Reagents, methods, and libraries for bead-based sequencing
a technology of bead-based sequencing and reagents, which is applied in the field of bead-based sequencing reagents, methods, and libraries, can solve the problems of limiting the speed and accuracy of sequencing in many contexts, requiring a time-consuming separation step, and proving difficult to identify reversible terminators that can be incorporated by polymerase with high efficiency, so as to achieve high throughput sequencing and efficient implementation of methods
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example 1
Efficient Cleavage and Ligation of Phosphorothiolated Oligonucleotides
[0347]This example describes an experiment demonstrating efficient ligation and cleavage of extension oligonucleotides containing a 3′-S phosphorothiolate linkage.
[0348]Materials and Methods
[0349]Ligation Sequencing Protocol
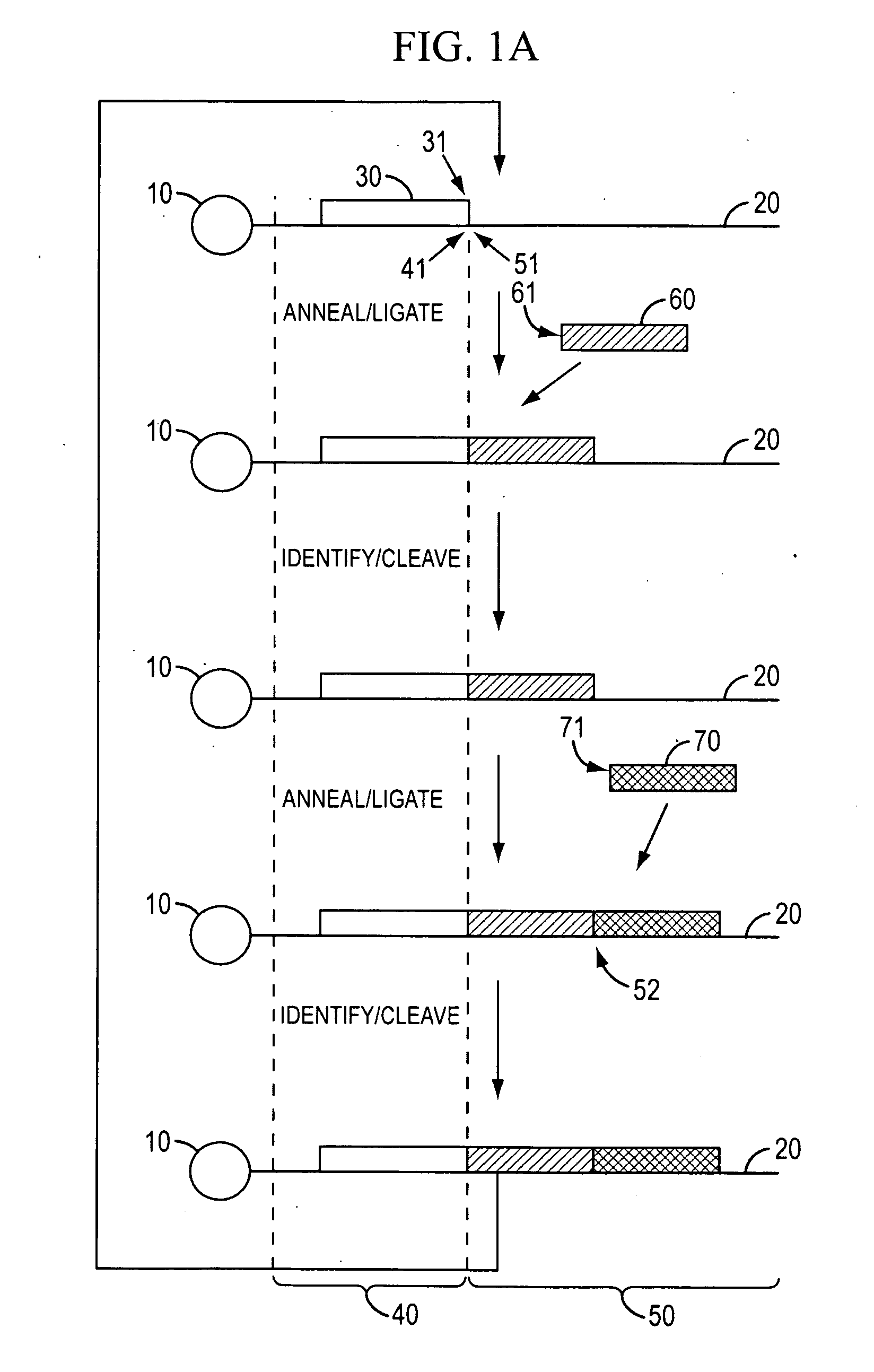
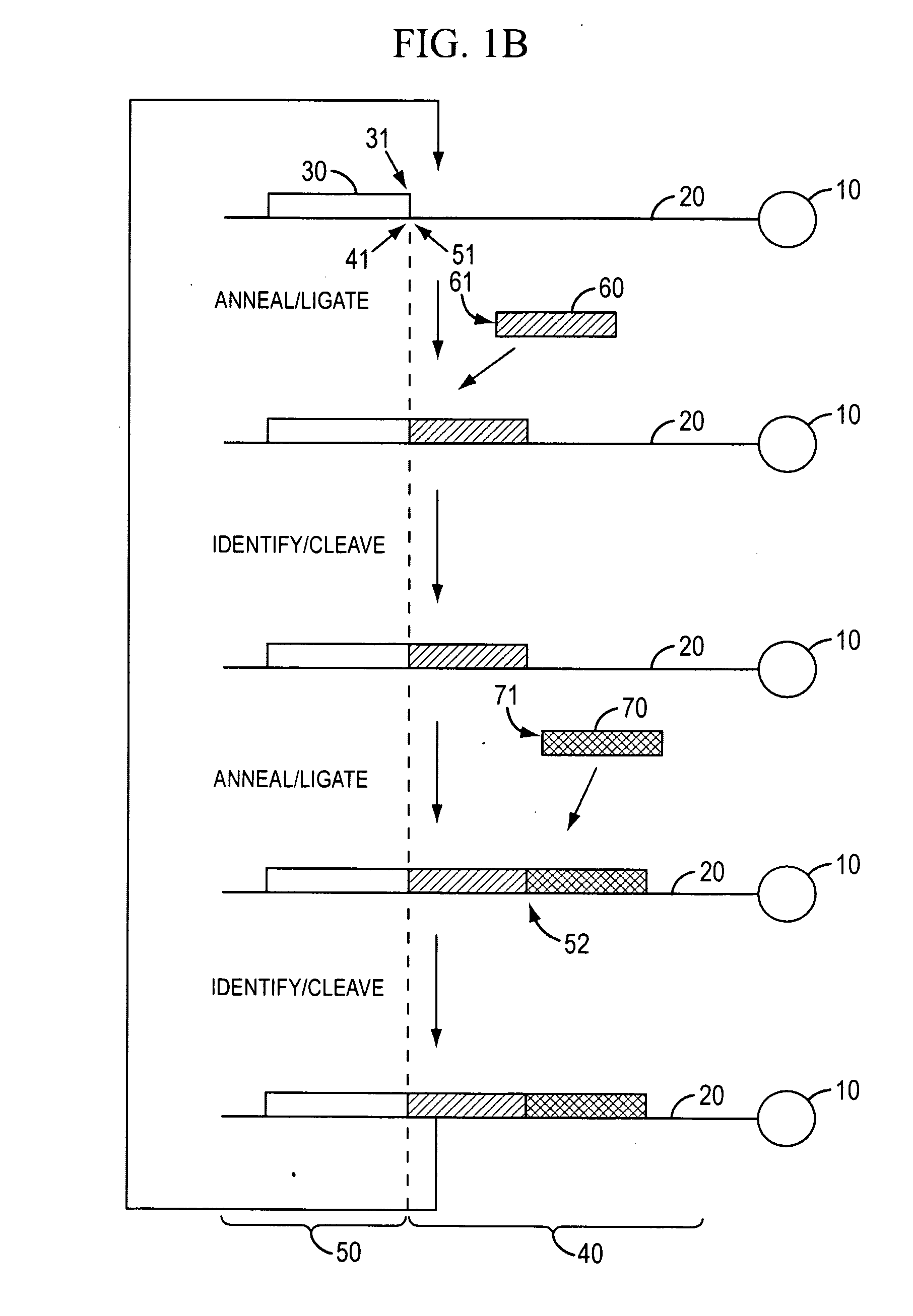
[0350]Template Preparation: To demonstrate evaluate the potential of sequencing by cycled oligonucleotide ligation and cleavage and to explore the effect of variations in certain aspects of the method, two sets of model bead-based template populations were prepared. In preferred implementations, as described in the Examples, cycled oligonucleotide ligation and cleavage extends strands in the 3′→5′ direction. Therefore, to evaluate ligation efficiencies, model templates were bound to beads at the 5′ end and designed with the same binding region at the 3′ end. One set was comprised of short (70 bp) oligonucleotides bound to streptavidin-coated magnetic beads (1 micron) via a dual biotin moiety. E...
example 2
Efficient Cleavage and Ligation of Phosphorothiolated Oligonucleotides Containing Degeneracy-Reducing Nucleotides
[0363]A competing consideration to probe length, however, is the fidelity of the extended oligonucleotide and its effect on subsequent ligation efficiency. The fidelity of T4 DNA ligase has been shown to decrease rapidly following the 5th base after the junction (Luo et al., Nucleic Acid Res., 24: 3071-3078 and 3079-3085, 1996). If mismatches are introduced at the 5′ side of a new ligation junction, the ligation efficiency may be reduced by attrition, however, no dephasing or increase in background signal will be generated (a major obstacle encountered in polymerase-based sequencing by synthesis methods).
[0364]Probe sets should preferably be capable of hybridizing to any DNA sequence in order to permit de novo sequencing of uncharacterized DNA. However, the complexity of a labeled probe set grows exponentially with the length and number of 4-fold degenerate bases. In addi...
example 3
Fidelity of Probe Ligation
[0369]Bacterial NAD-dependent ligases, such as Taq DNA ligase, have been reported to have high sequence fidelity across ligation junctions, with mismatches on the 3′ side having essentially no nick-closure activity, but mismatches on the 5′ side being tolerated to some degree (Luo et al., Nucleic Acid Res., 24: 3071-3078 and 3079-3085, 1996). T4 DNA ligase, on the other hand, has been reported to be somewhat less stringent, allowing mismatches on both the 3′- and 5′-sides of the junction. It was therefore of interest to evaluate the fidelity of probe ligation with T4 DNA ligase in comparison to Taq DNA ligase in the context of our system.
[0370]We developed two methods to evaluate the sequence fidelity of ligated oligonucleotides using standard ABI sequencing technology. The first method was designed to clone and sequence ligation products. In this method, ligation extension products were attached to adapter sequences, cloned and transformed into bacteria. I...
PUM
| Property | Measurement | Unit |
|---|---|---|
| diameter | aaaaa | aaaaa |
| diameter | aaaaa | aaaaa |
| diameters | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More - R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com