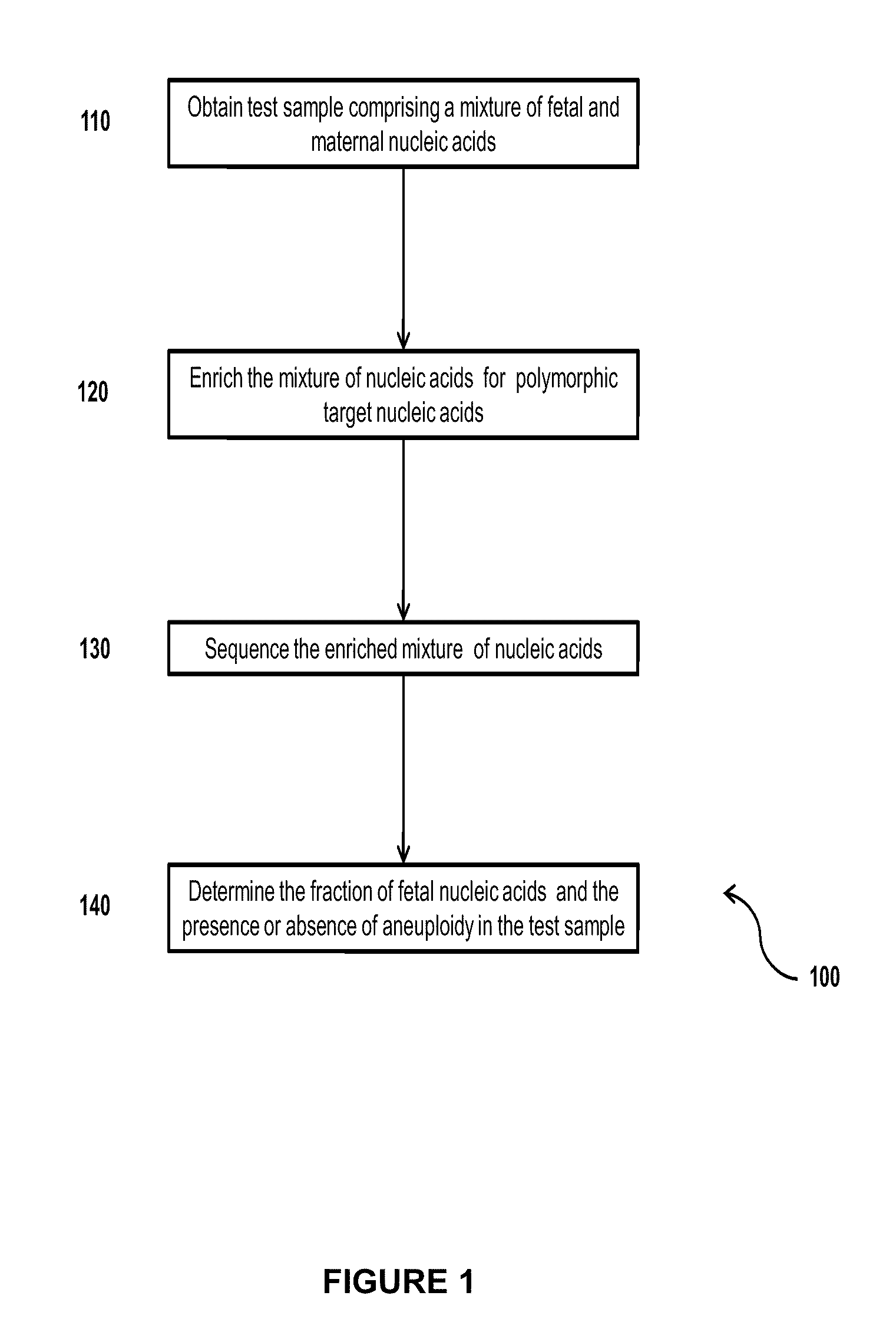
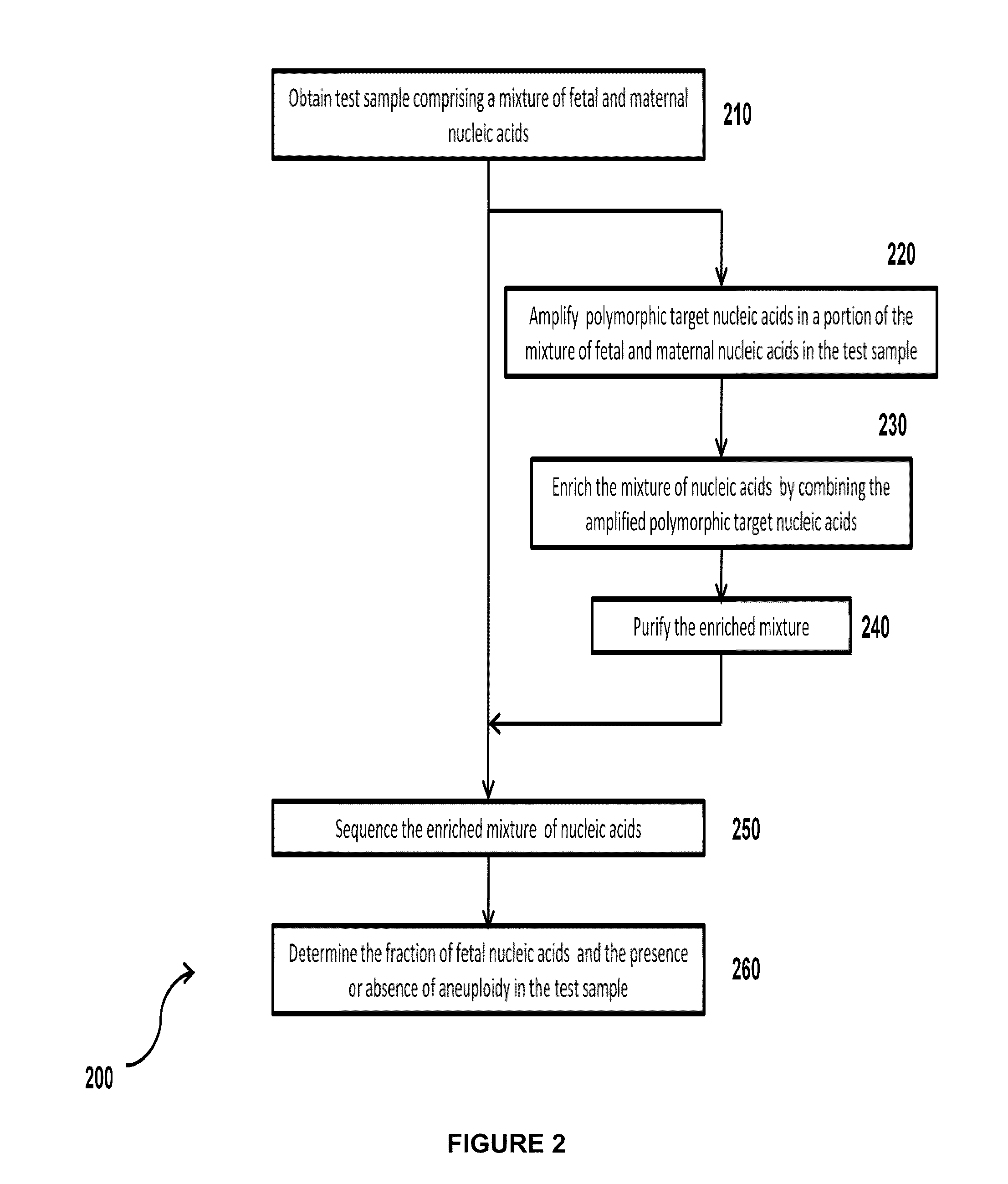
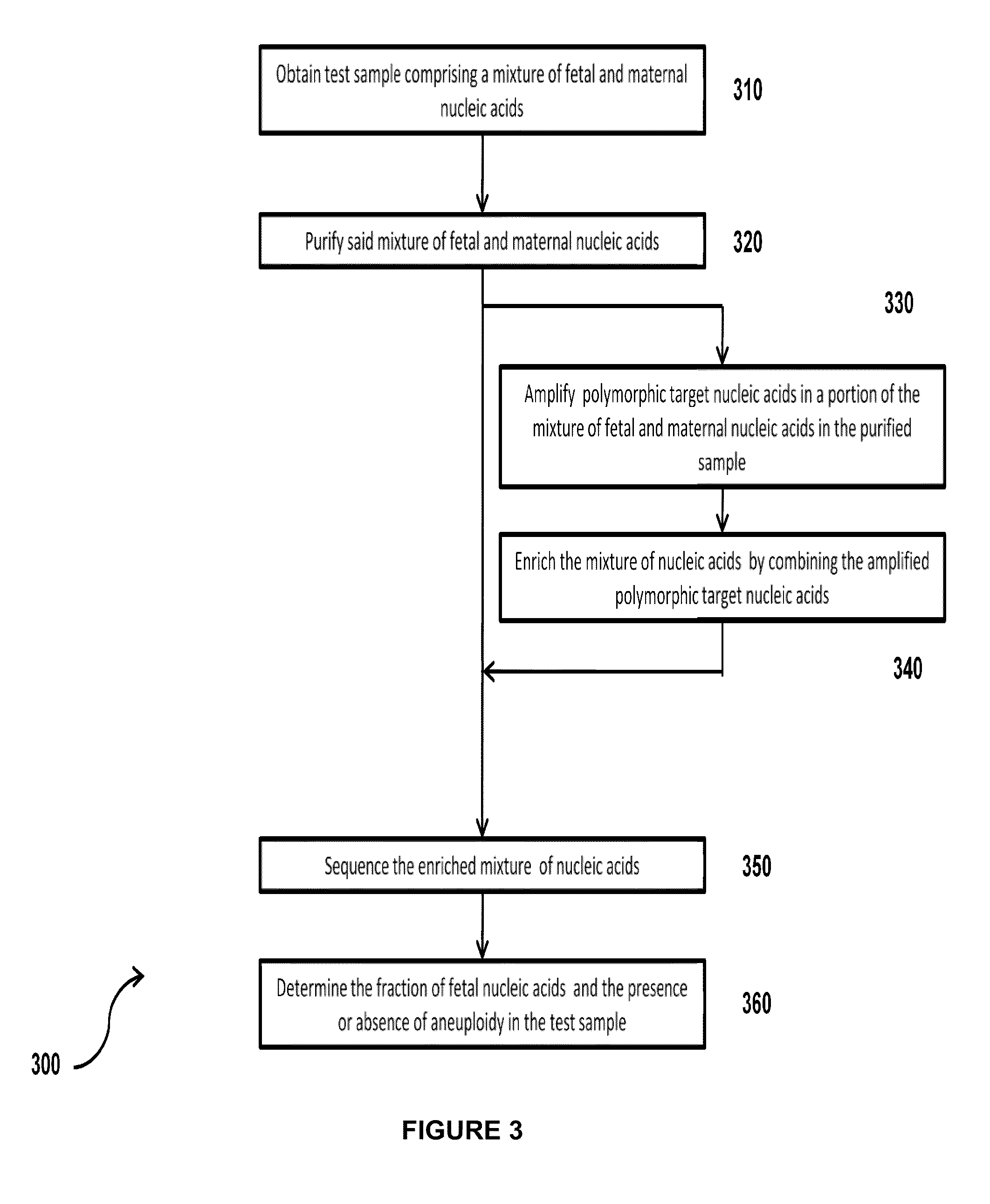
Simultaneous determination of aneuploidy and fetal fraction
InactiveUS20110224087A1Nucleotide librariesMicrobiological testing/measurementFetal aneuploidyBiology
The invention provides compositions and methods for simultaneously determining the presence or absence of fetal aneuploidy and the relative amount of fetal nucleic acids in a sample obtained form a pregnant female. The method encompasses the use of sequencing technologies and exploits the occurrence of polymorphisms to provide a streamlined noninvasive process applicable to the practice of prenatal diagnostics.
Owner:VERINATA HEALTH INC
Oligonucleotide arrays and their use for sorting, isolating, sequencing, and manipulating nucleic acids
InactiveUS6322971B1Increase in hybridization specificityBioreactor/fermenter combinationsMaterial nanotechnologyHybridization ArrayBioinformatics
Ligation methods for manipulating nucleic acid stands and oligonucleotides utilizing hybridization arrays of immobilized oligonucleotides. The oligonucleotide arrays may be plain or sectioned, comprehensive or non-comprehensive. The immobilized oligonucleotides may in some cases be binary oligonucleotides having constant as well as variable segments. Some embodiments include amplification of ligated products.
Owner:UNIV OF MEDICINE & DENTISTRY OF NEW JERSEY
Microarray fabrication system and method
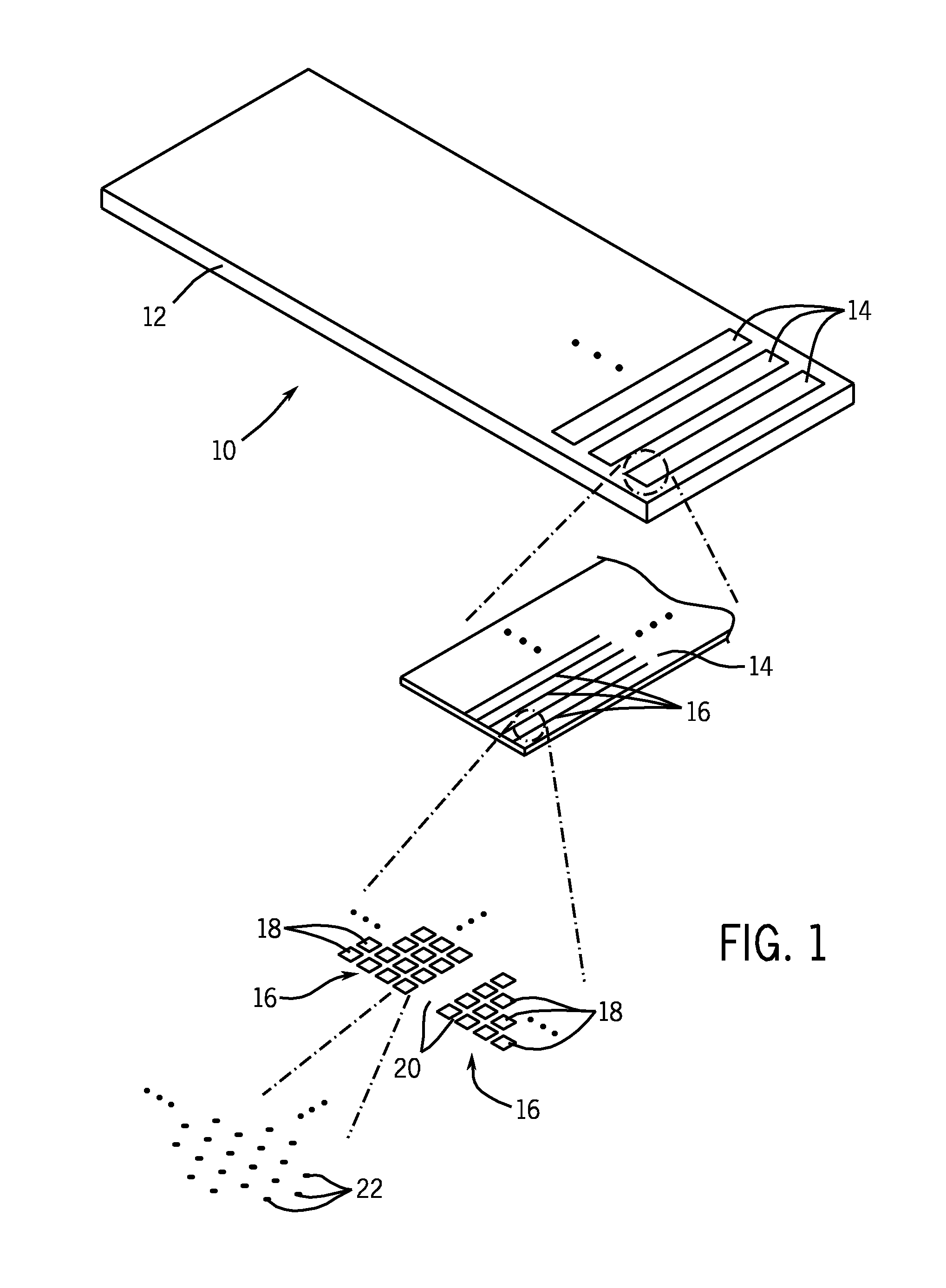
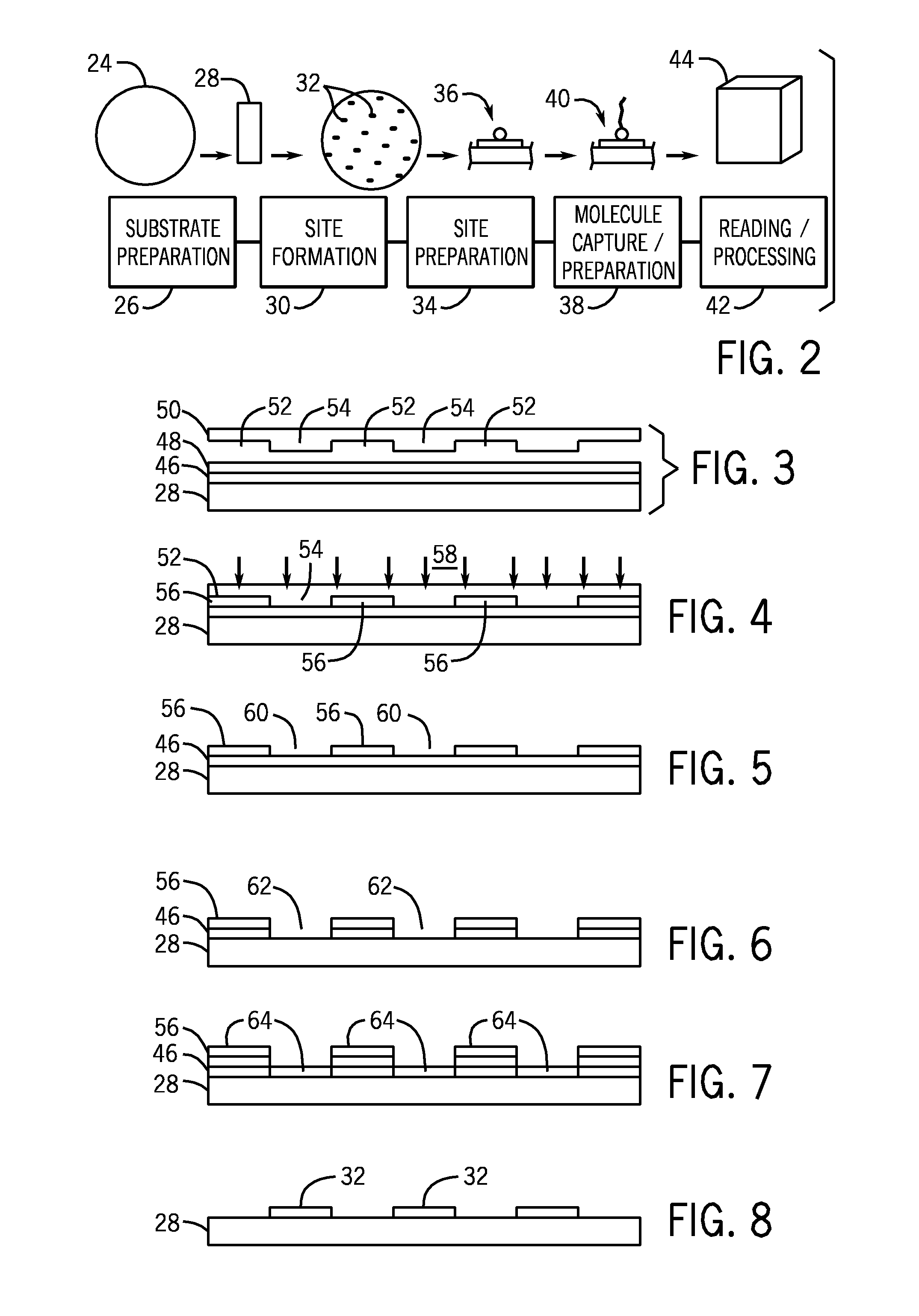
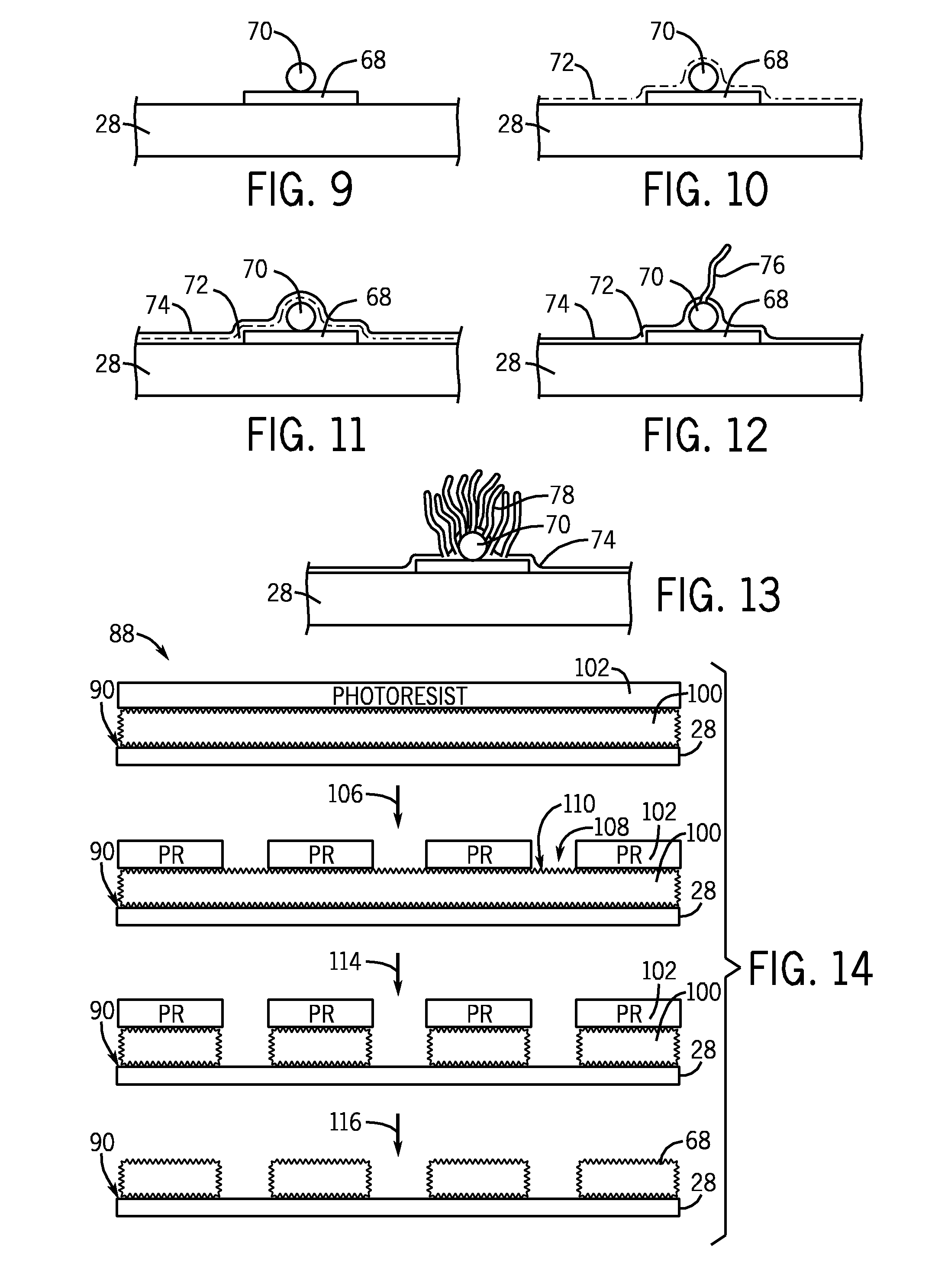
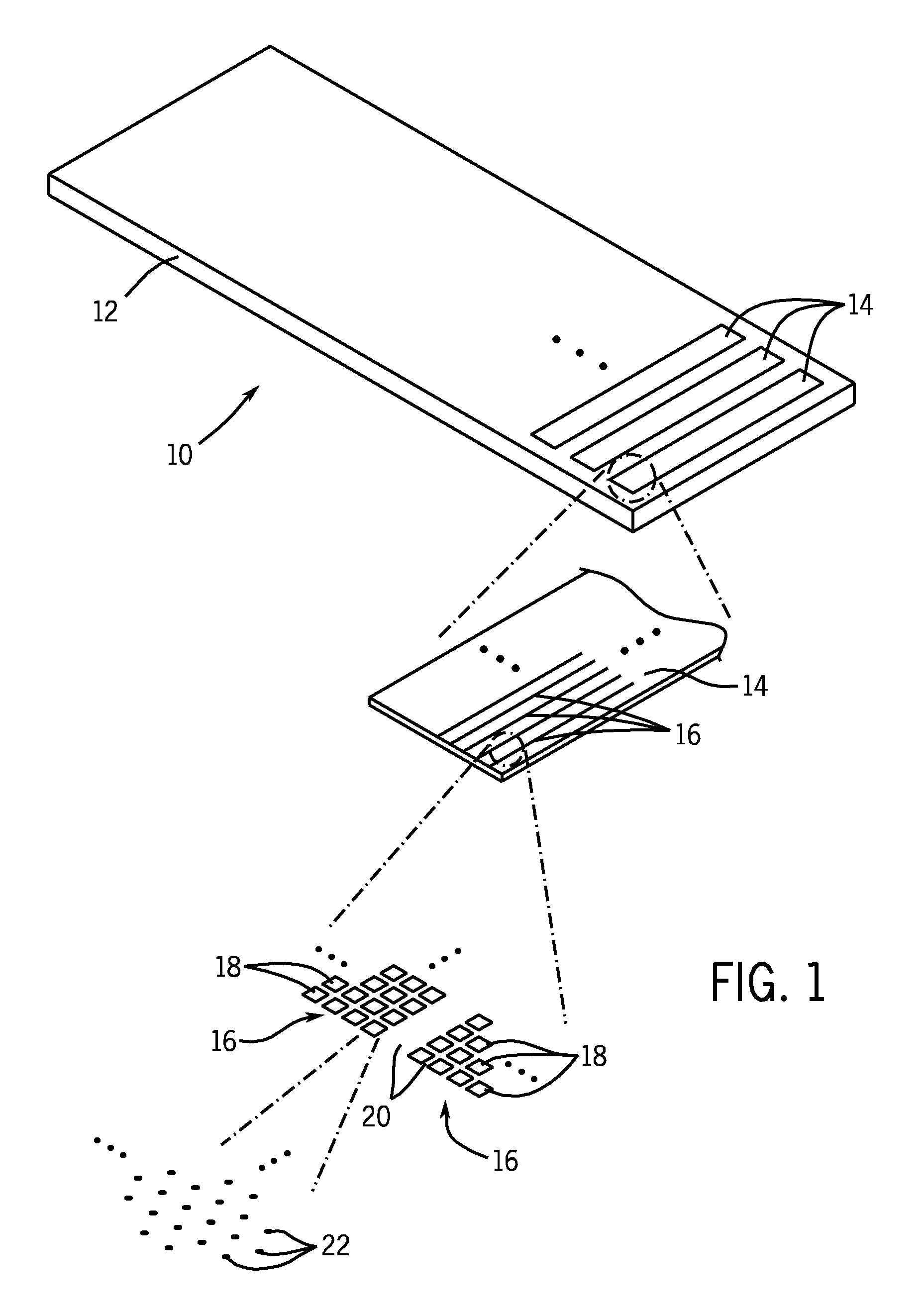
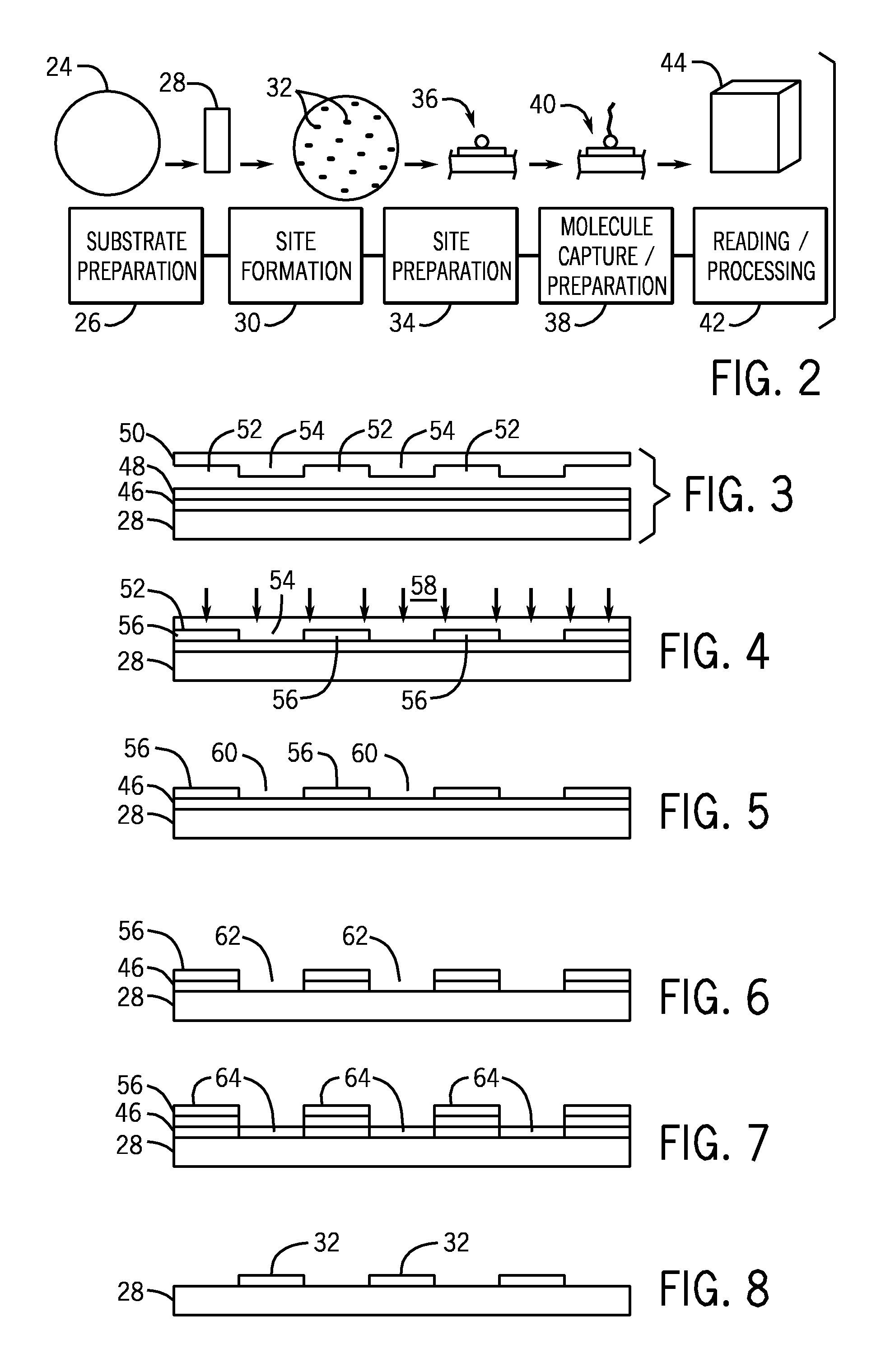
A microarray is designed capture one or more molecules of interest at each of a plurality of sites on a substrate. The sites comprise base pads, such as polymer base pads, that promote the attachment of the molecules at the sites. The microarray may be made by one or more patterning techniques to create a layout of base pads in a desired pattern. Further, the microarrays may include features to encourage clonality at the sites.
Owner:ILLUMINA INC
Nucleic acid encoding reactions
ActiveUS20130005585A1Avoid substantial annealingNucleotide librariesMicrobiological testing/measurementNucleotideBarcode
Described herein are methods useful for incorporating one or more adaptors and / or nucleotide tag(s) and / or barcode nucleotide sequence(s) one, or typically more, target nucleotide sequences. In particular embodiments, nucleic acid fragments having adaptors, e.g., suitable for use in high-throughput DNA sequencing are generated. In other embodiments, information about a reaction mixture is encoded into a reaction product. Also described herein are methods and kits useful for amplifying one or more target nucleic acids in preparation for applications such as bidirectional nucleic acid sequencing. In particular embodiments, methods of the invention entail additionally carrying out bidirectional DNA sequencing. Also described herein are methods for encoding and detecting and / or quantifying alleles by primer extension.
Owner:FLUIDIGM CORP
System for mixing fluids by coalescence of multiple emulsions
ActiveUS20110053798A1High-confidence resultLower the volumeSequential/parallel process reactionsHeating or cooling apparatusEmulsionChemistry
System, including methods, apparatus, compositions, and kits, for the mixing of small volumes of fluid by coalescence of multiple emulsions.
Owner:BIO RAD LAB INC
Microarray fabrication system and method
A microarray is designed capture one or more molecules of interest at each of a plurality of sites on a substrate. The sites comprise base pads, such as polymer base pads, that promote the attachment of the molecules at the sites. The microarray may be made by one or more patterning techniques to create a layout of base pads in a desired pattern. Further, the microarrays may include features to encourage clonality at the sites.
Owner:ILLUMINA INC
Kits for amplifying and detecting nucleic acid sequences, including a probe
InactiveUS6040166AStrong specificityReduce manpowerNanotechSequential/parallel process reactionsOligonucleotide primersNucleic acid sequencing
The present invention is directed to a process for amplifying any target nucleic acid sequence contained in a nucleic acid or mixture thereof using a thermostable enzyme. The process comprises treating separate complementary strands of the nucleic acid with a molar excess of two oligonucleotide primers, extending the primers with a thermostable enzyme to form complementary primer extension products which act as templates for synthesizing the desired nucleic acid sequence, and detecting the sequence so amplified. The steps of the reaction can be repeated as often as desired and involve temperature cycling to effect hybridization, promotion of activity of the enzyme, and denaturation of the hybrids formed.
Owner:ROCHE MOLECULAR SYST INC
Gel patterned surfaces
ActiveUS20140243224A1Easy to adaptSequential/parallel process reactionsNucleotide librariesBiologyNucleic acid
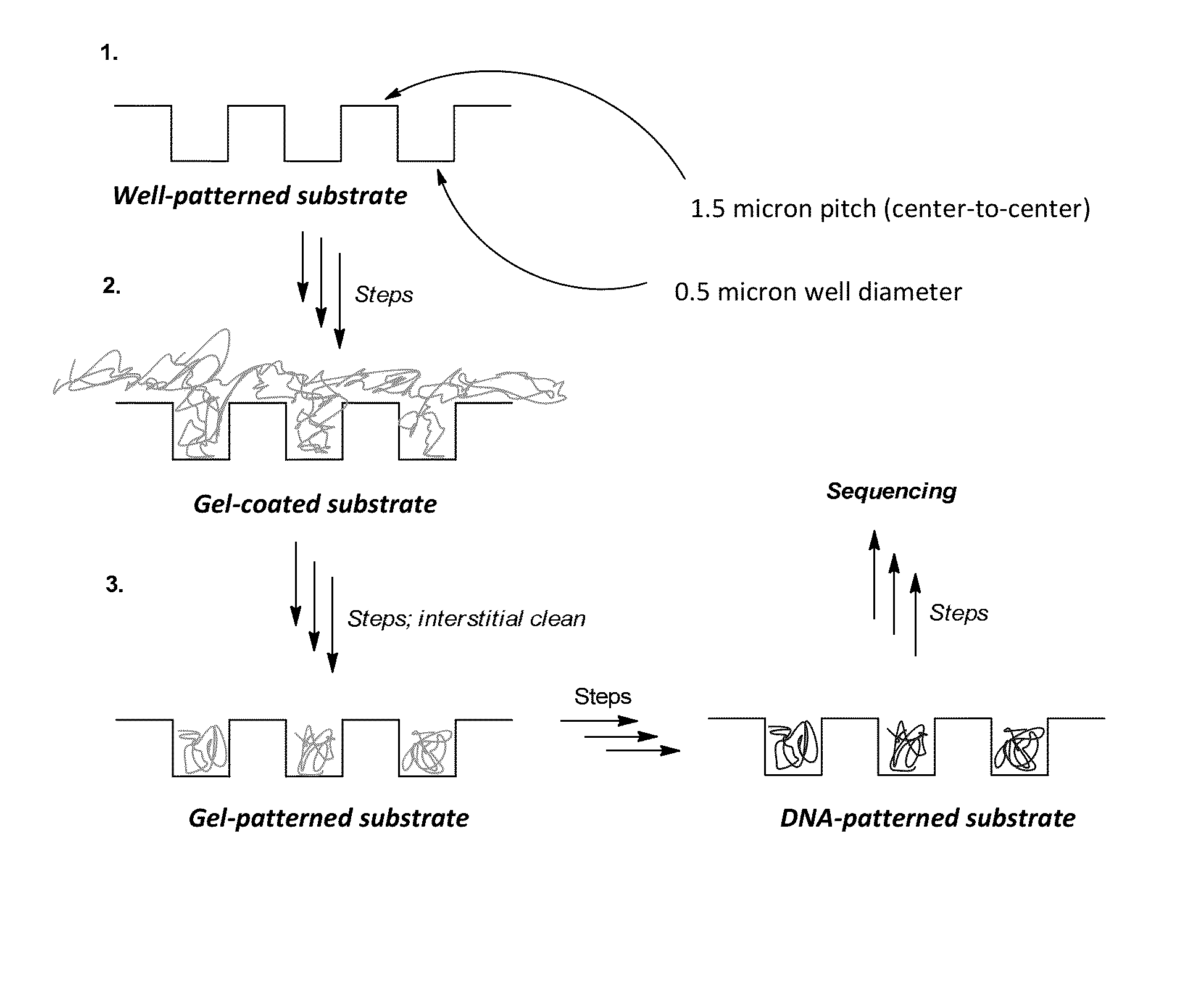
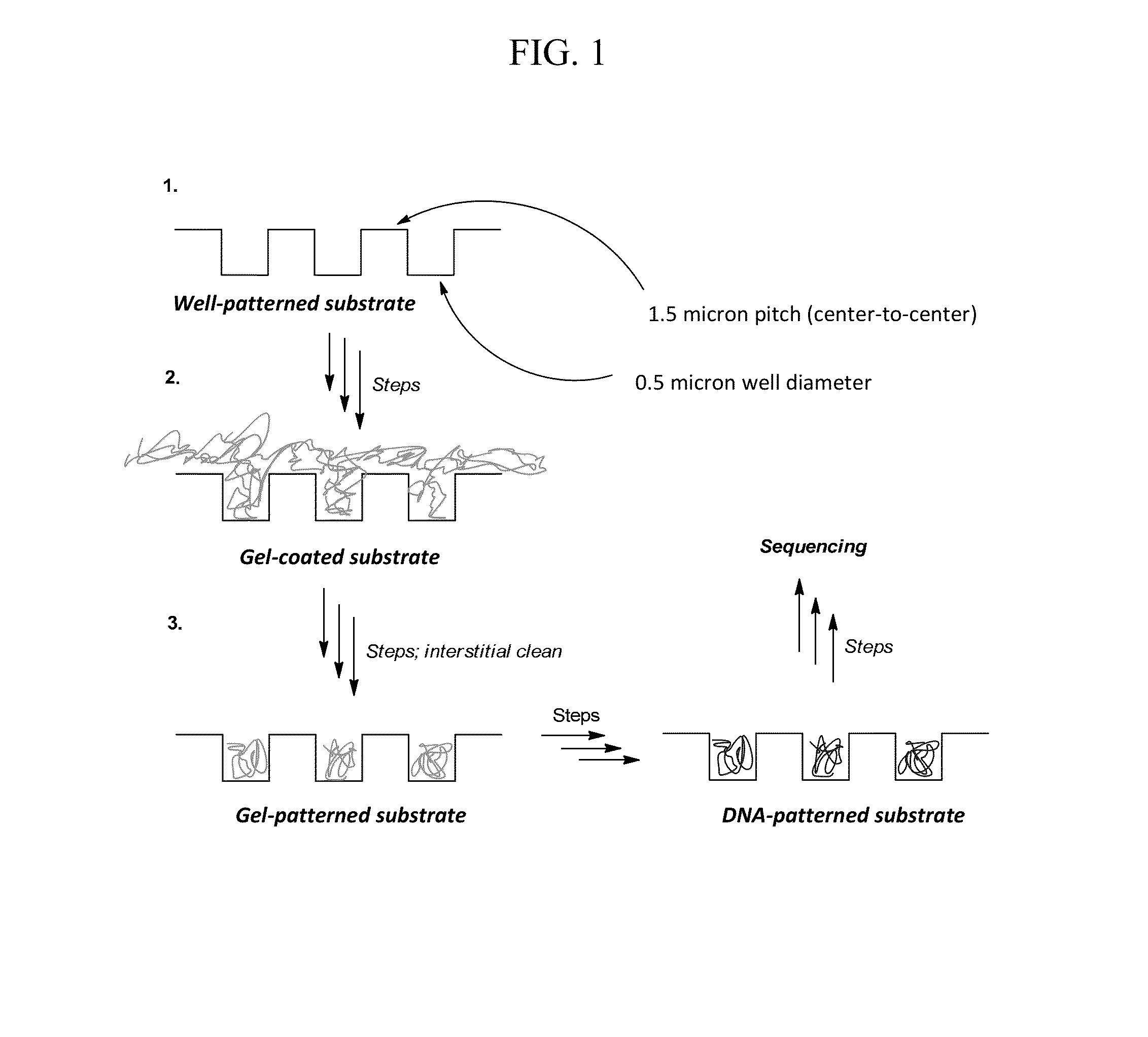
Provided is an array including a solid support having a surface, the surface having a plurality of wells, the wells containing a gel material, the wells being separated from each other by interstitial regions on the surface, the interstitial regions segregating the gel material in each of the wells from the gel material in other wells of the plurality; and a library of target nucleic acids in the gel material, wherein the gel material in each of the wells comprises a single species of the target nucleic acids of the library. Methods for making and using the array are also provided.
Owner:ILLUMINA INC
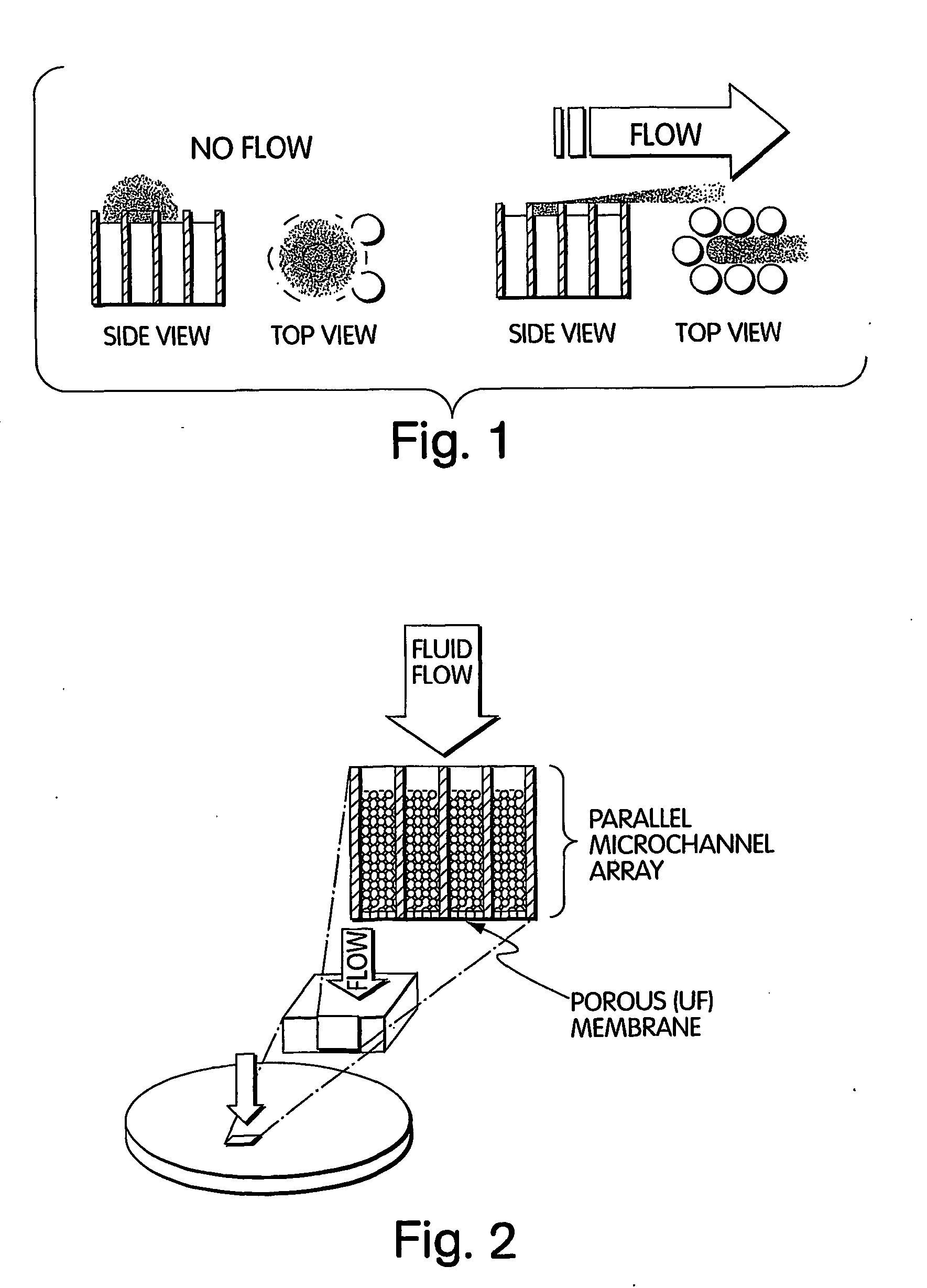
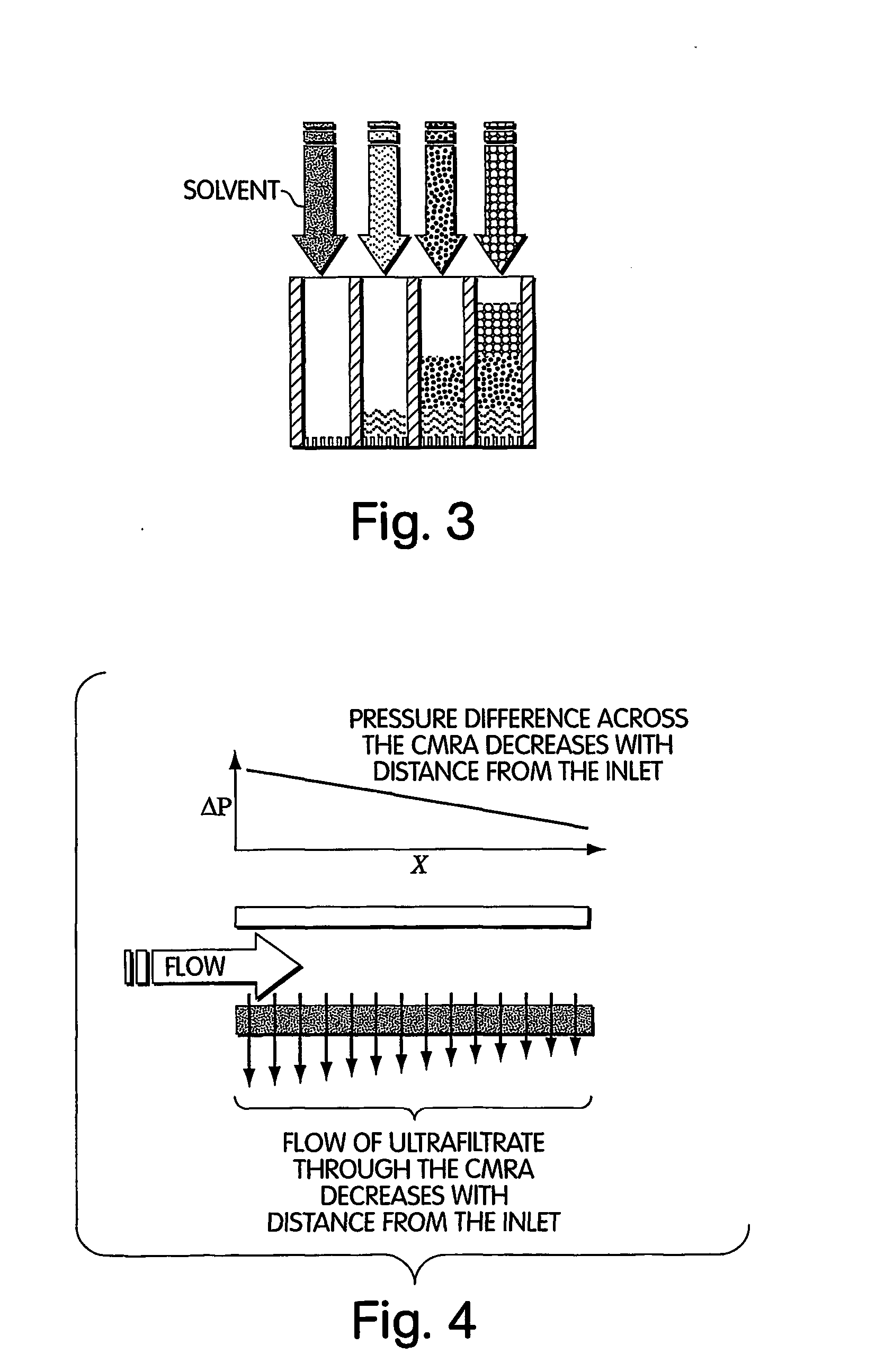
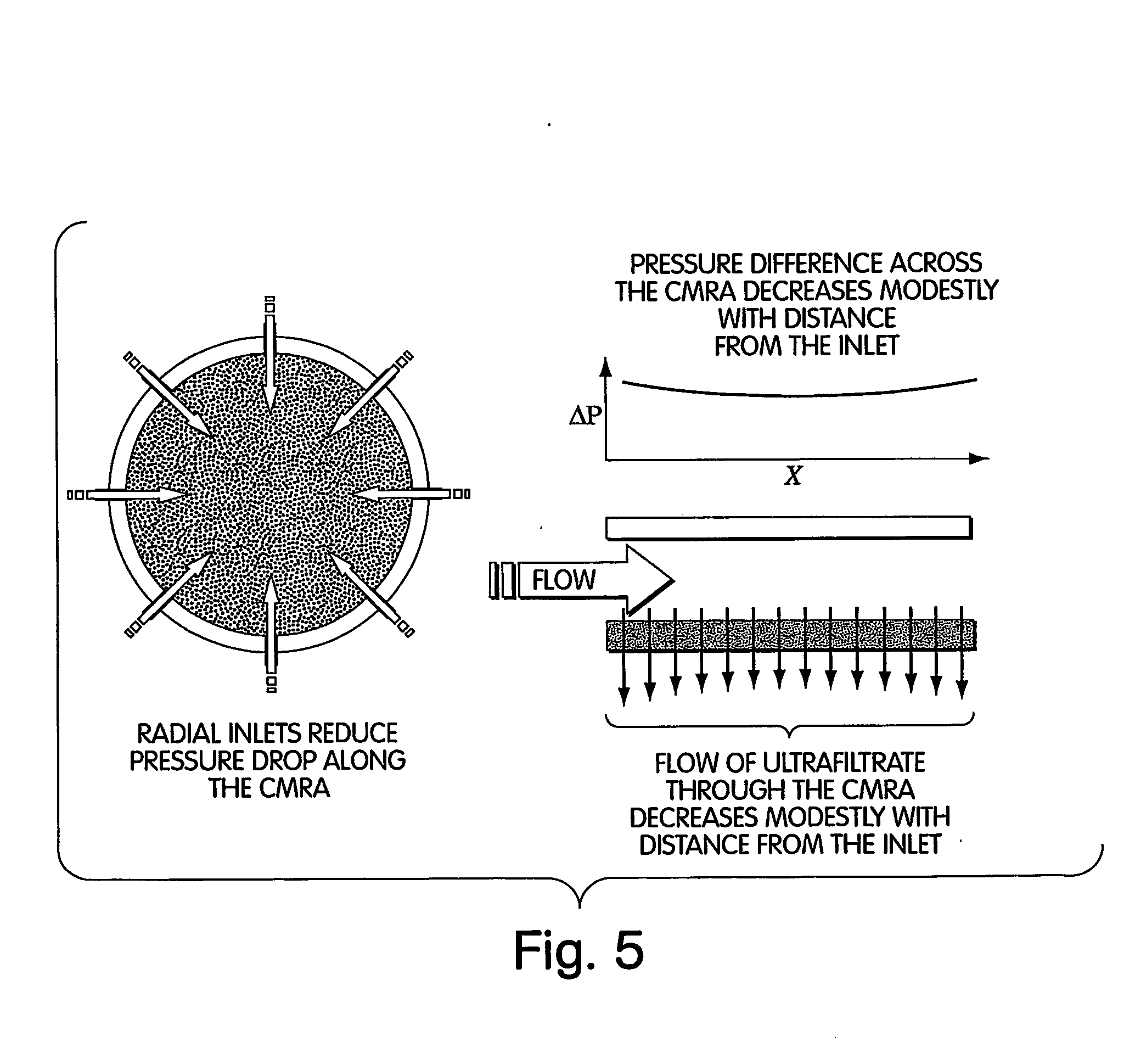
Method for isolation of independent, parallel chemical micro-reactions using a porous filter
InactiveUS20050009022A1Fast deliveryFaster more complete removalBioreactor/fermenter combinationsPeptide librariesChemical reactionCompound (substance)
The present invention relates to the field of fluid dynamics. More specifically, this invention relates to methods and apparatus for conducting densely packed, independent chemical reactions in parallel in a substantially two-dimensional array. Accordingly, this invention also focuses on the use of this array for applications such as DNA sequencing, most preferably pyrosequencing, and DNA amplification.
Owner:WEINER MICHAEL P +4
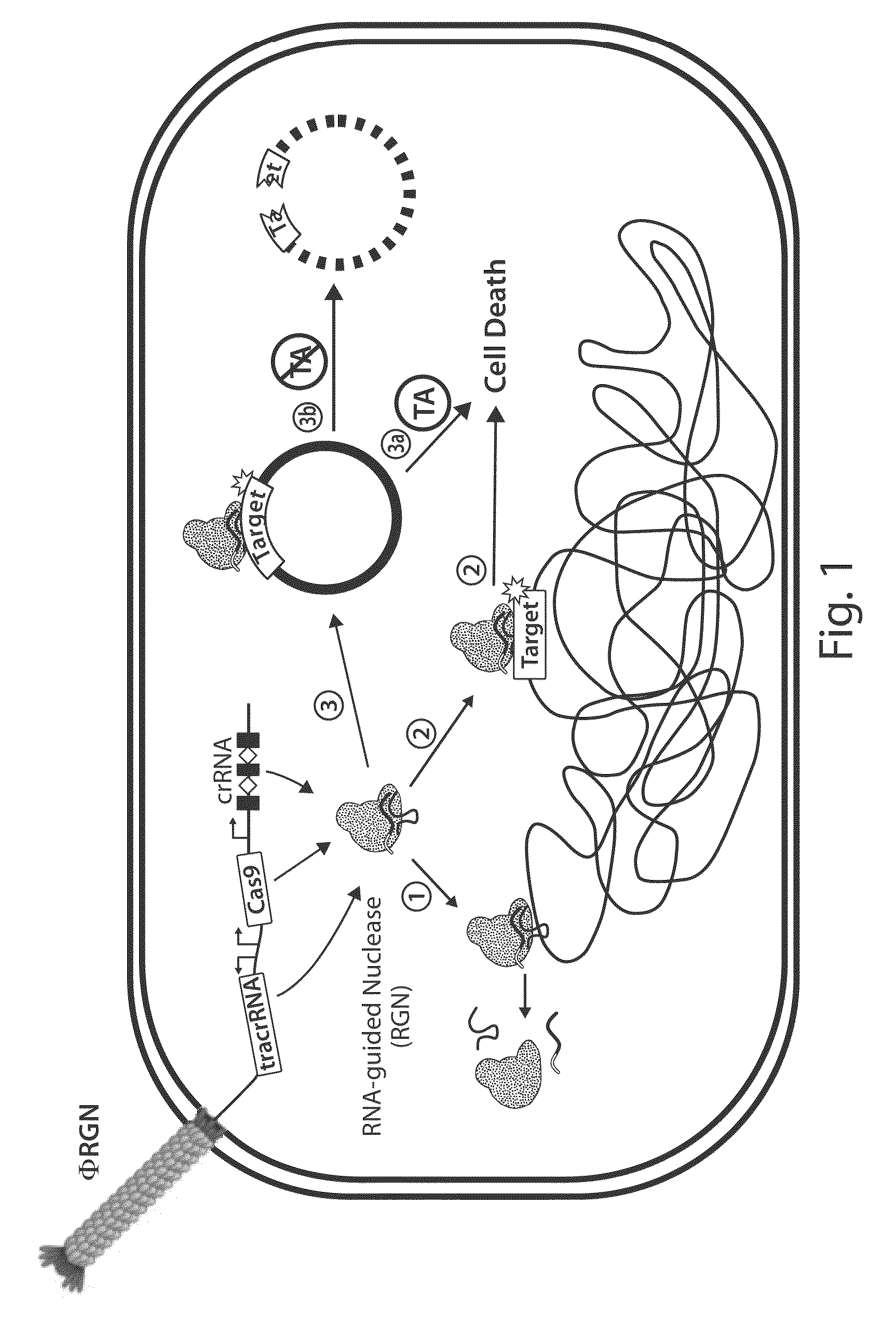
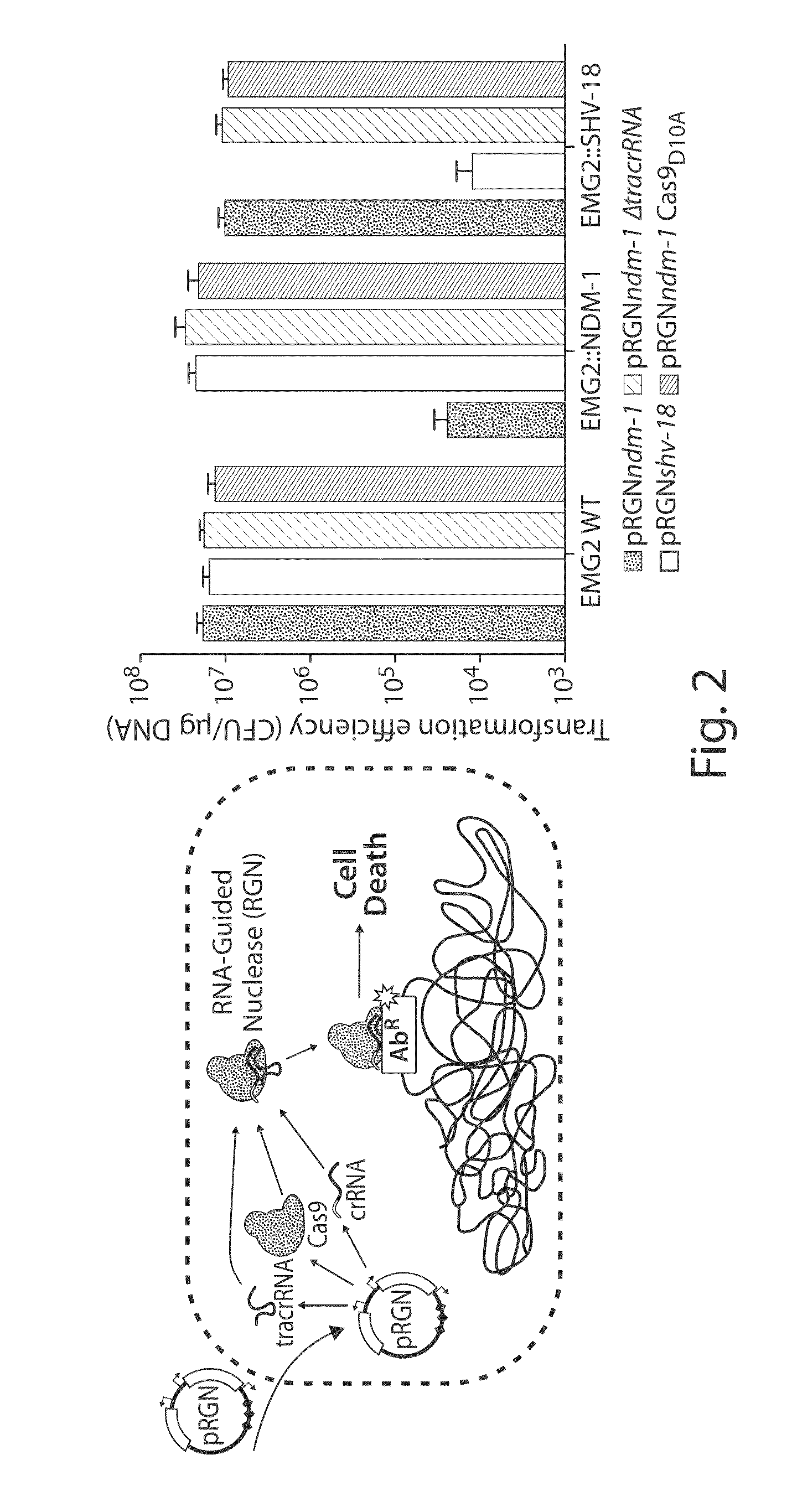
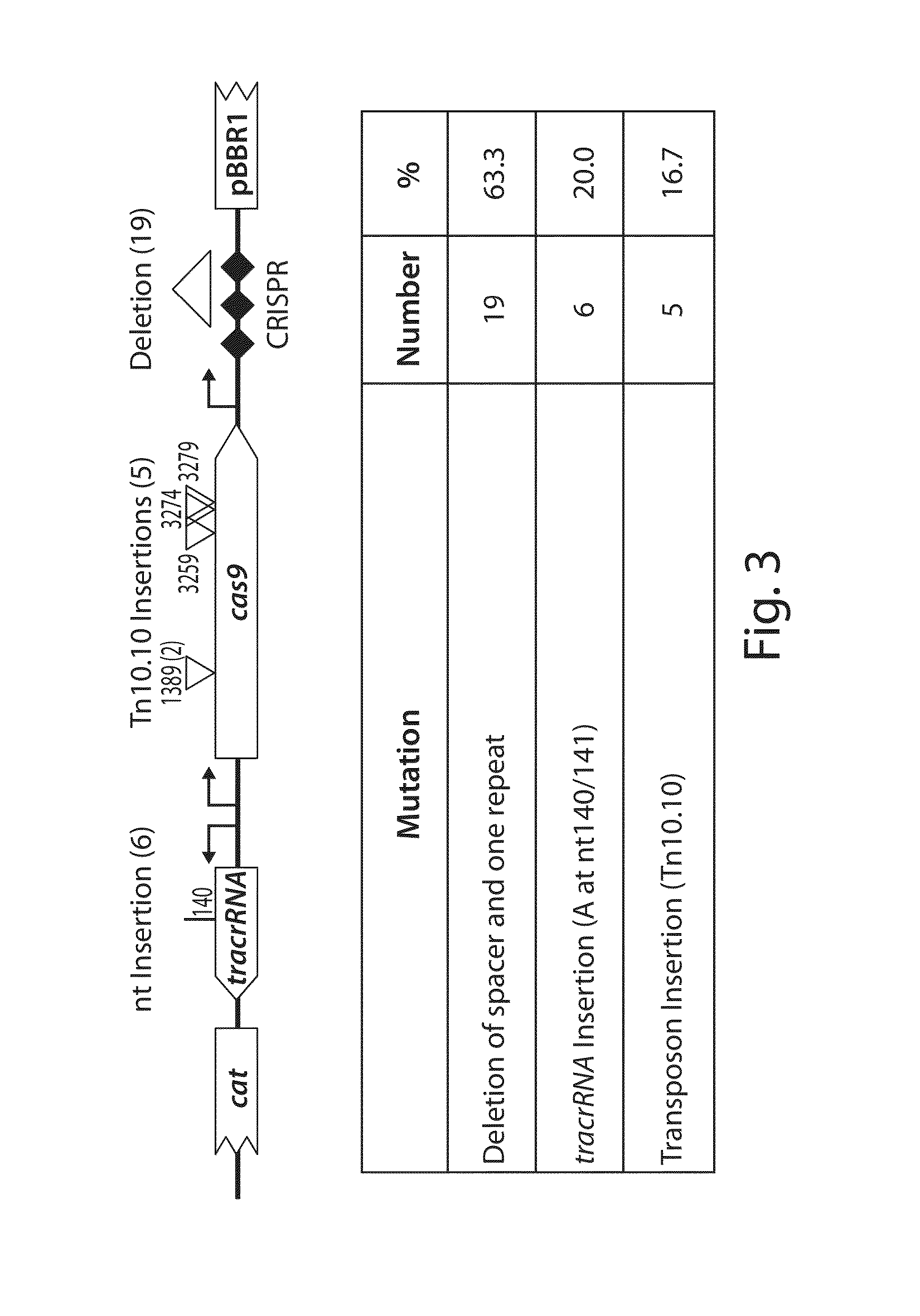
Tuning microbial populations with programmable nucleases
ActiveUS20150064138A1Efficiently and rapidly encoded into synthetic constructBroad deliveryBiocideSugar derivativesMicroorganismVirulent characteristics
Various aspects and embodiments of the invention are directed to methods and compositions for reversing antibiotic resistance or virulence in and / or destroying pathogenic microbial cells such as, for example, pathogenic bacterial cells. The methods include exposing microbial cells to a delivery vehicle with at least one nucleic acid encoding an engineered autonomously distributed circuit that contains a programmable nuclease targeted to one or multiple genes of interest.
Owner:MASSACHUSETTS INST OF TECH
Gene expression analysis in single cells
The present invention provides methods and compositions for the analysis of gene expression in single cells or in a plurality of single cells. The invention provides methods for preparing a cDNA library from individual cells by releasing mRNA from each single cell to provide a plurality of individual mRNA samples, synthesizing cDNA from the individual mRNA samples, tagging the individual cDNA, pooling the tagged cDNA samples and amplifying the pooled cDNA samples to generate a cDNA library. The invention also provides a cDNA library produced by the methods described herein. The invention farther provides methods for analyzing gene expression in a plurality of cells by preparing a cDNA library as described herein and sequencing the library.
Owner:ILLUMINA INC
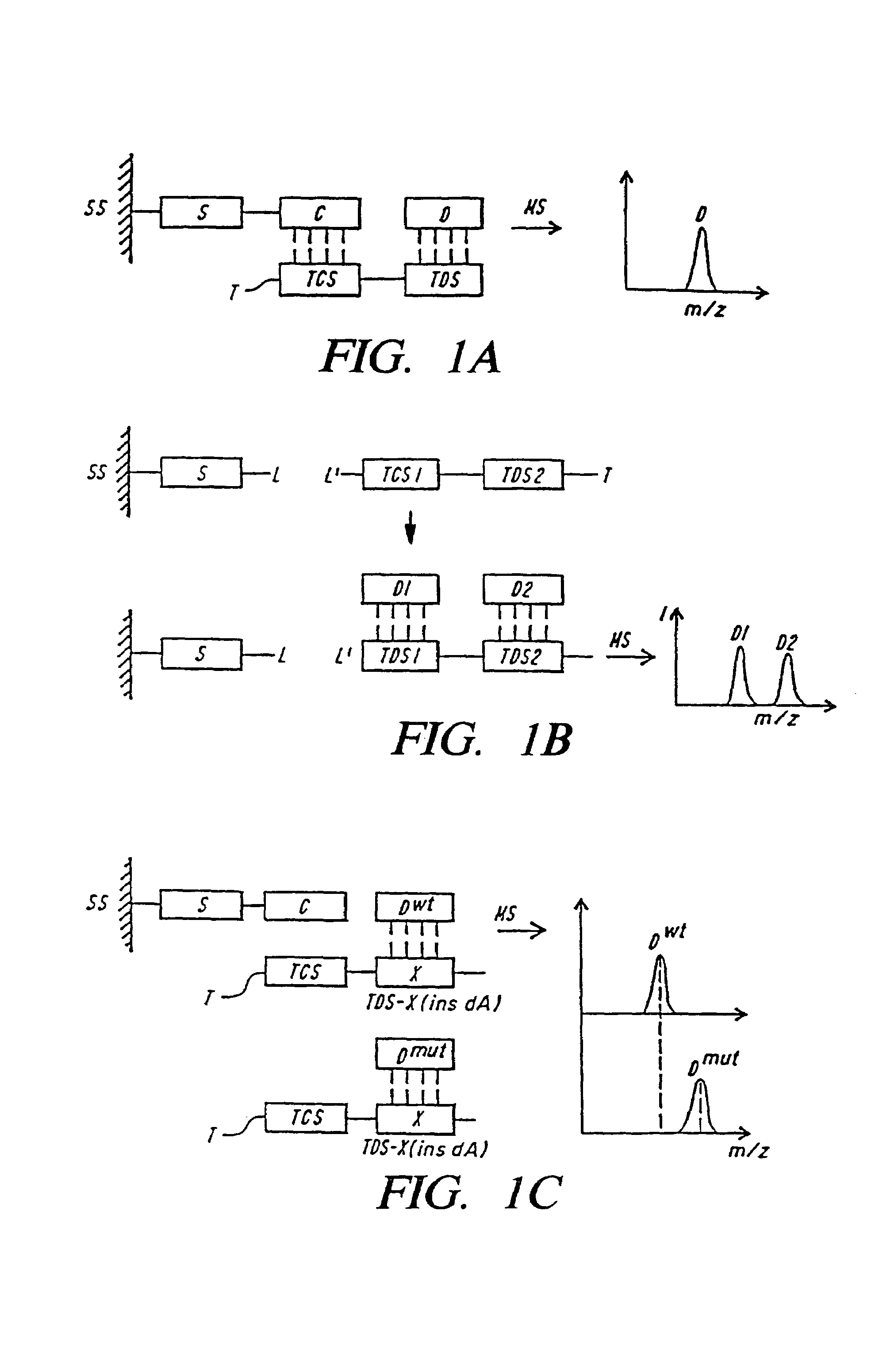
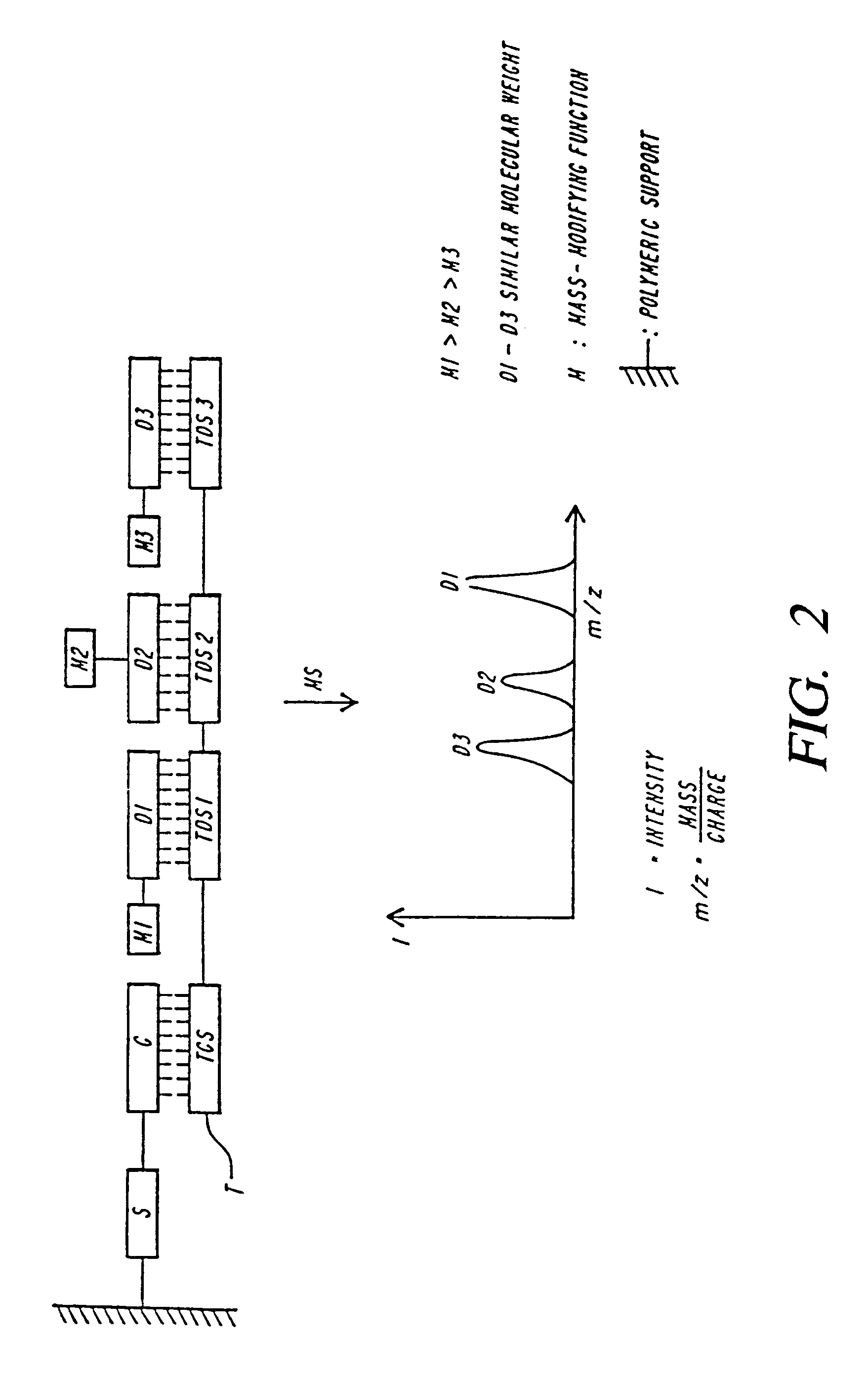
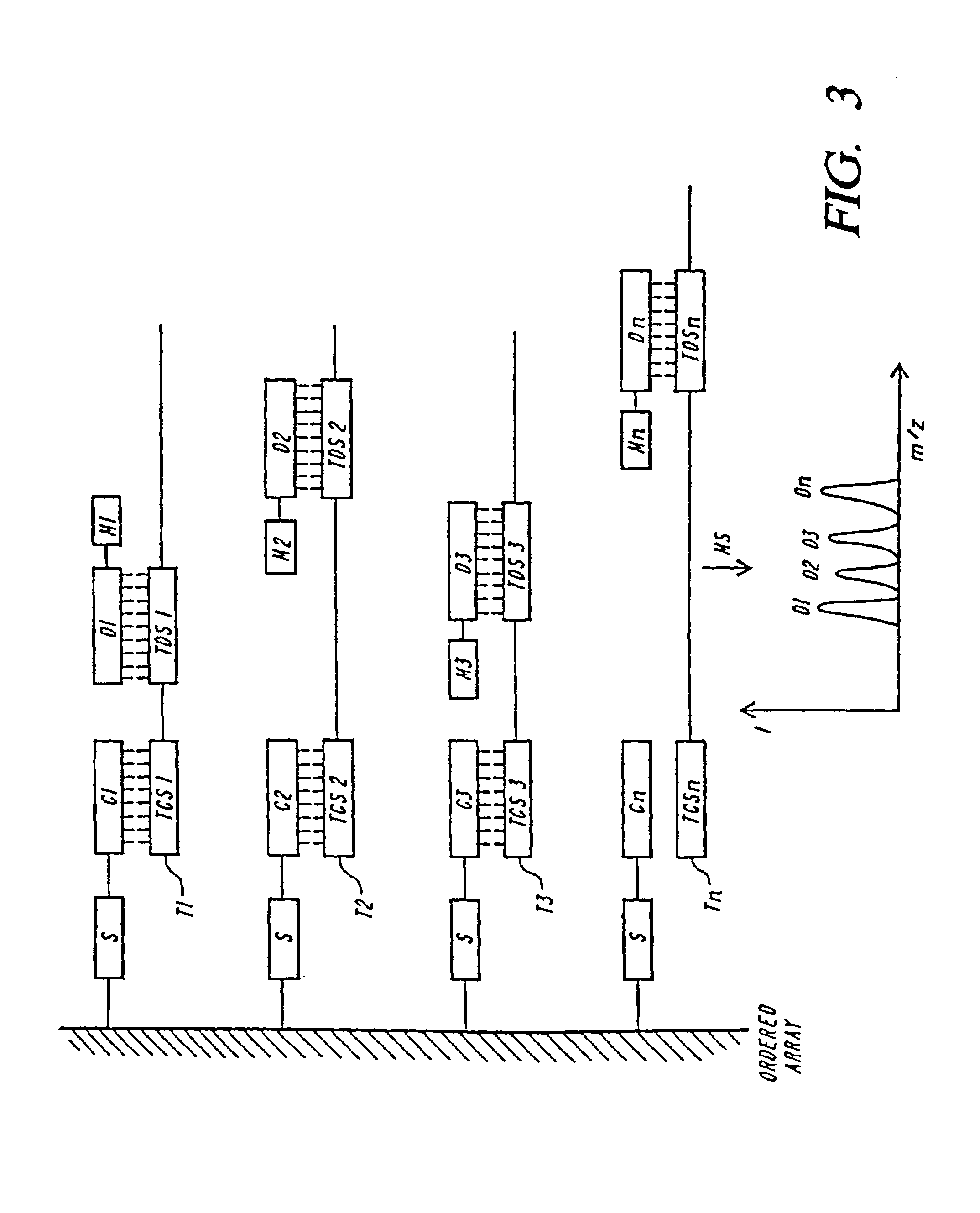
DNA diagnostics based on mass spectrometry
InactiveUS7198893B1Increase mass resolution massIncrease mass mass accuracyPeptide librariesSequential/parallel process reactionsMass spectrometry imagingOrganism
Fast and highly accurate mass spectrometry-based processes for detecting a particular nucleic acid sequence in a biological sample are provided. Depending on the sequence to be detected, the processes can be used, for example, to diagnose a genetic disease or chromosomal abnormality; a predisposition to a disease or condition, infection by a pathogenic organism, or for determining identity or heredity.
Owner:AGENA BIOSCI
Nucleic acid sequence analysis from single cells
ActiveUS20160053253A1Reduce cascadeNucleotide librariesMicrobiological testing/measurementBarcodeNucleic acid sequencing
Presented herein are methods and compositions for multiplexed single cell gene expression analysis. Some methods and compositions include the use of droplets and / or beads bearing unique barcodes such as unique molecular barcodes (UMI).
Owner:ILLUMINA INC
Single molecule loading methods and compositions
ActiveUS20100009872A1Improve throughputImprove efficiencySequential/parallel process reactionsNucleotide librariesAnalyte moleculeMolecular physics
Methods, compositions and arrays for non-random loading of single analyte molecules into array structures are provided.
Owner:PACIFIC BIOSCIENCES
Nucleic acid amplification with continuous flow emulsion
InactiveUS7927797B2Rapid and economical mannerReduce nozzle cloggingHeating or cooling apparatusFlow mixersMicroreactorGenetic Materials
Embodiments of the present invention are directed to methods and devices / systems for amplifying genetic material and may include providing a water-in-oil emulsion in a continuous flow. The emulsion may include a plurality of water droplets comprising microreactors. Each of the plurality of microreactors may include a single bead capable of capturing a nucleic acid template, a single species nucleic acid template and sufficient reagents to amplify the copy number of the nucleic acid template. The method also includes flowing the emulsion across a first temperature zone and a second lower temperature zone to thermally process the microreactors to amplify the nucleic acid template by polymerase chain reaction.
Owner:454 LIFE SCIENCES CORP
Analysis of genetic polymorphisms and gene copy number
InactiveUS6468744B1Material nanotechnologySequential/parallel process reactionsCytochrome P450Drug biotransformation
The invention provides methods for detecting variations in polymorphic sites and / or variations in gene copy number. The methods are particularly useful for analysis of biotransformation genes, such as cytochromes P450.
Owner:AFFYMETRIX INC
Features
- R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
Why Patsnap Eureka
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Social media
Patsnap Eureka Blog
Learn More Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com