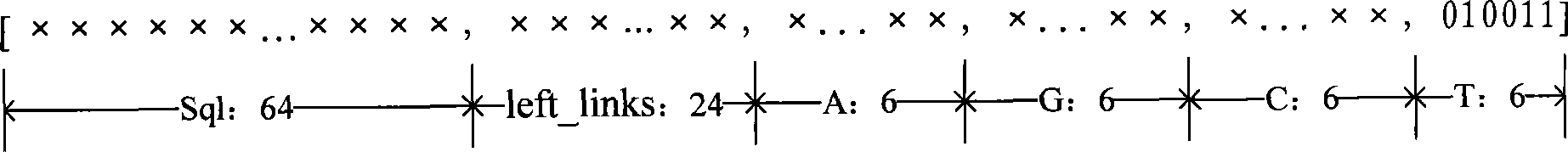Patents
Literature
Hiro is an intelligent assistant for R&D personnel, combined with Patent DNA, to facilitate innovative research.
52 results about "Short string" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Language identification from short strings
ActiveUS20160357728A1Natural language translationSemantic analysisNatural language processingShort string
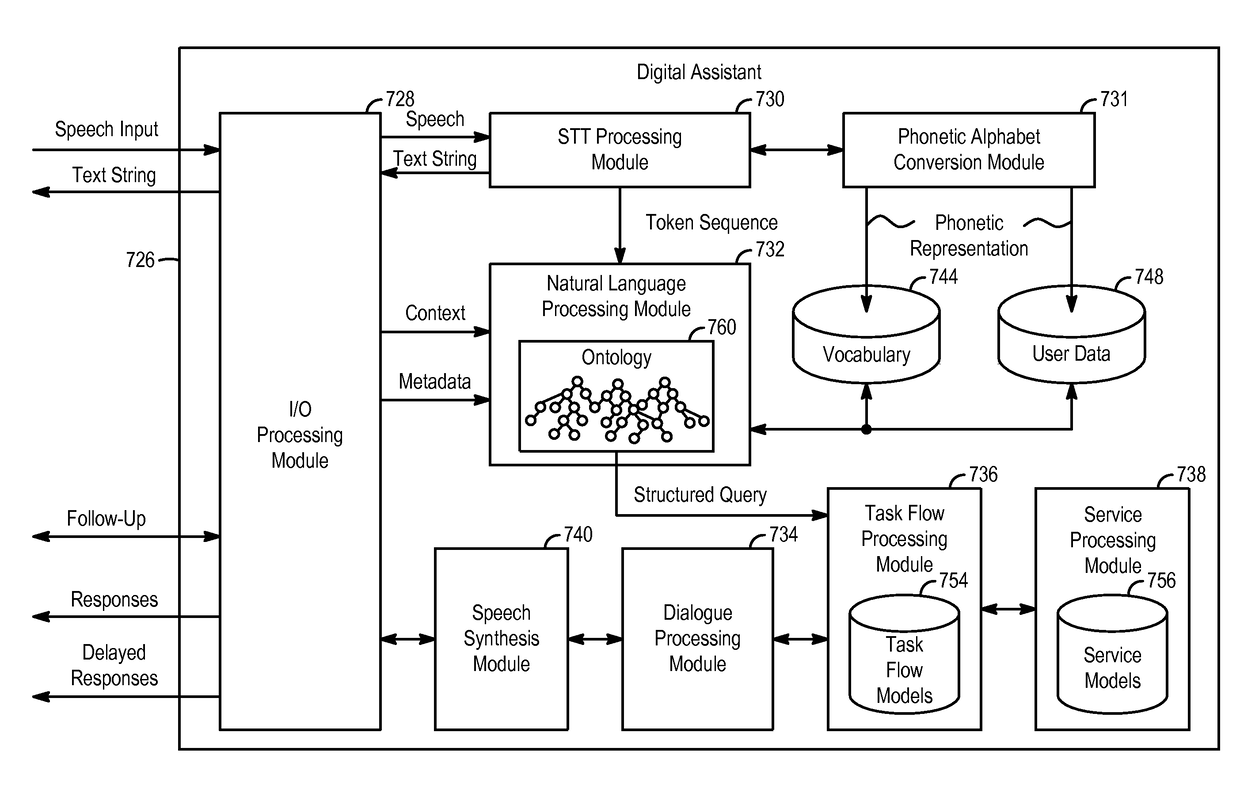
Systems and processes for language identification from short strings are provided. In accordance with one example, a method includes, at a first electronic device with one or more processors and memory, receiving user input including an n-gram and determining a similarity between a representation of the n-gram and a representation of a first language. The representation of the first language is based on an occurrence of each of a plurality of n-grams in the first language and an occurrence of each of the plurality of n-grams in a second language. The method further includes determining whether the similarity between the representation of the n-gram and the representation of the first language satisfies a threshold.
Owner:APPLE INC
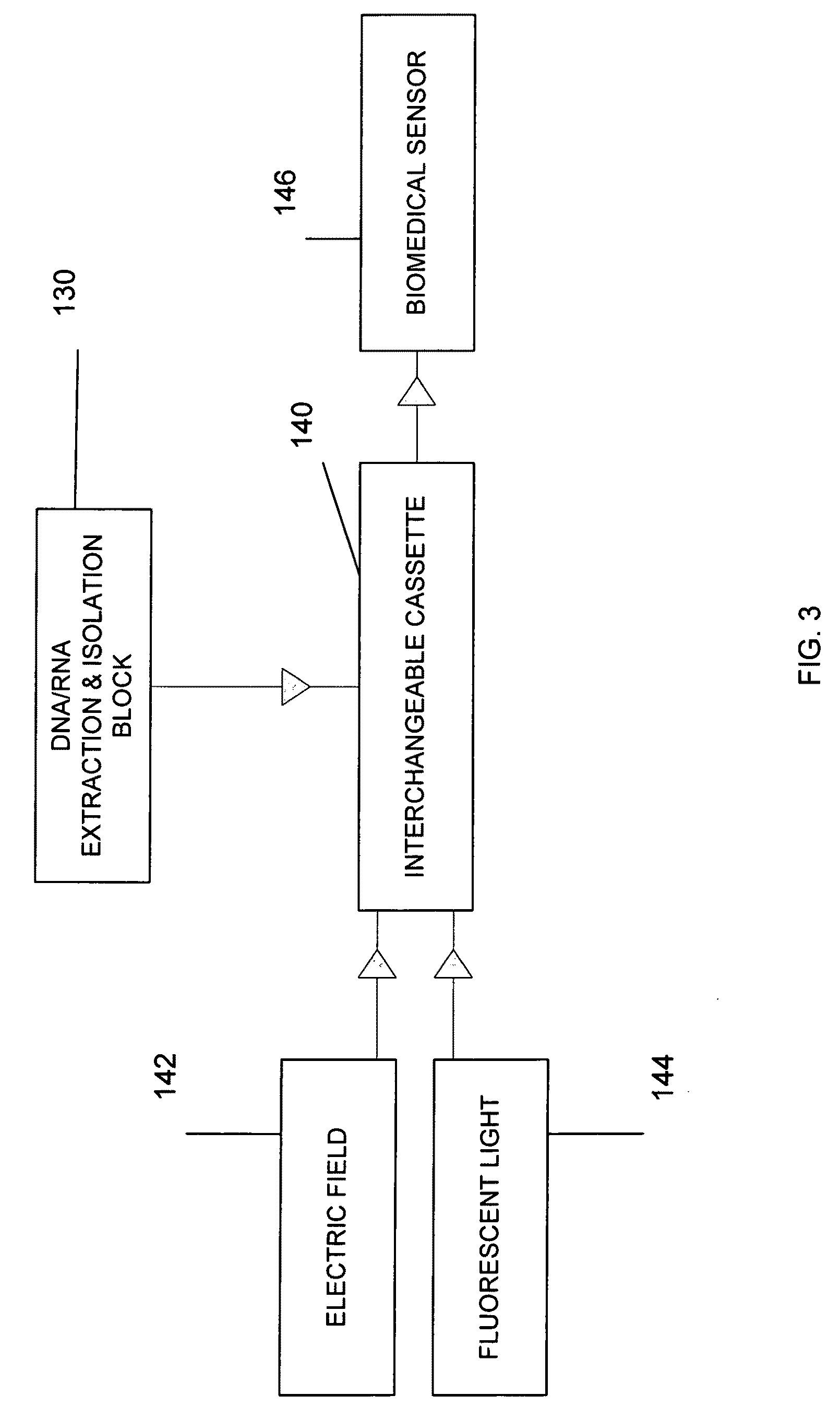
Direct identification and measurement of relative populations of microorganisms with direct DNA sequencing and probabilistic methods
ActiveUS8478544B2Quick identificationMicrobiological testing/measurementBiostatisticsGenomic SegmentProbabilistic method
Owner:COSMOSID INC
Direct identification and measurement of relative populations of microorganisms with direct DNA sequencing and probabilistic methods
ActiveUS20120004111A1Quick identificationMicrobiological testing/measurementLibrary screeningGenomic SegmentProbabilistic method
The present invention relates to systems and methods capable of characterizing populations of organisms within a sample. The characterization may utilize probabilistic matching of short strings of sequencing information to identify genomes from a reference genomic database to which the short strings belong. The characterization may include identification of the microbial community of the sample to the species and / or sub-species and / or strain level with their relative concentrations or abundance. In addition, the system and methods may enable rapid identification of organisms including both pathogens and commensals in clinical samples, and the identification may be achieved by a comparison of many (e.g., hundreds to millions) metagenomic fragments, which have been captured from a sample and sequenced, to many (e.g., millions or billions) of archived sequence information of genomes (i.e., reference genomic databases).
Owner:COSMOSID INC
Genome identification system
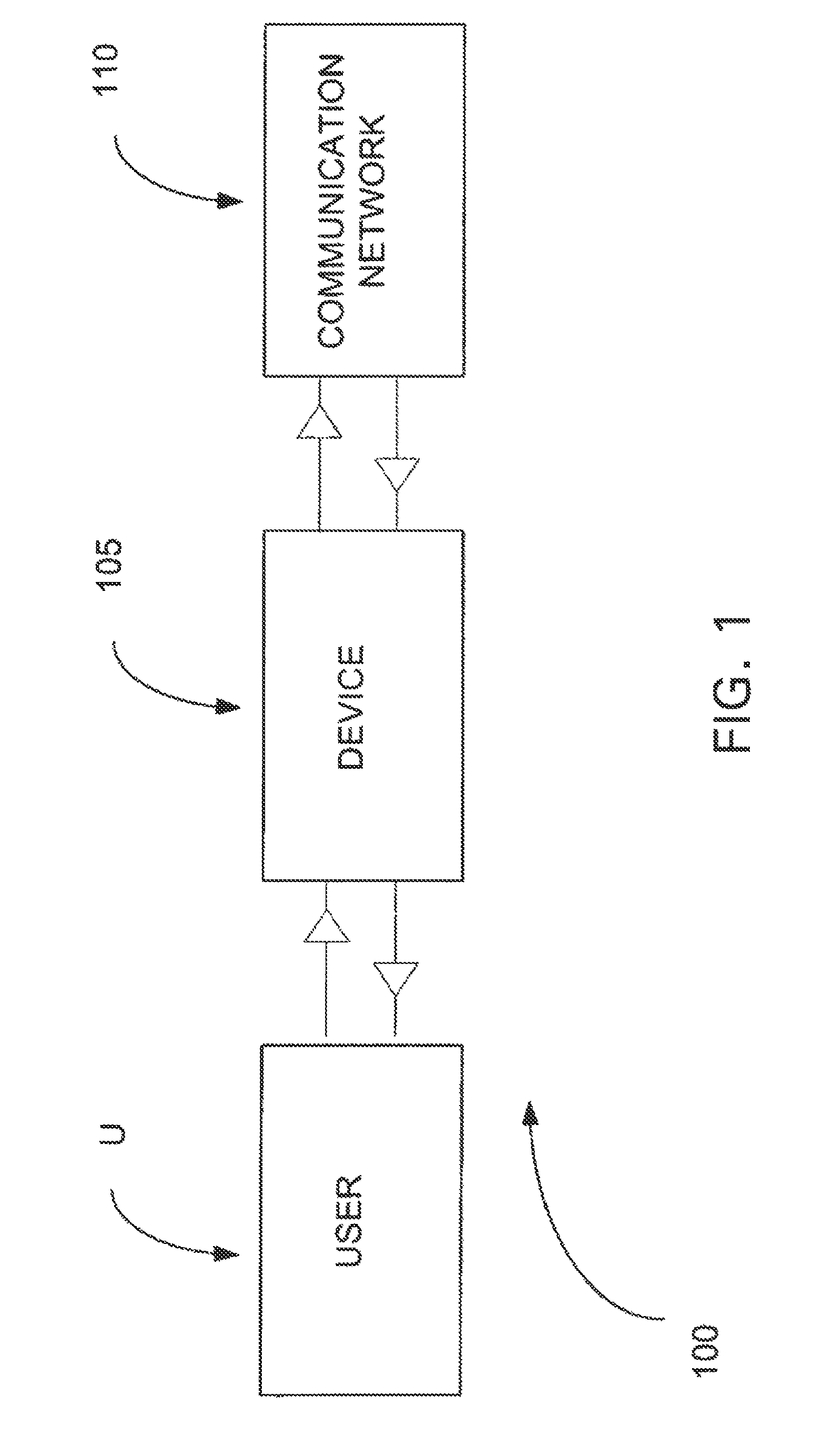
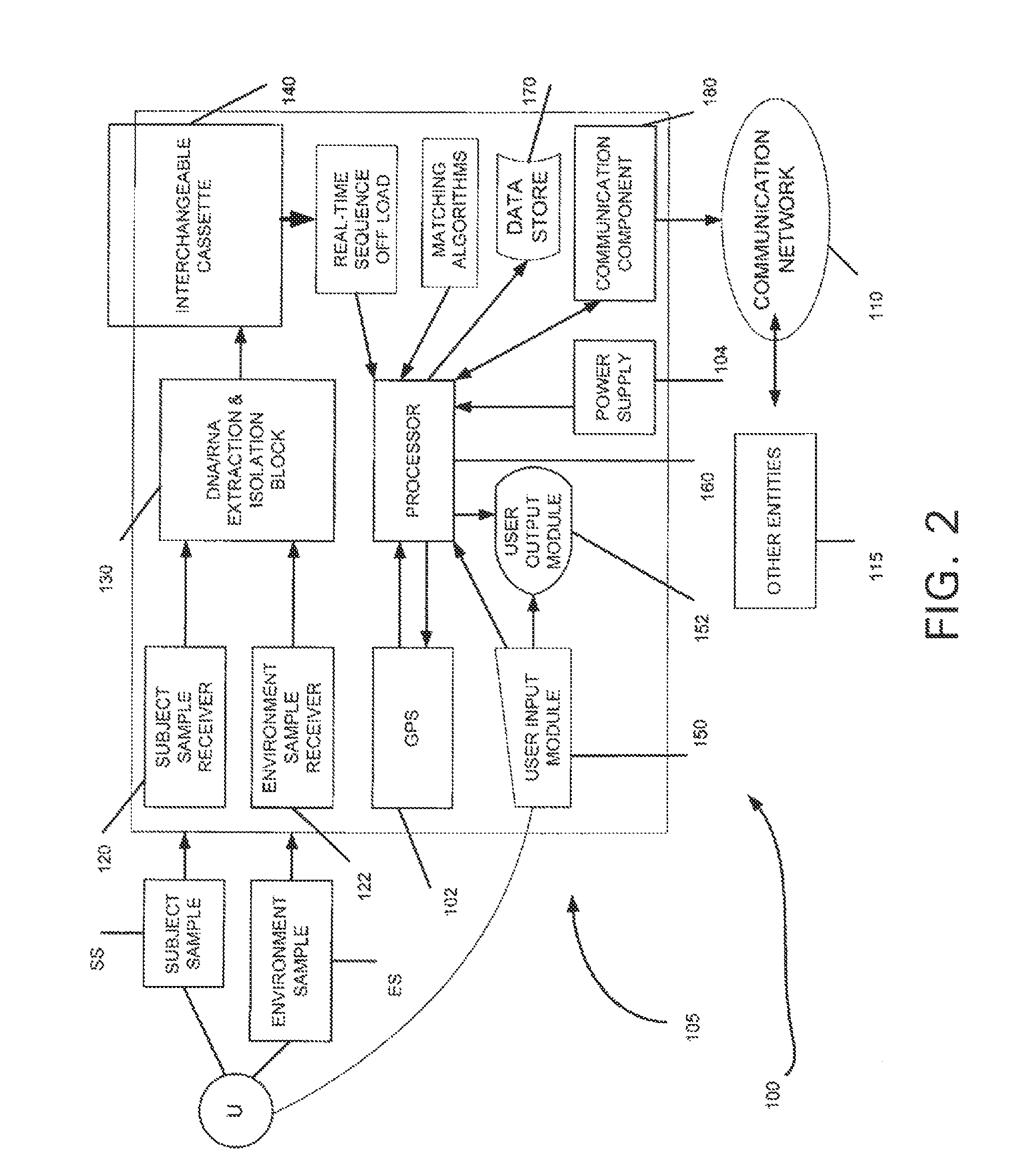
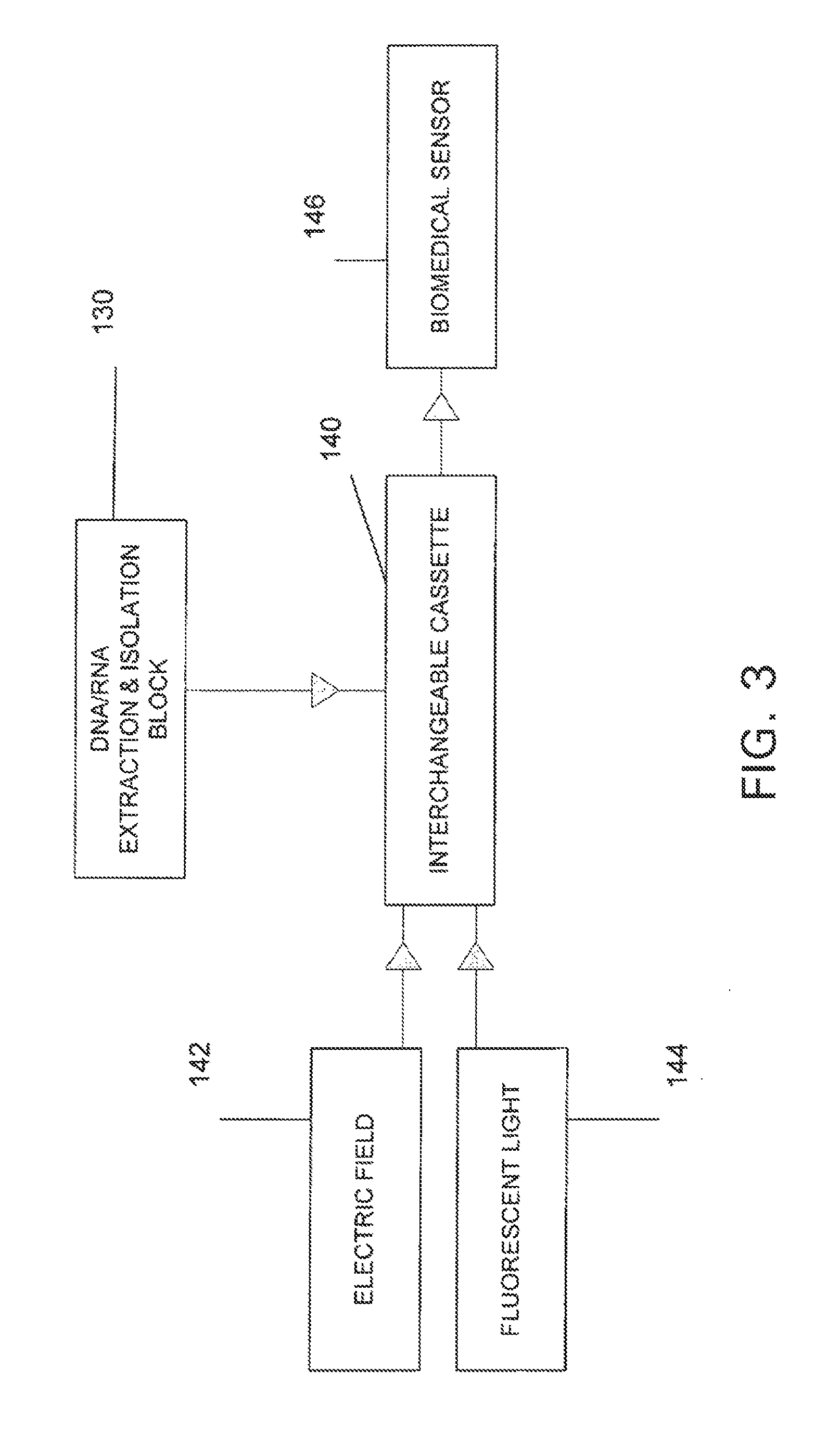
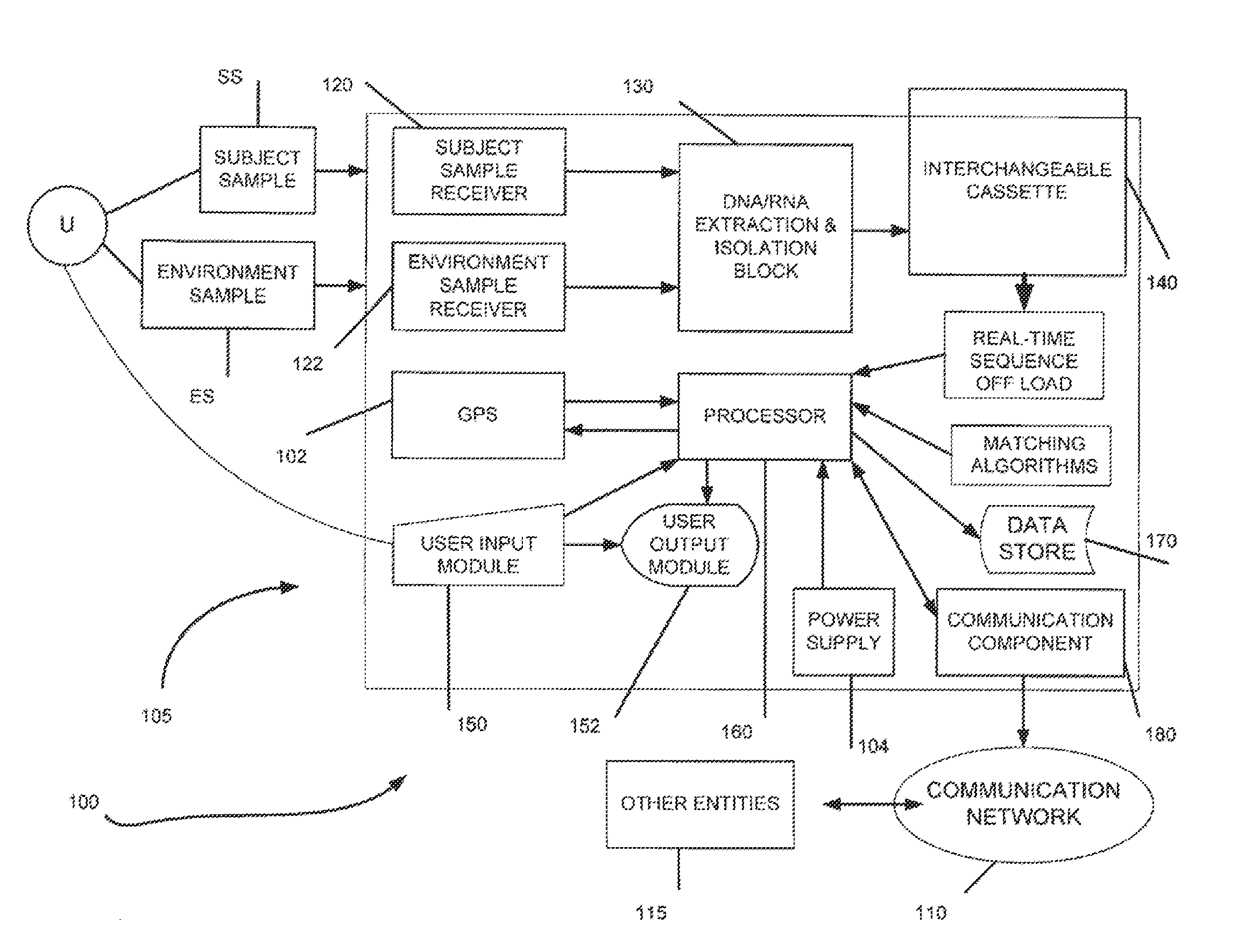
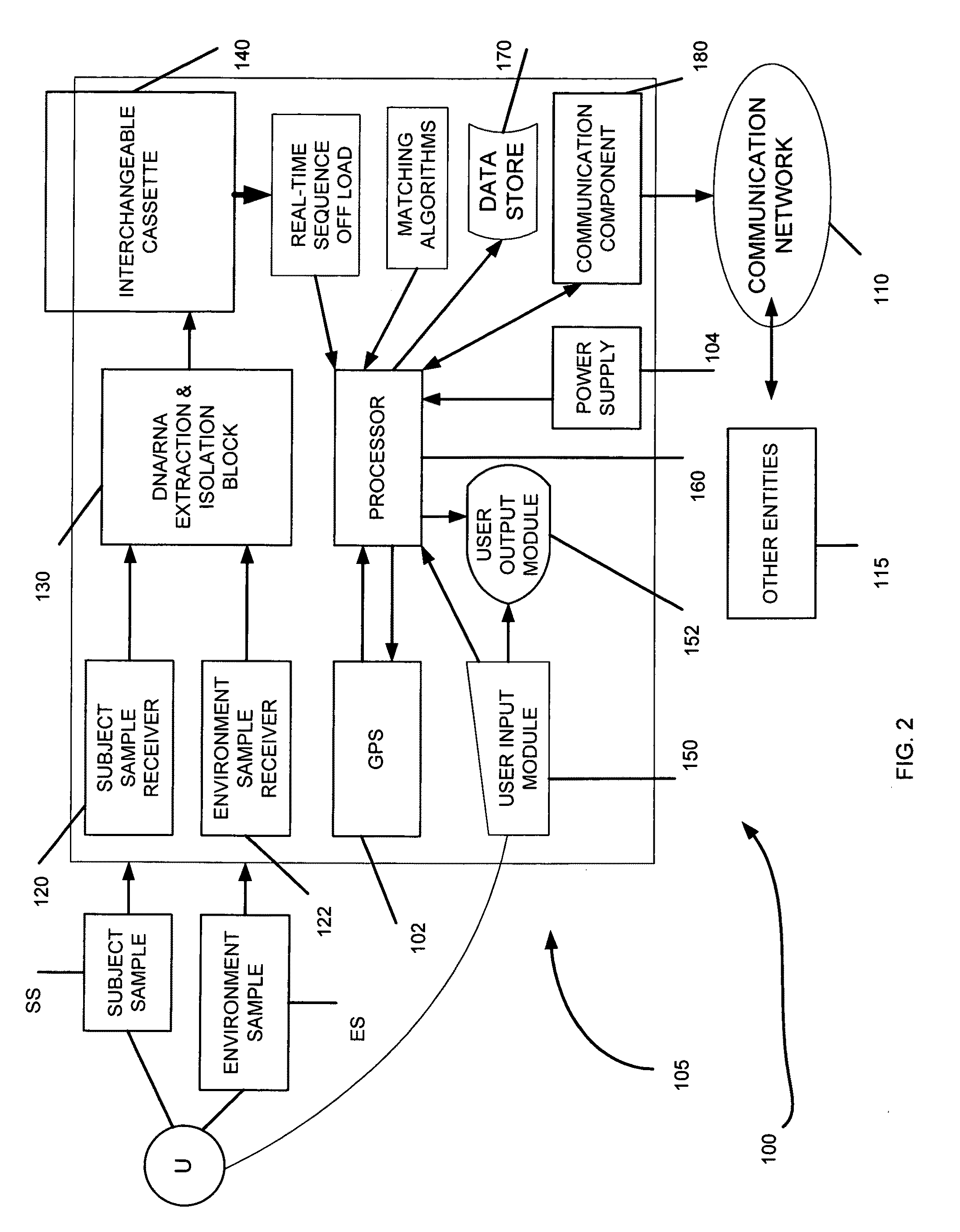
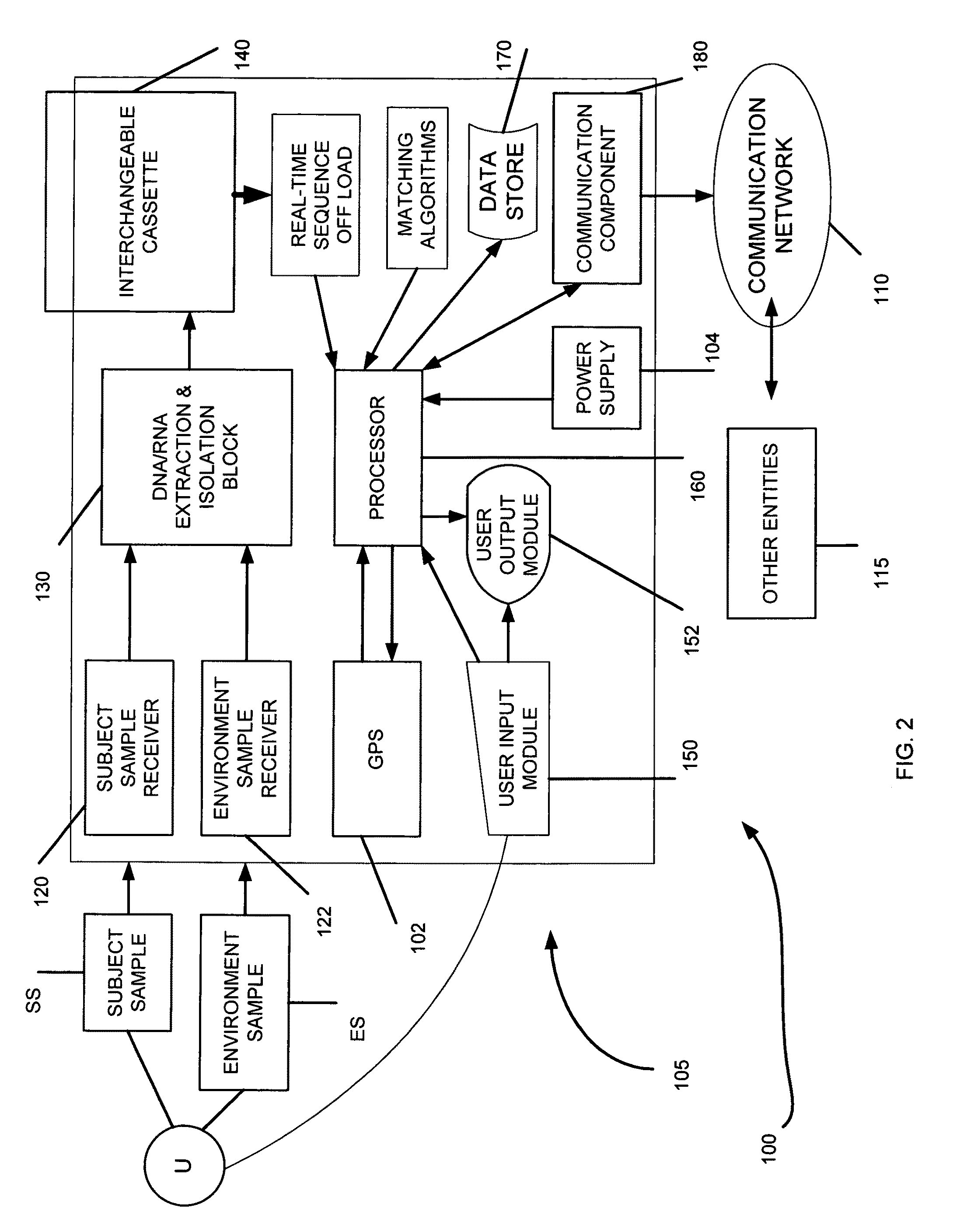
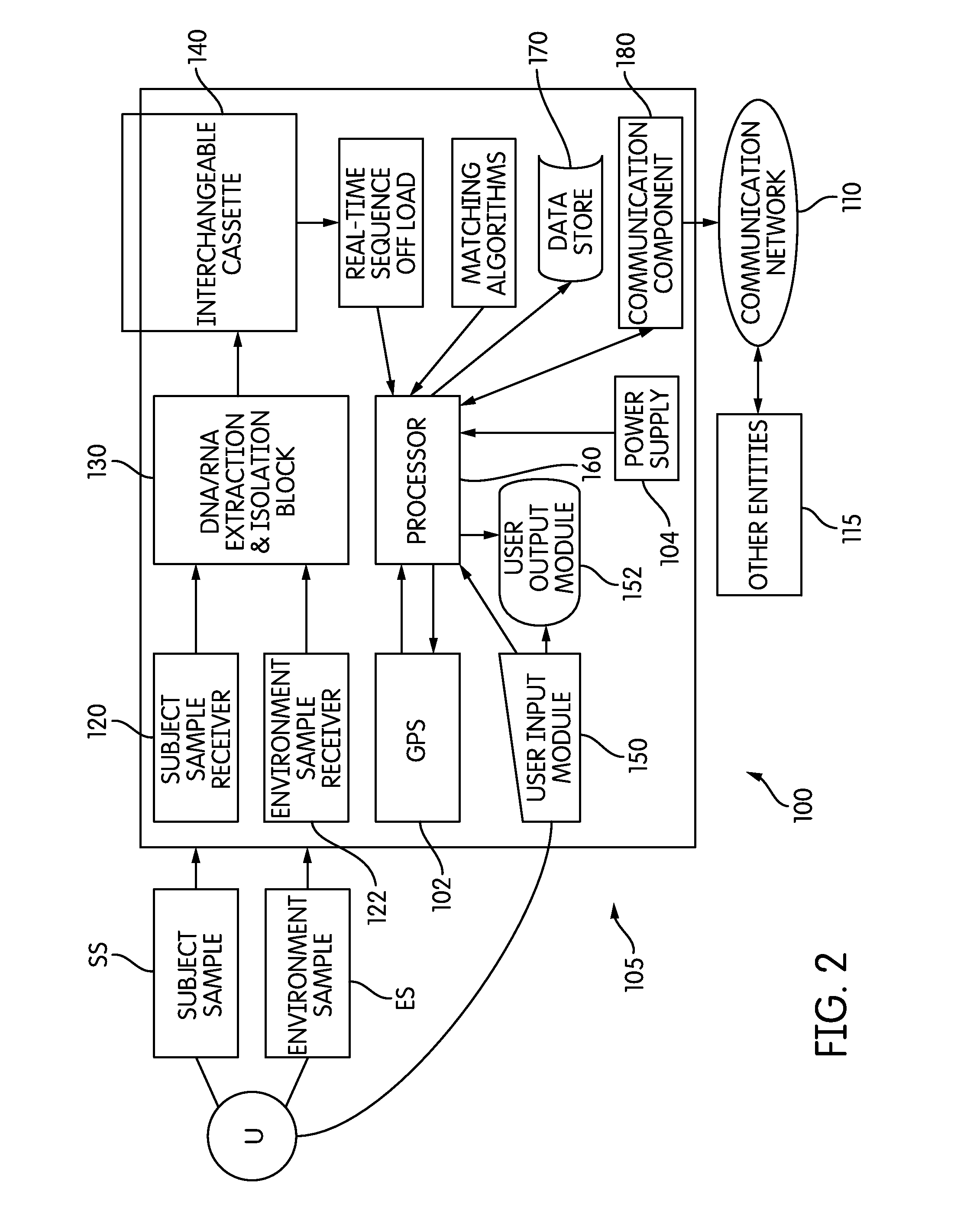
The present invention belongs to the field of genomics and nucleic acid sequencing. It involves a novel method of sequencing biological material and real-time probabilistic matching of short strings of sequencing information to identify all species present in said biological material. It is related to real-time probabilistic matching of sequence information, and more particular to comparing short strings of a plurality of sequences of single molecule nucleic acids, whether amplified or unamplied, whether chemically synthesized or physically interrogated, as fast as the sequence information is generated and in parallel with continuous sequence information generation or collection.
Owner:COSMOSID INC
Language identification from short strings
Systems and processes for language identification from short strings are provided. In accordance with one example, a method includes, at a first electronic device with one or more processors and memory, receiving user input including an n-gram and determining a similarity between a representation of the n-gram and a representation of a first language. The representation of the first language is based on an occurrence of each of a plurality of n-grams in the first language and an occurrence of each of the plurality of n-grams in a second language. The method further includes determining whether the similarity between the representation of the n-gram and the representation of the first language satisfies a threshold.
Owner:APPLE INC
Method and system for genome identification
The present invention belongs to the field of genomics and nucleic acid sequencing. It involves a novel method of sequencing biological material and real-time probabilistic matching of short strings of sequencing information to identify all species present in said biological material. It is related to real-time probabilistic matching of sequence information, and more particular to comparing short strings of a plurality of sequences of single molecule nucleic acids, whether amplified or unamplied, whether chemically synthesized or physically interrogated, as fast as the sequence information is generated and in parallel with continuous sequence information generation or collection.
Owner:COSMOSID INC
Query to task mapping
InactiveUS20050262058A1Great mapping qualityQuality improvementMetadata text retrievalDigital data processing detailsShort stringTask mapping
Owner:MICROSOFT TECH LICENSING LLC
Two strings private key (symmetric) encryption and decryption method
InactiveUS20100202606A1Easy and secure and affordable meanData stream serial/continuous modificationSecret communicationPlaintextShort string
Two strings encryption algorithm where a long and a short string are used. The byte values of the short string points to a location of the long string and the plaintext is aligned with the long string's location and encryption is performed using the long string's byte values and the plaintext the process is repeated for all bytes of the short string pointing to the long string and aligned a byte to encrypt with the long string.
Owner:PILOANDIA LLC
System and Method for Matching Data Using Probabilistic Modeling Techniques
InactiveUS20140052688A1Penalizing the similarity scoreFuzzy logic based systemsSpecific program execution arrangementsProbit modelExact match
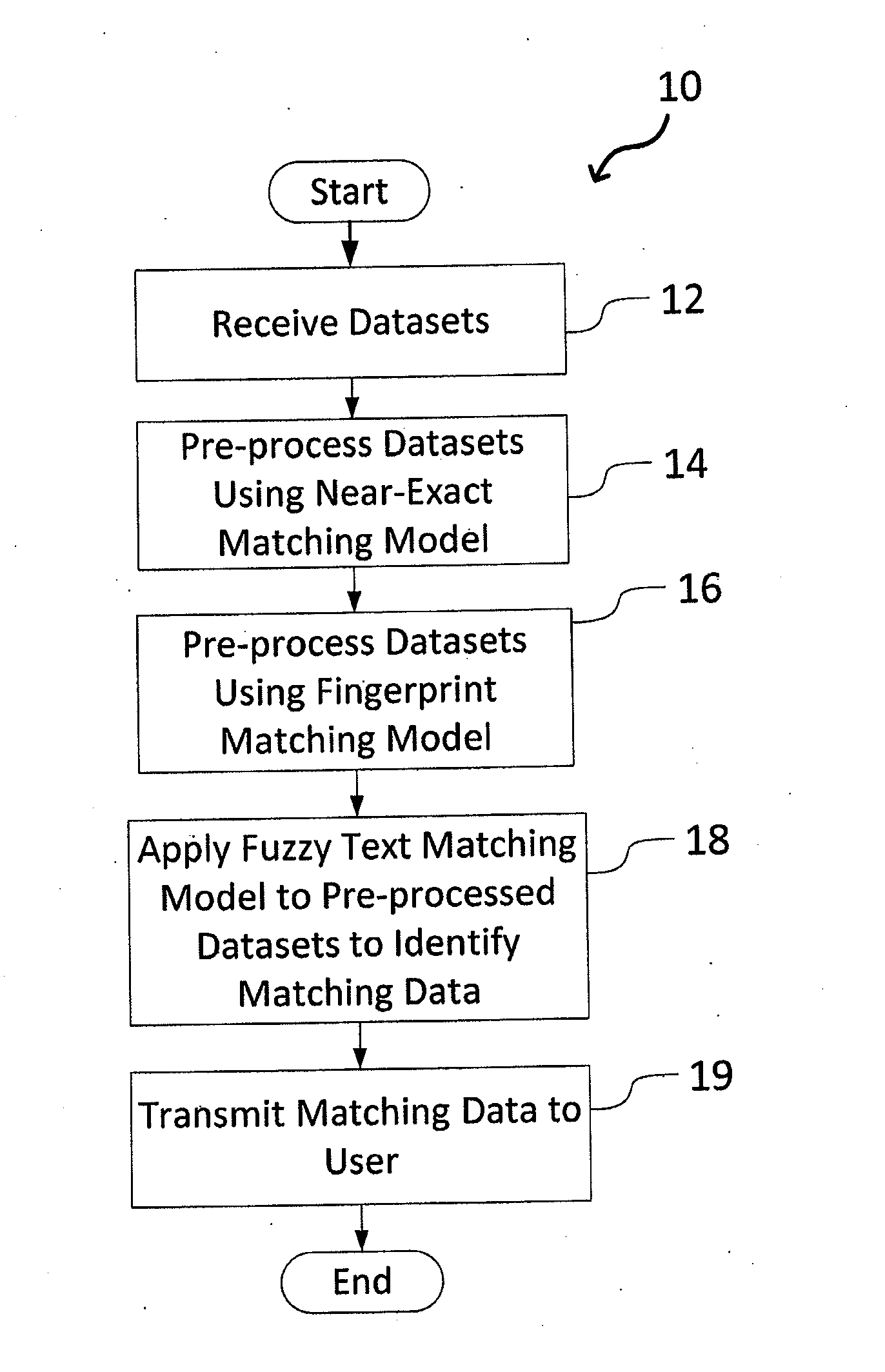
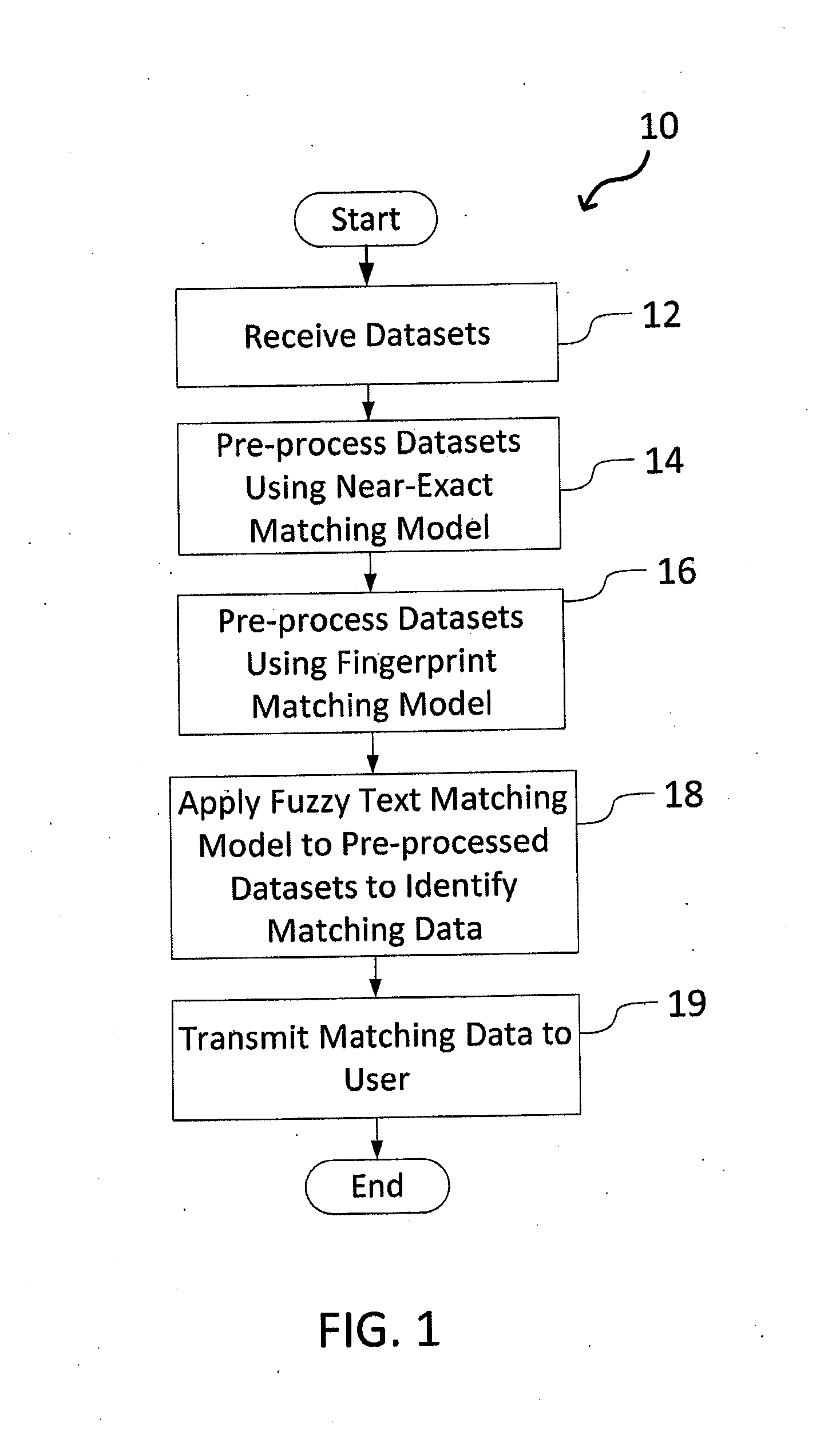
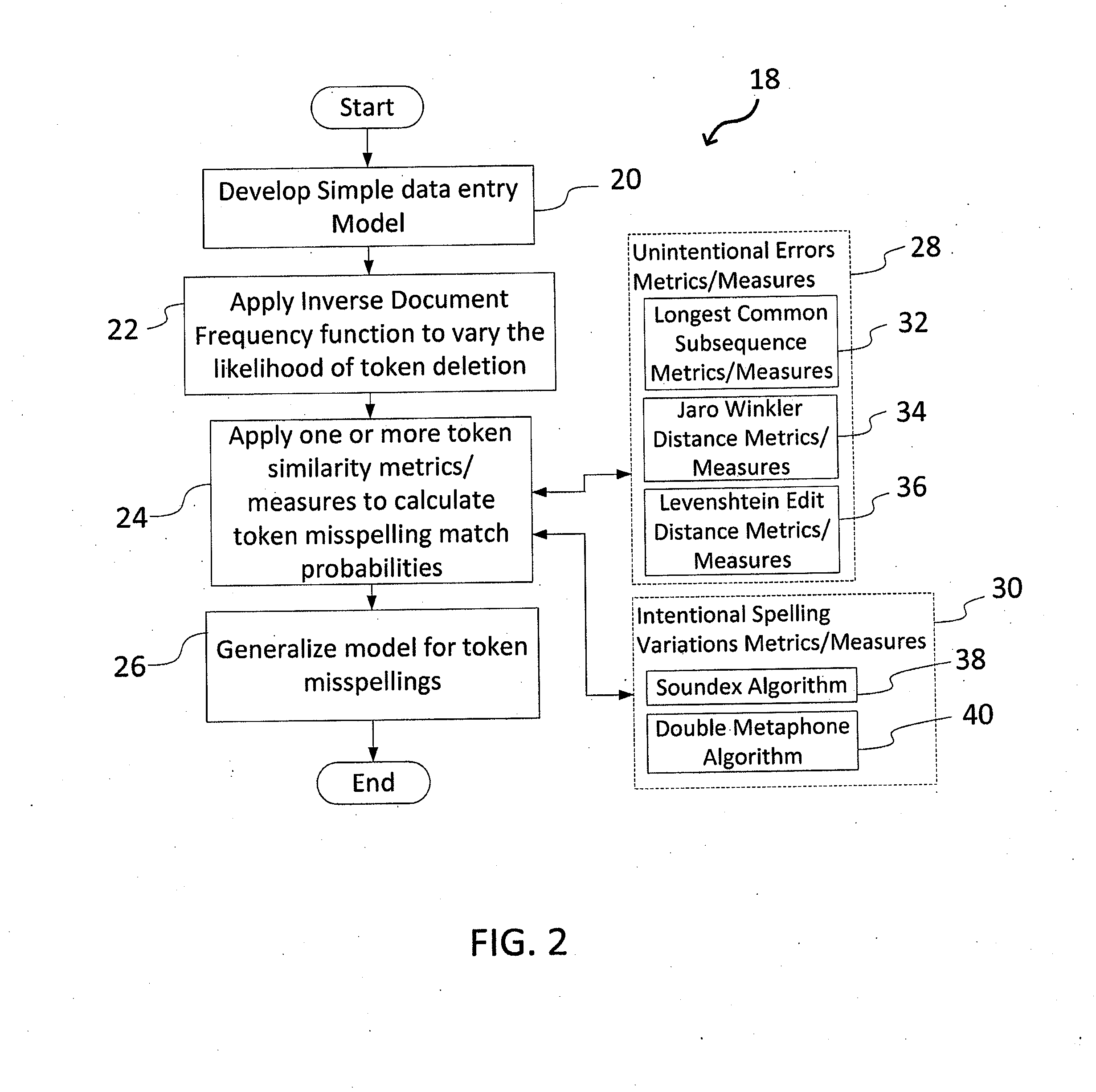
A system and method for matching data using probabilistic modeling techniques is provided. The system includes a computer system and a data matching model / engine. The present invention precisely and automatically matches and identifies entities from approximately matching short string text (e.g., company names, product names, addresses, etc.) by pre-processing datasets using a near-exact matching model and a fingerprint matching model, and then applying a fuzzy text matching model. More specifically, the fuzzy text matching model applies an Inverse Document Frequency function to a simple data entry model and combines this with one or more unintentional error metrics / measures and / or intentional spelling variation metrics / measures through a probabilistic model. The system can be autonomous and robust, and allow for variations and errors in text, while appropriately penalizing the similarity score, thus allowing dataset linking through text columns.
Owner:OPERA SOLUTIONS U S A LLC
Method and system for drawing construction in short sequence assembly
ActiveCN101430742ASmall footprintHigh speedMicrobiological testing/measurementSequence analysisShort stringConnection number
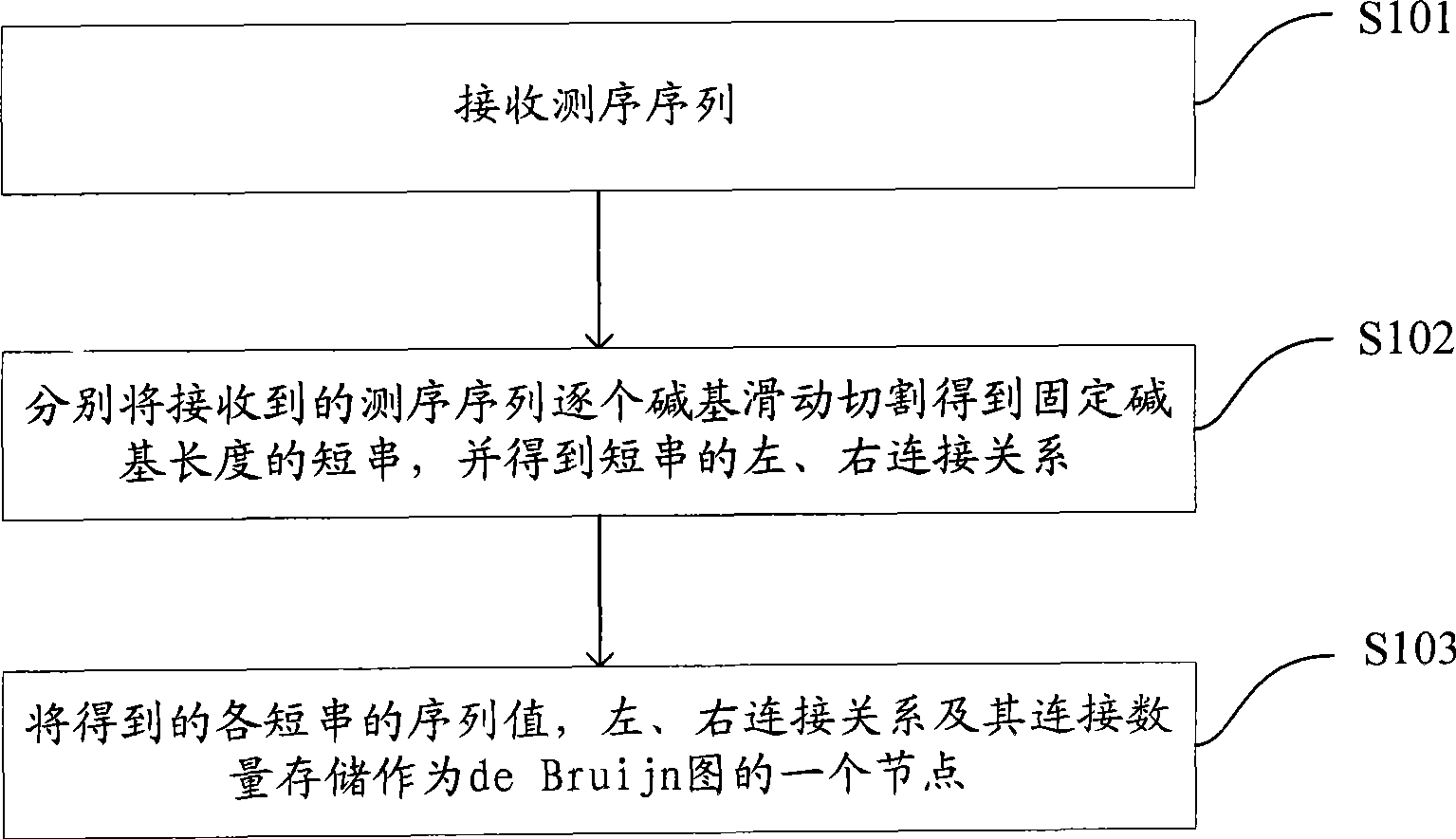
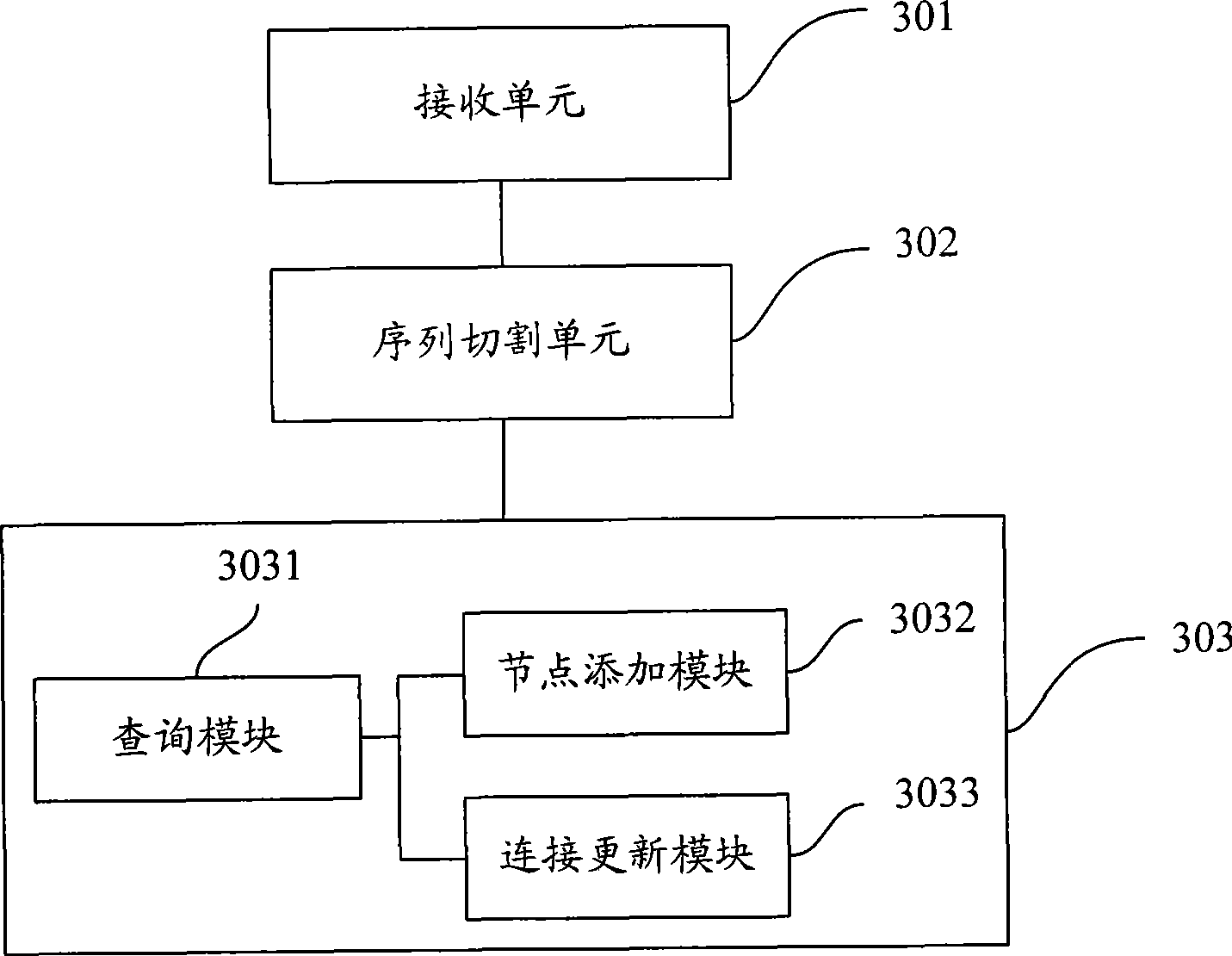
The invention is applicable to the technical field of gene engineering, and provides a method for constructing a graph in a short sequence assembly and a system thereof. The method comprises the following steps: receiving an order-checking sequence; carrying out sliding cutting on each base of the received order-checking sequence to obtain a short string with a fixed base length and a left and right connecting relation of the short string; storing a sequence value of the obtained short string, the left and right connecting relation and a connection number as a node of a de Bruijn graph. In the invention, the method for constructing the graph in the short sequence assembly can be realized by slidingly cutting the base of the received order-checking sequence one by one to obtain the short string with the fixed base length and the left and right connecting relation of the short string, and storing the sequence value of the obtained short string, the left and right connecting relation and the connection number as the node of the de Bruijn graph. The method can assemble a large genome with small occupied memory and fast speed.
Owner:BGI TECH SOLUTIONS
Method for carrying out harmful content recognition on network text and short message service
InactiveCN101876968AImprove efficiencyHigh speedSpecial data processing applicationsShort stringShort Message Service
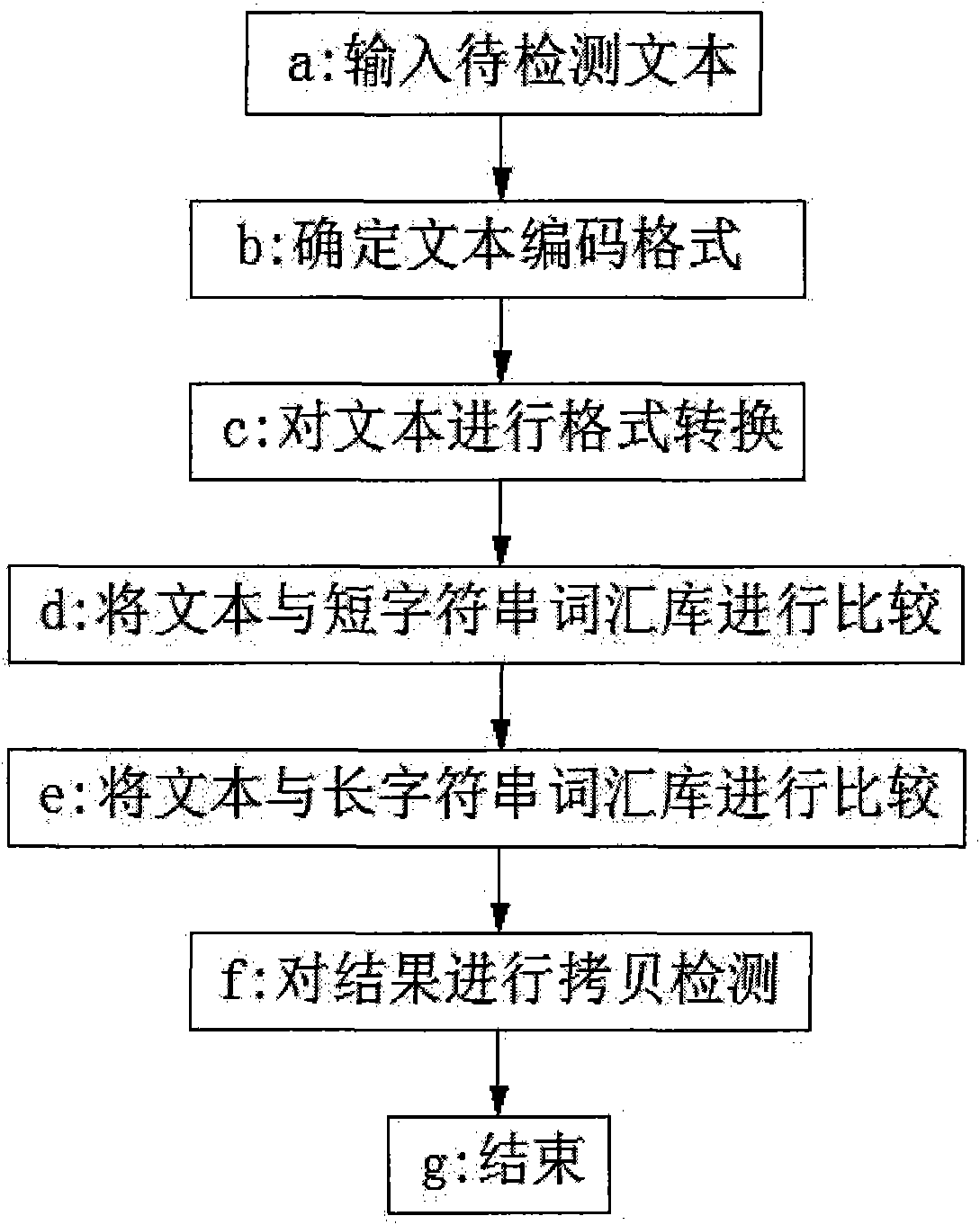
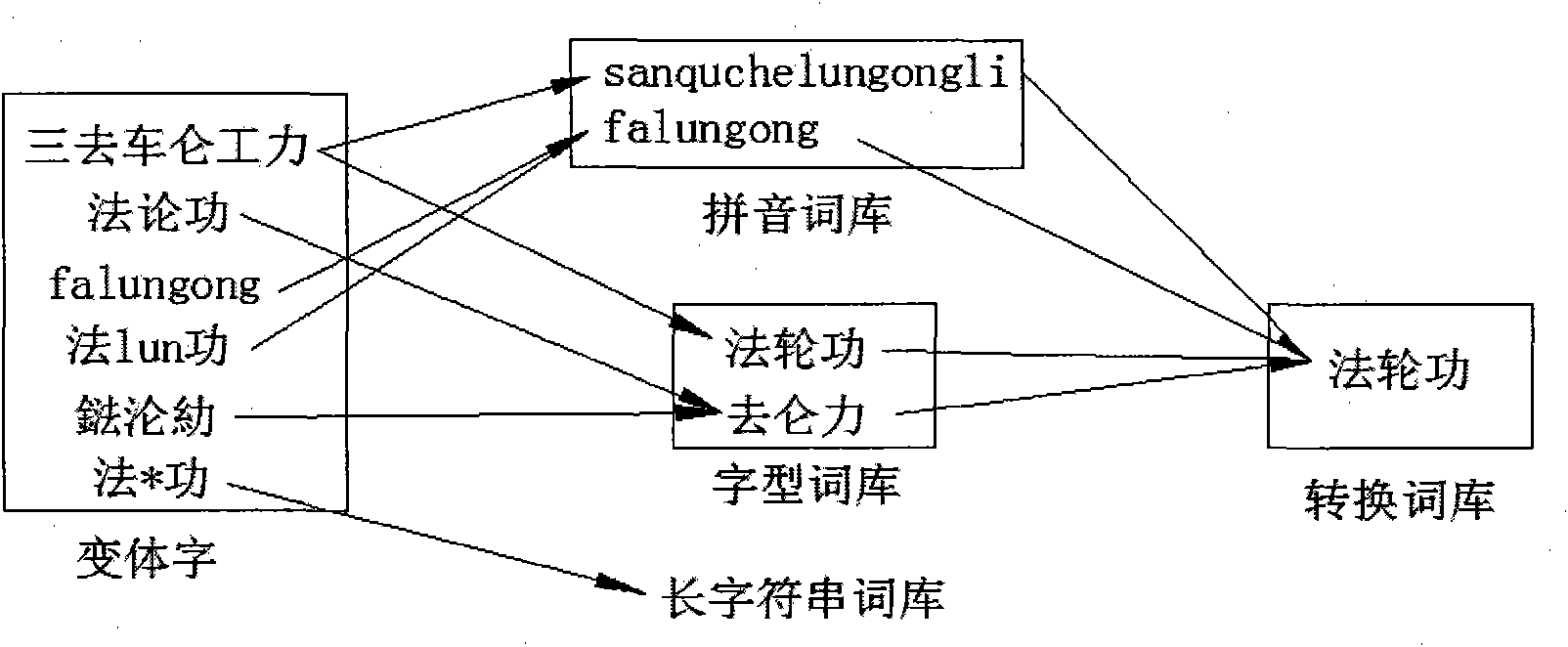
The invention belongs to the technical field of text processing, in particular to a method for carrying out harmful content recognition on network text and short message service, which comprises the following steps of: inputting a text to be detected, determining a text coding format, carrying out format conversion on the text, comparing the text with a short string word bank, comparing the text with a long string word bank, carrying out copy detection on a result, and displaying a final result. The method can be used for the detection and the filtration on harmful, violent and reactionary texts in the internet, inhibits the spreading of the harmful content, and protects physical and psychological health of youngsters.
Owner:FUDAN UNIV
A rapid fuzzy matching algorithm for strings in mass audio data
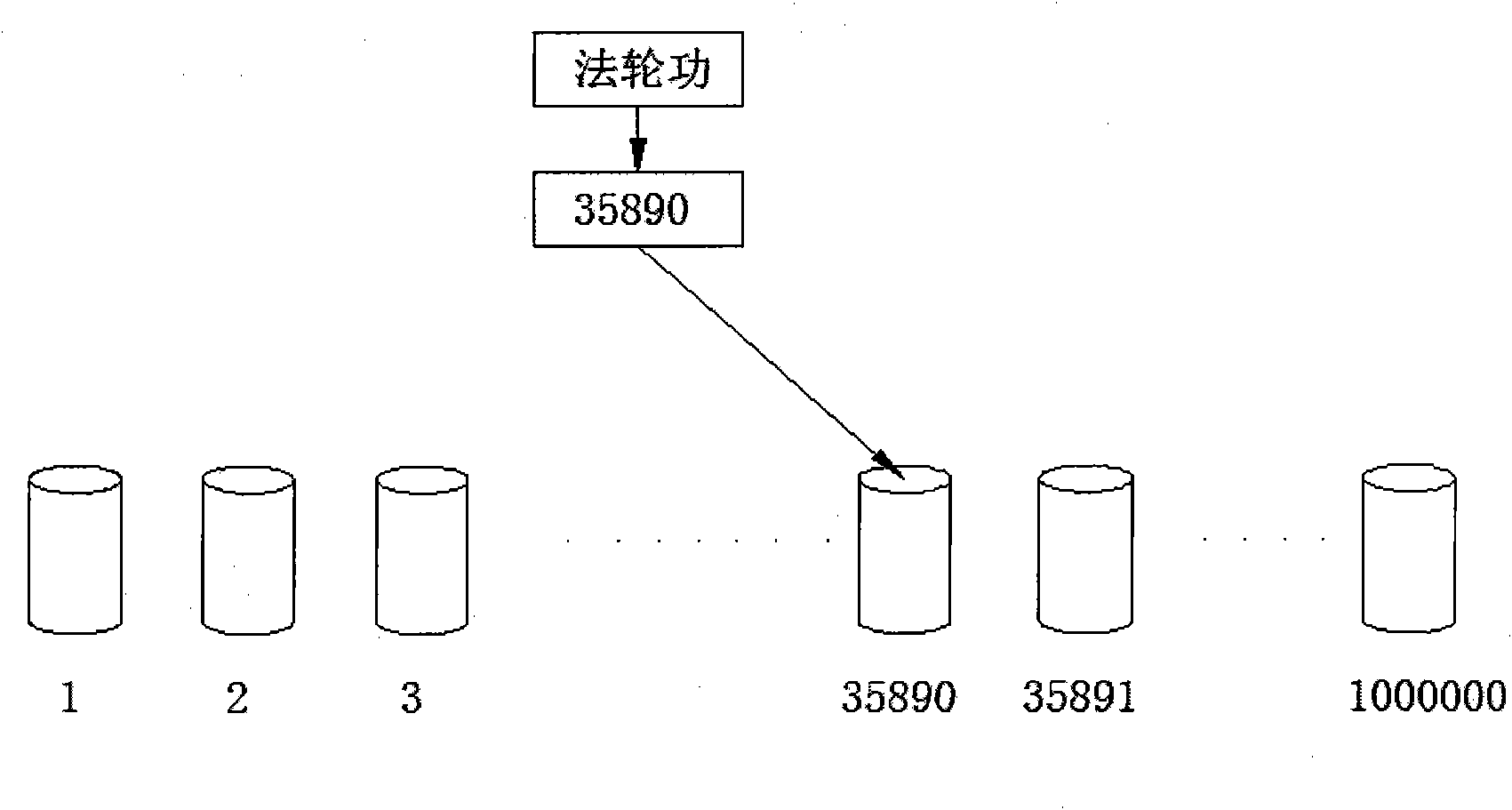
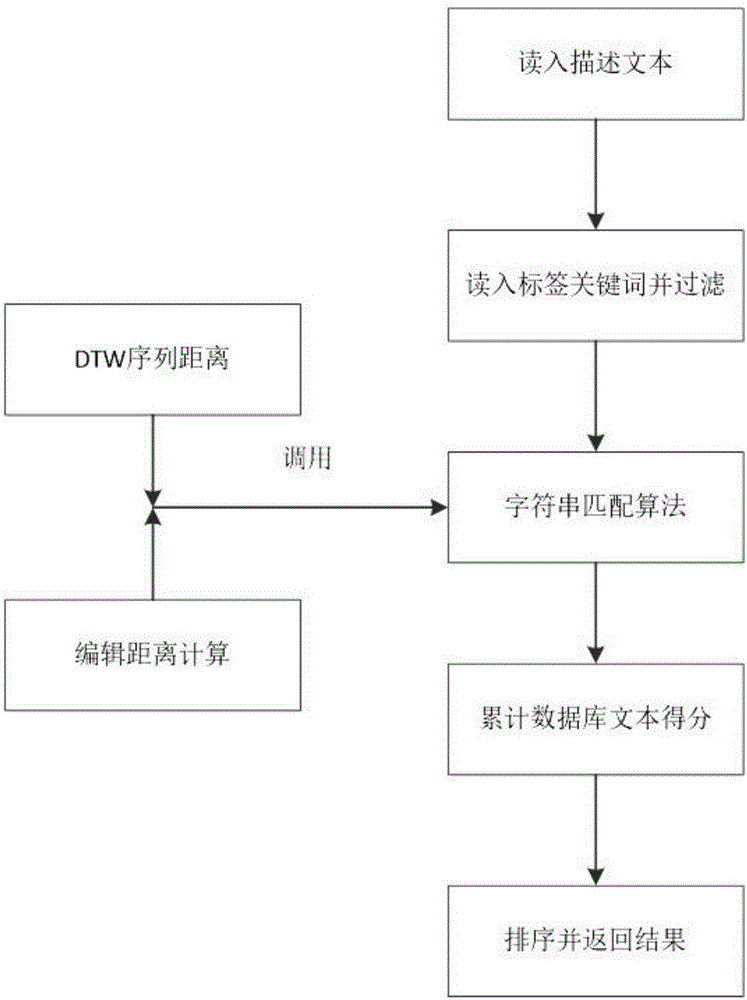
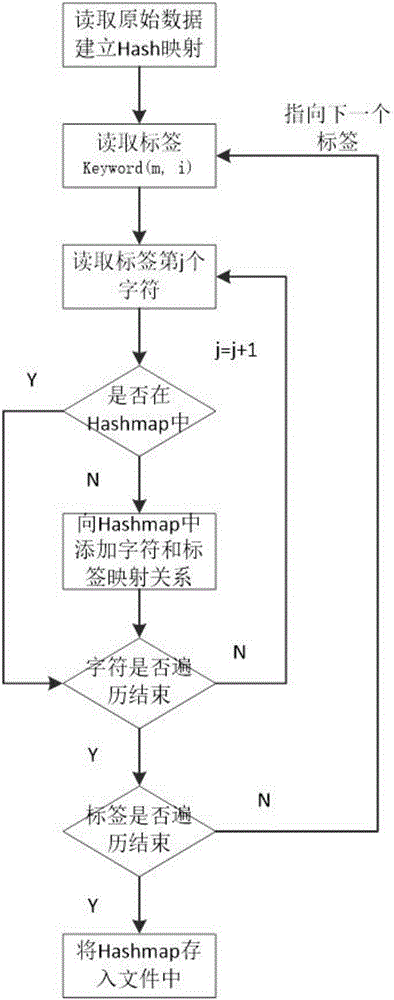
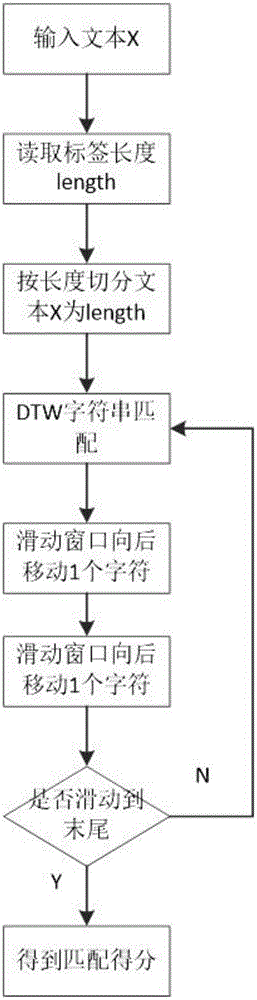
InactiveCN106528599ASupport searchSupport matchingSpecial data processing applicationsChinese charactersShort string
The invention provides a rapid fuzzy matching algorithm for strings. According to the invention, firstly data preprocessing is performed on texts in a database to obtain a statistical model and an index is established via Hash. An input text is a shorter string. The algorithm traverses all Chinese characters therein, activates the positions of corresponding Chinese characters in a finite character complete set, and maps the activation state of the finite character complete set to each tag to filter tags. A few filtered tags are used for matching the texts and the DTW algorithm is used for approximate string matching. The algorithm also comprises the steps of performing scoring and sorting according to the result of the degree of approximation of matching and returning to a search result. Through the efficient tag filtering method, the calculation efficiency of the string matching algorithm is greatly increased; in a process of input text matching, a fuzzy matching effect is achieved and a good matching performance is guaranteed for fuzzy languages.
Owner:深圳凡豆信息科技有限公司
Method and system for genome identification
The present invention belongs to the field of genomics and nucleic acid sequencing. It involves a novel method of sequencing biological material and real-time probabilistic matching of short strings of sequencing information to identify all species present in said biological material. It is related to real-time probabilistic matching of sequence information, and more particular to comparing short strings of a plurality of sequences of single molecule nucleic acids, whether amplified or unamplied, whether chemically synthesized or physically interrogated, as fast as the sequence information is generated and in parallel with continuous sequence information generation or collection.
Owner:COSMOSID INC
Multi-string matching method
InactiveCN101901257AMax jump distance increasedWill not missSpecial data processing applicationsShort stringTheoretical computer science
The invention relates to a multi-string matching method, belonging to the technical field of string matching. The invention separates long strings from short strings in a rule set based on the conventional Wu-Manber method and further processes the long strings and short strings in the rule set in different ways when a SHIFT table is created, thus ensuring the maximum table entry of the SHIFT table to be free from the limit of the length of the short strings and overcoming the disadvantage that the maximum skipping distance of the maximum table entry is limited by the length of the shortest string in the rule set; and by introducing the HOT table and using the method for HOT search in the matching process, the invention increases the maximum skipping distance of the window without skipping the short strings. The method of the invention achieves higher matching efficiency.
Owner:BEIJING INSTITUTE OF TECHNOLOGYGY
Short sequence mapping method and system
InactiveCN101430741AExtended processing timeLength prefix short processing timeMicrobiological testing/measurementSpecial data processing applicationsContigShort string
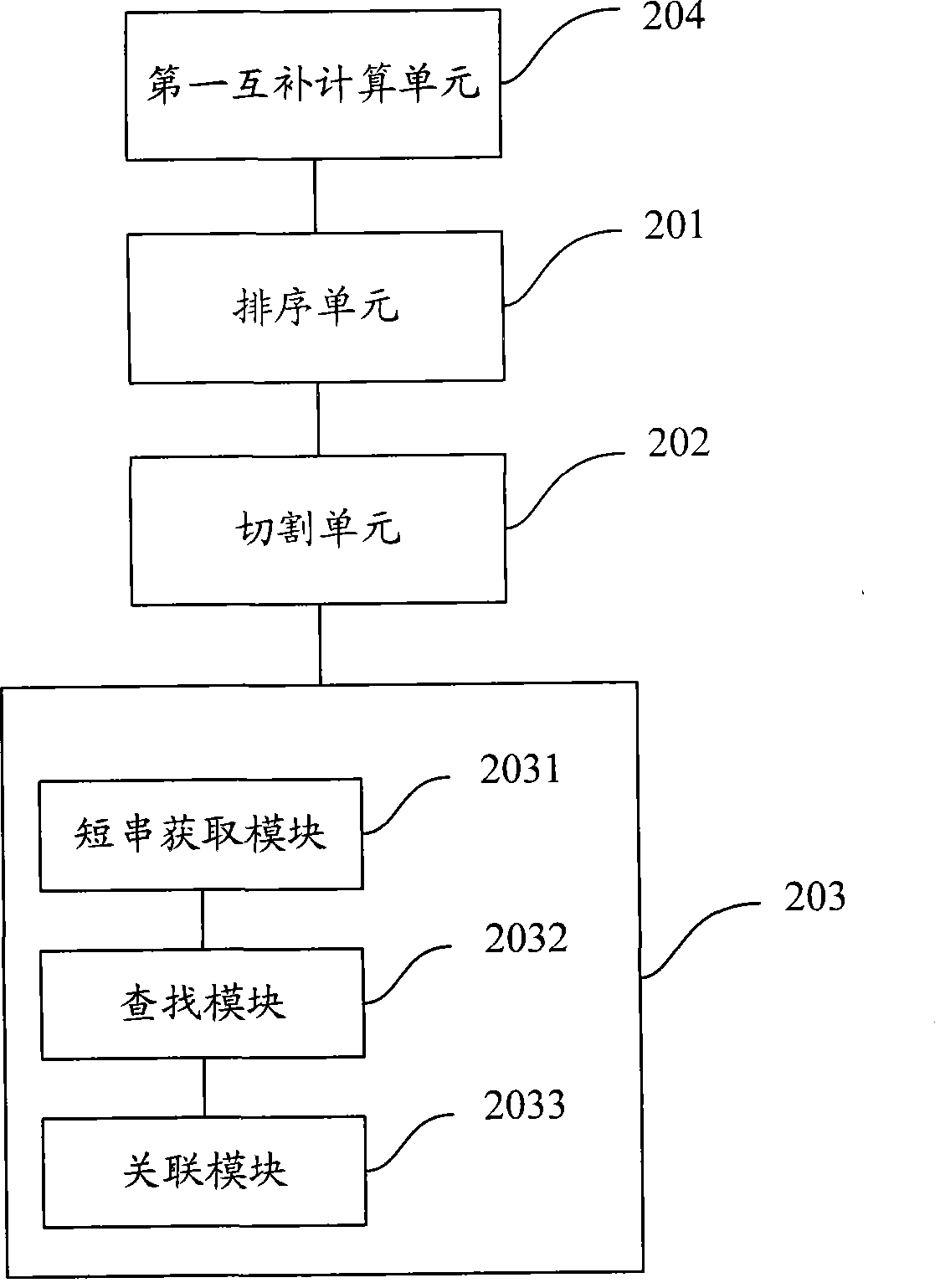
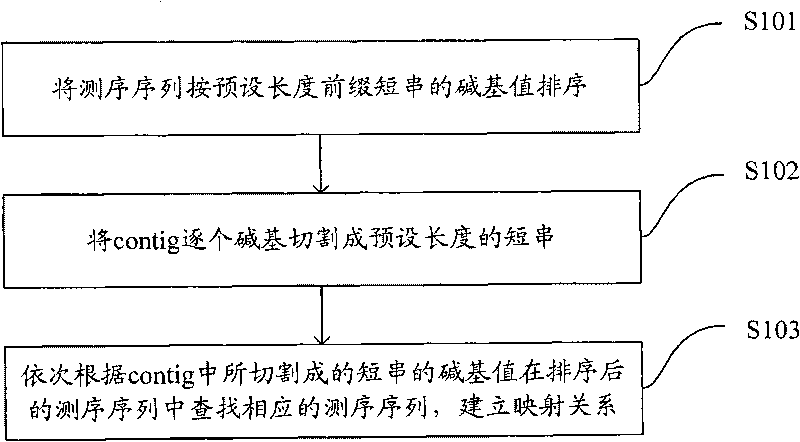
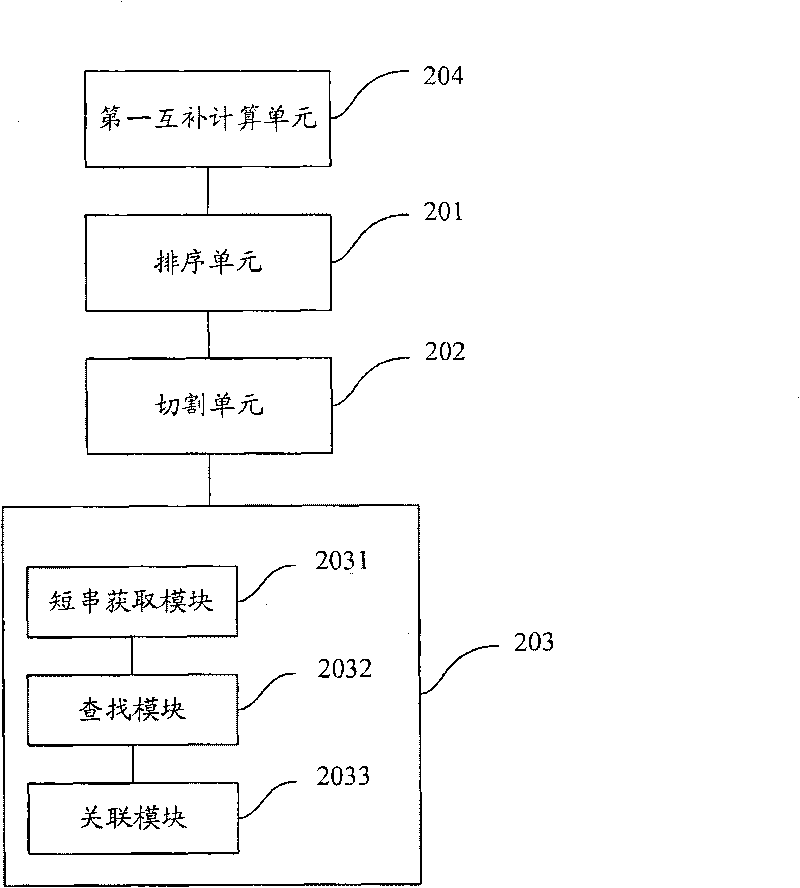
The invention is applicable to the technical field of gene engineering, and provides a method for mapping a short sequence and a system thereof. The method comprises the following steps: ordering an order-checking sequence according to base values of prefixed short strings with predetermined length; cutting each base of a contig to a short string with the predetermined length; searching a corresponding order-checking sequence in an ordered order-checking sequence in sequence according to the base value of the cut short string in the contig so as to establish a mapping relation. In the invention, the method for mapping the short sequence used in a short sequence assembly is realized by ordering the order-checking sequence according to the base values of the prefixed short strings with the predetermined length, cutting each base of the contig to the short string with the predetermined length and searching the corresponding order-checking sequence in the ordered order-checking sequence in sequence according to the base value of the cut short string in the contig so as to establish the mapping relation. Therefore, the method has short treatment time and high efficiency.
Owner:SHENZHEN HUADA GENE INST
Method and system for fast processing genome short sequence mapping
ActiveCN101751517AReduce processing timeImprove efficiencyMicrobiological testing/measurementSpecial data processing applicationsContigShort string
Being applicable to the technical field of genetic engineering, the invention provides a method and a system for fast processing genome short sequence mapping, comprising the following steps: ranking sequencing sequence according to base number of short strings of preset length; cutting basic groups of sequence contig into short strings of preset length; searching corresponding sequencing sequence in ranked sequencing sequence according to base number of short strings cut from the sequence contig; then establishing mapping relation. In the invention, the sequencing sequence is ranked according to base number of short strings of preset strings and basic groups of sequence contig are cut into short strings of preset length; in addition, the corresponding sequencing sequence in ranked sequencing sequence is searched according to base number of short strings cut from the sequence contig; finally mapping relation is established; so that short sequence mapping applied to short sequence assembling is realized, processing time is short and processing efficiency is high.
Owner:BGI TECH SOLUTIONS
Method and apparatus for identifying conversation in multiple strings
Techniques for identifying conversations in multiple short strings include determining from a first plurality of strings associated with a first contact of a user, based on time separations between successive strings, a first conversation portion and a different second conversation portion. The first conversation portion (snippet) comprises a plurality of strings of the first plurality; and the second snippet comprises a different pluralty of strings of the first plurality. A first semantic content for the first snippet and a second semantic content for the second snippet are determined. It is determined whether to merge the first snippet and the second snippet into a first conversation that includes the first snippet based, at least in part, on a similarity of the first semantic content to the second semantic content.
Owner:NOKIA TECHNOLOGLES OY
Keyword extraction method and device
ActiveCN107885717AImprove accuracyNatural language data processingSpecial data processing applicationsShort stringMorpheme
The embodiment of the invention provides a keyword extraction method and device. The method comprises the steps that multiple candidate morphemes are extracted from a to-be-extracted document, and theimportance of each candidate morpheme is calculated based on a morpheme importance model; permutation and combination are performed on the candidate morphemes according to preset rules, multiple candidate short strings are generated, and the integrity of each candidate short string is calculated based on a short string integrity model; candidate morphemes in a first quantity are selected from thecandidate morphemes according to the order of the importance; candidate short strings in a second quantity are selected from the candidate short strings according to the order of the integrity; and the candidate morphemes in the first quantity and the candidate short strings in the second quantity are determined as keywords of the to-be-extracted document. By the adoption of the keyword extraction method and device, the morphemes with high importance and the short strings with high integrity in the to-be-extracted document are extracted, and therefore the accuracy of the extracted keywords isimproved.
Owner:TENCENT TECH (SHENZHEN) CO LTD
Search algorithm based on DNA k-mer index problem four-node list trie tree
InactiveCN106484865ASave node spaceEasy to returnSpecial data processing applicationsNODALRound complexity
The invention relates to the field of data structures and big data processing, in particular to a novel quick search algorithm based on a trie tree, comprising: establishing a four-node trie tree model, and using four bases of a DNA sequence as system inputs; establishing a trie tree terminal search list, determining a terminal end mark, not distinguishing base sequences, and establishing a model for reversely deducting sequence numbers and base pair numbers upon query; establishing a DNA sequence index and analyzing its complexity; acquiring positions of substrings, hooking a search list to leaf sub-node, and storing position data; querying k-mer short strings, and analyzing their complexity. The longer a common prefix of a word, the higher the query speed of the trie tree; the complexity varies with k differences, is substantially a constant and is nearly not affected by data quantity. Letter mapping is applied to original data, 26 sub-nodes of the trie tree are decreased to 4, and node space is saved.
Owner:HARBIN ENG UNIV
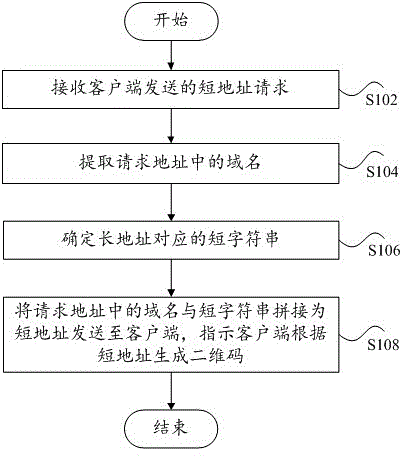
Quick response (QR) code generating method and response method of QR code scanning event
The invention discloses a QR code generating method, device and system and a response method, device and system of a QR code scanning event. The QR code generating method comprises that a short address request is received from a client, parameters transmitted by the short address request includes a long address, a domain name of the address requested by the short address request is one selected from multiple domain names, and the multiple domain names all direct to the address of a service end; the domain name is extracted from the request address; a short string corresponding to the long address is determined; and the domain name in the request address is spliced with the short strong to form a short address, the short address is sent to the client, and the client is indicated to generate a QR code according to the short address. According to the invention, the domain name of the short address of the generated QR code can change with the domain name in the requested address of the short address request and thus, is not unique.
Owner:BEIJING UNION VOOLE TECH
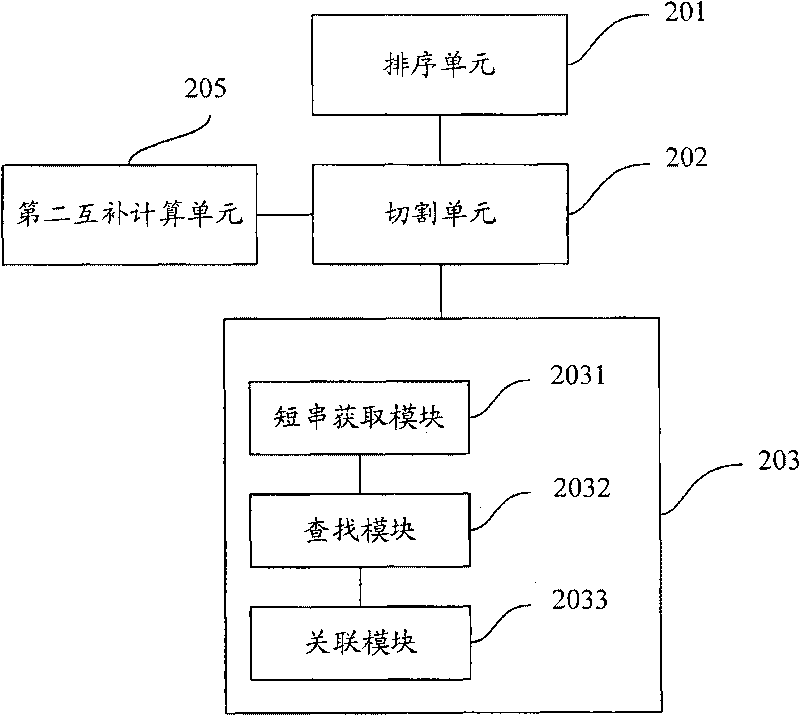
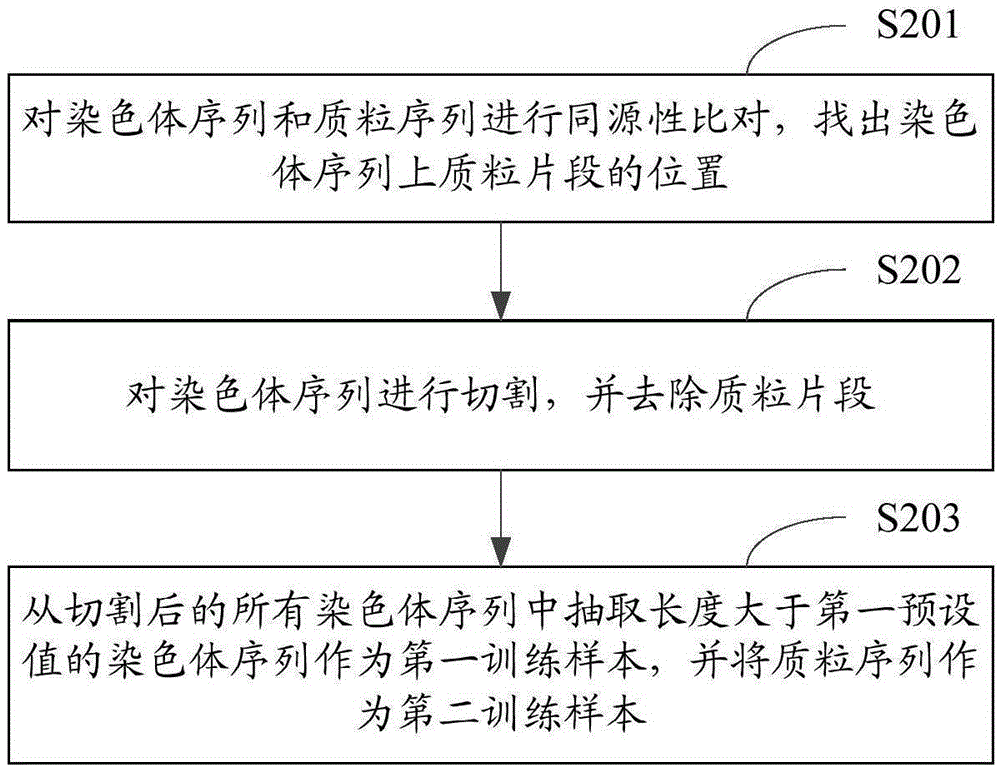
Method and device for classifying chromosome sequences and plasmid sequences
ActiveCN105631464AImprove accuracyImprove training efficiencyRecognition of DNA microarray patternAlgorithmShort string
The invention is applicable to the technical field of data mining and provides a method and a device for classifying chromosome sequences and plasmid sequences. The method comprises steps: chromosome sequences and plasmid sequences are acquired, and a first training sample and a second training sample are obtained; frequency characteristics of all k character short strings and reverse complementary sequence pairs thereof are extracted and a first frequency characteristic table and a second frequency characteristic table are generated, wherein k is no less than 2 but no more than 5; a training set and a test set are extracted from the first frequency characteristic table and the second frequency characteristic table, and a chi-square test algorithm is adopted to calculate weight values of all characteristic data in the training set; a random forests algorithm is adopted and according to the characteristic data whose weight values meet preset conditions, a classification model is trained; and according to the classification model, the chromosome sequences and the plasmid sequences are classified. Thus, the training efficiency and the training effects of the classification model are improved, and accuracy on classification on the chromosome sequences and the plasmid sequences is improved.
Owner:SHENZHEN INST OF ADVANCED TECH
LED light string circuit having a plurality of short strings connected as long string
InactiveCN102196614AAvoid flickeringImplement overvoltage protectionElectric light circuit arrangementEnergy saving control techniquesPower flowShort string
The invention discloses a device, which comprises a first input end for connecting an alternating power source; a second input end for connecting an alternating power source, wherein the first input end and the second input end have an input pressure difference; a rectifier connected with the first input end and the second input end; a first light string with at least one light emitting diode (LEDs); and a second light string with at least one LEDs, wherein a first current flows via the first light string but does not flow via the second light string within a first time interval; a second current flows via the first and second light strings within a second time interval, and the input pressure difference between the first input end and the second input end is higher in the second time interval than in the first time interval.
Owner:ACTIVE SEMI SHANGHAI +1
Short-string parallel-dc optimizer for photovoltaic systems
ActiveUS20160285264A1Photovoltaic supportsSingle network parallel feeding arrangementsElectricityShort string
This disclosure generally relates to an energy generation system. In one embodiment, the energy generation system comprises a plurality of solar panels that are connected in a series electrical connection. The energy generation system further includes a short-string optimizer which outputs direct current electricity to a direct current bus.
Owner:ZYNTONY INC
Implementation method of short dynamic code and application thereof
ActiveCN103425797AHigh precisionEnhance memorySpecial data processing applicationsShort stringTheoretical computer science
The invention discloses an implementation method of a short dynamic code and application thereof. The implementation method is based on a distributed storage database, and relates to a long string and a short string differing in digit length. The method comprises a code decreasing step for mapping the long string with the short string and a code returning step for returning the short string into the long string, wherein the short string is valid within a certain period of time in the two steps. A time stamp is added into the record storage in the database, and a judgment on whether failure time is surpassed is made, so that a mapping relation of the short dynamic code is generated accurately. After the application of the implementation method disclosed by the invention, the failure time is designed, and a mapping correspondence relation is established between the long string and the short string which is switched dynamically by rolling and is relatively short within a certain duration, so that convenience is brought to memory of accounts and the safety and practicability are enhanced via the short dynamic code; moreover, due to the introduction of the time stamp, the advantages of data disaster tolerance and system scale of the distributed storage database are brought into full play, and the accuracy of the short dynamic code is increased.
Owner:北京通付盾人工智能技术有限公司
Method of two strings private key (symmetric) encryption and decryption algorithm
InactiveUS20080165965A1Easy and secure and affordable meanData stream serial/continuous modificationPublic key for secure communicationPlaintextShort string
Two strings encryption algorithm where a long and a short strings are used. The byte values of the short string points to a location of the long string and the plaintext is aligned with the long string and encryption is performed using the long string's byte values and the plaintext the process is repeated for all bytes of the short string. The short string defines the encryption strength by pointing to the long string encrypting at first and re-encrypting thereafter. At the end, once the encryption is finished, the short key byte's values are used and once again pointing to the long string and removing a byte from the location and the removed byte(s) form a third string. The process will repeat until all the bytes values from the first string are used and a third string of equal length as of the short string is formed. Finally, the third string will perform the same process and its byte's values are used to point the long string and insert a byte of the short string into the location where they're pointing to, the process will repeat until all bytes of the short string are inserted into the long string. The long and short (third) strings are now unbalanced and the third (short) string becomes the private content's key. The reverse process is used to remove the short string, insert the third string and have two balanced string and decryption can be performed thereafter.
Owner:UNOWEB
Method and device for extracting keywords in page
ActiveCN104679731AFix not workingImprove versatilitySpecial data processing applicationsShort stringWord search
The invention discloses a method and a device for extracting keywords in a page. The method comprises the following steps: performing character string analysis on the title content of the page to obtain candidate words, and constructing a candidate word search table by the obtained candidate words; performing page analysis on the page to obtain a character combination, and constructing a short string set by the obtained character combination; performing character string analysis on the short string set to obtain character strings, and constructing an original weight pool by the obtained character strings; performing weighted voting on the candidate words in the candidate word search table through the character strings according to the sequence of the quantities of words included in each character string in the original weight pool, and increasing the weight values of the candidate words if the character strings are consistent with the candidate words in the candidate word search table; sequencing according to the weighted values of the candidate words from large to small, and extracting a preset quantity of candidate words in the front as keywords according to the sequence. By adopting the method and the device, the universality of a keyword extraction technology can be enhanced, and a way for extracting the keywords is more intelligent and efficient.
Owner:BAIDU ONLINE NETWORK TECH (BEIJIBG) CO LTD
Retrieval system and method for continuous characters and fuzzy characters
InactiveCN106446062AImprove portabilityFast retrievalSpecial data processing applicationsAlgorithmShort string
The invention provides a retrieval system and method for continuous characters and fuzzy characters. The retrieval system comprises a continuous character string mode matching module, wherein a KMP character string mode matching algorithm is used for matching the continuous characters in a short string with a long string; when the character string mode matching module fails in matching, the retrieval system furthermore comprises a fuzzy character matching module used for taking out Unicodes one by one from the long string to be matched with the short string, and the characters corresponding to the Unicodes are Chinese characters or ASCII characters. Through the retrieval system and method, matching retrieval of the continuous characters and the fuzzy characters and polyphone retrieval of Chinese characters are achieved, wherein continuous character string mode matching is also suitable for retrieval of multi-linguistic characters, and fuzzy character matching is suitable for retrieval of Chinese characters or combinations of Chinese characters and ASCII characters; the Chinese character pinyin mapping table has good portability, can be used across platforms and is high in retrieval speed and efficiency.
Owner:HUIZHOU DESAY SV AUTOMOTIVE
Method for bit-byte synchronization in sampling a data string
InactiveUS7388938B2Modification of read/write signalsError detection/correctionShort stringData field
Bit and byte synchronization for sampling and decoding a data string is provided a single data field u. The data string x has pre-pended to it a short string of 1s (ones), followed by u to yield a string y= . . . 1111, u, x. The string is pre-coded by convolution with 1 / (1⊕D2). PRML-sampling of y starts at an initial phase, and vectors are obtained from that string by sampling at pre-selected phases following the initial sampling point. The vectors of y are compared with vectors corresponding to PRML samples of an initial set of bits in u obtained at predetermined phases. The pair of y, u vectors exhibiting the minimum Euclidian distance yields a sampling correction value by which the initial sampling phase is corrected and a new initial sampling point preceding x is determined. Here, bit and byte synchronization have been achieved and sampling of x proceeds at the corrected phase, from the new initial sampling point.
Owner:WESTERN DIGITAL TECH INC
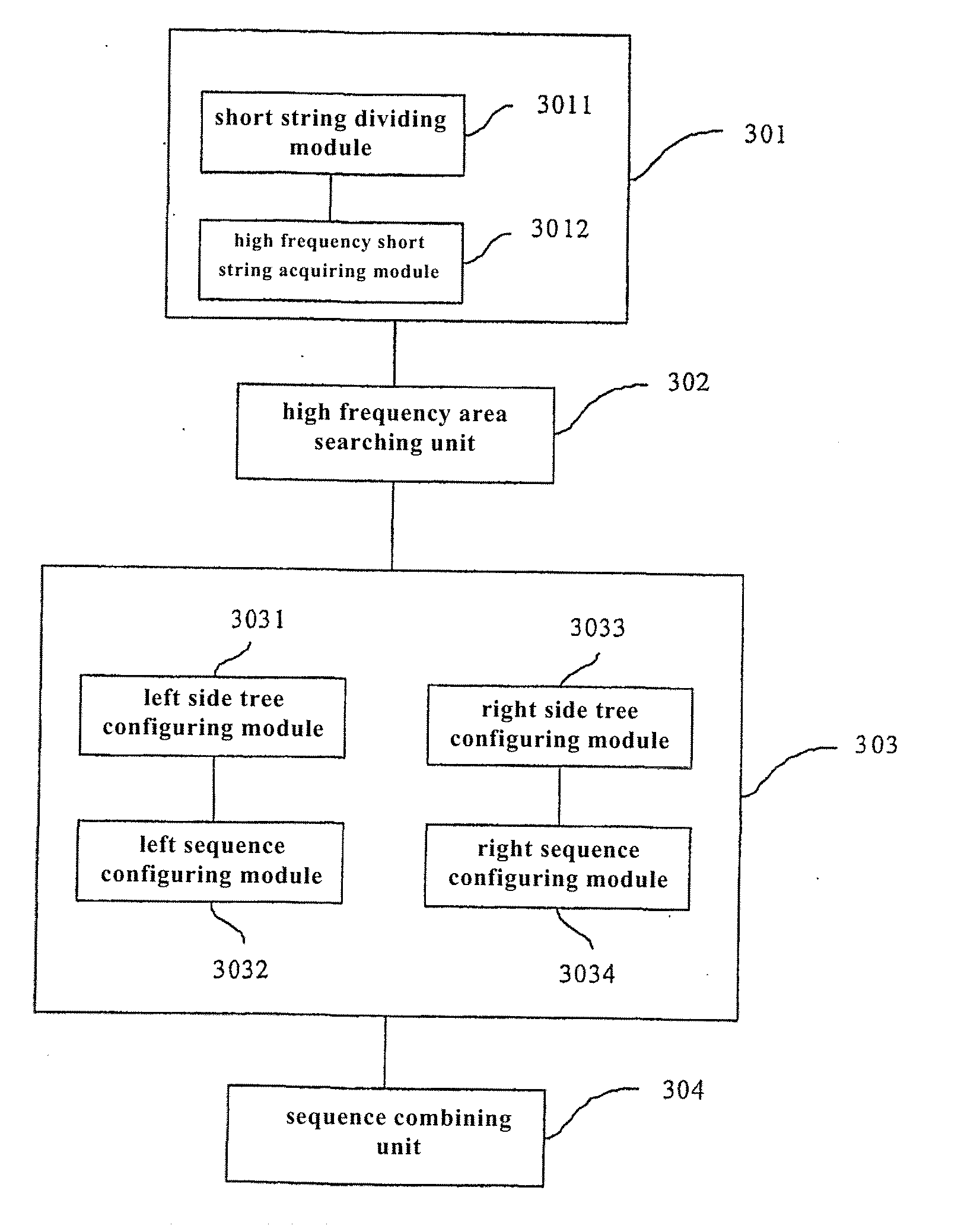
Error correcting method of test sequence, corresponding system and gene assembly equipment
ActiveUS20110295784A1Increase profitReduce memory usageMicrobiological testing/measurementGenetic modelsShort stringGene assembly
The present invention provides an error correcting method of test sequence, which involves receiving test sequences, configuring high frequency short string list based on a preset high frequency threshold value, traversing each received test sequence, searching an area with the largest number of continuous high frequency short strings on each test sequence in combination with high frequency short string list, configuring whole left sequence and / or right sequence of high frequency short strings at left side and / or right side of searched area according to corresponding received test sequence and high frequency short string list, and constituting corresponding test sequence according to configured left and / or right sequence and searched area. The present invention also provides corresponding error correcting system of test sequence and gene assembly equipment.
Owner:BGI TECH SOLUTIONS
Comparison gene sequencing data compression method and system and computer readable medium
ActiveCN110021368AIncrease the compression ratioShort compression timeCode conversionSequence analysisData compressionData stream
The invention discloses a comparison gene sequencing data compression method and system and a computer readable medium. The compression method comprises the steps that initial gene character string CS0 is selected for each read R in a gene sequencing data sample; a short string K-mer with the length being k is generated according to the sequence, the short string K-mer and a reference-based genomeare sequentially compared so as to obtain the adjacent predicting characters c of the short string K-mer in the plus strand or the minus strand of the reference-based genome, and a predicting character set PS composed of all predicting characters c is obtained; invertible computation is conducted through an invertible function after the Lr-k locus of the read R and the predicting character set PSare encoded; and the plus / minus strand type d of the read R, CS0 and the invertible computation result serve as three data streams to be compressed and output. The method has the advantages of low compression rate, short compression time and stable compression property, does not need to conduct precise comparison on the gene data, and has the high computation efficiency, and the higher the precision degree is, the lower the compression rate is.
Owner:GENETALKS BIO TECH CHANGSHA CO LTD
Features
- R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
Why Patsnap Eureka
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Social media
Patsnap Eureka Blog
Learn More Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com