Synthetic antibody phage libraries
a technology of synthetic antibodies and phages, applied in the field of synthesized antibodies phage libraries, can solve the problems of inefficiency of production, reduced library size, and inability to apply in systematic and quantitative manner, and achieve high quality, high-quality target binding characteristics, and efficient generation.
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example 2
Library Design: L1, L2, L3, H1, H2
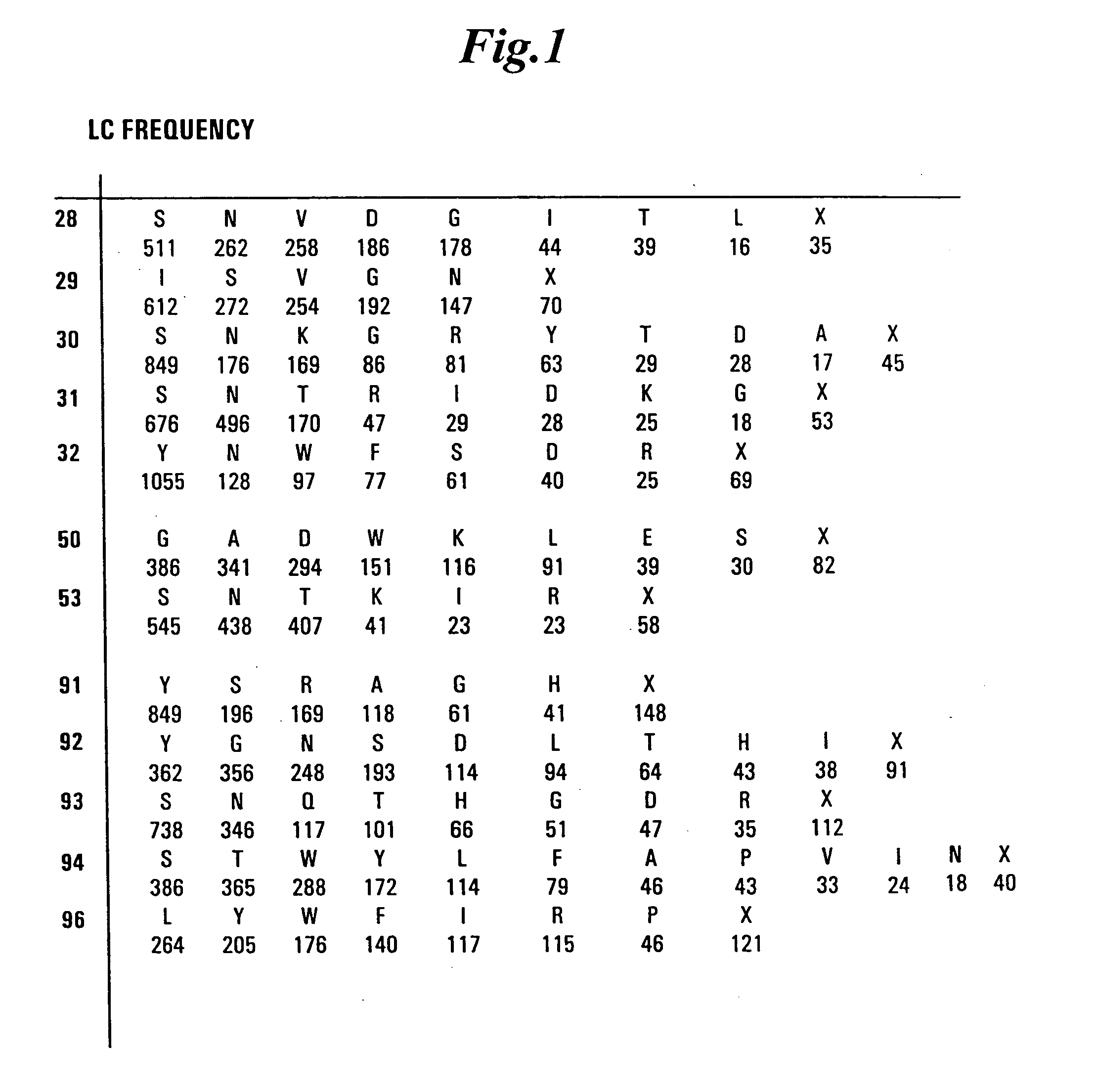
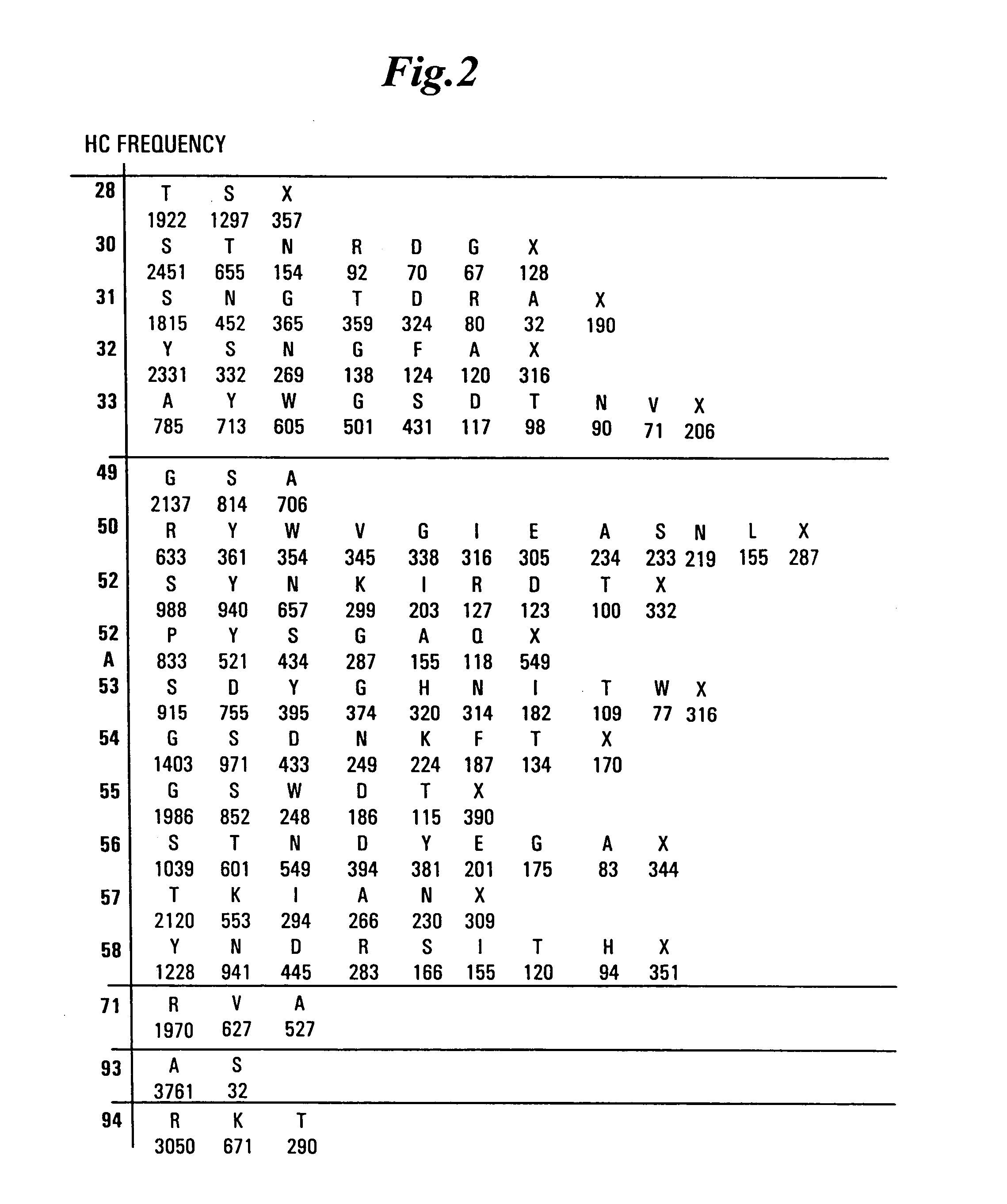
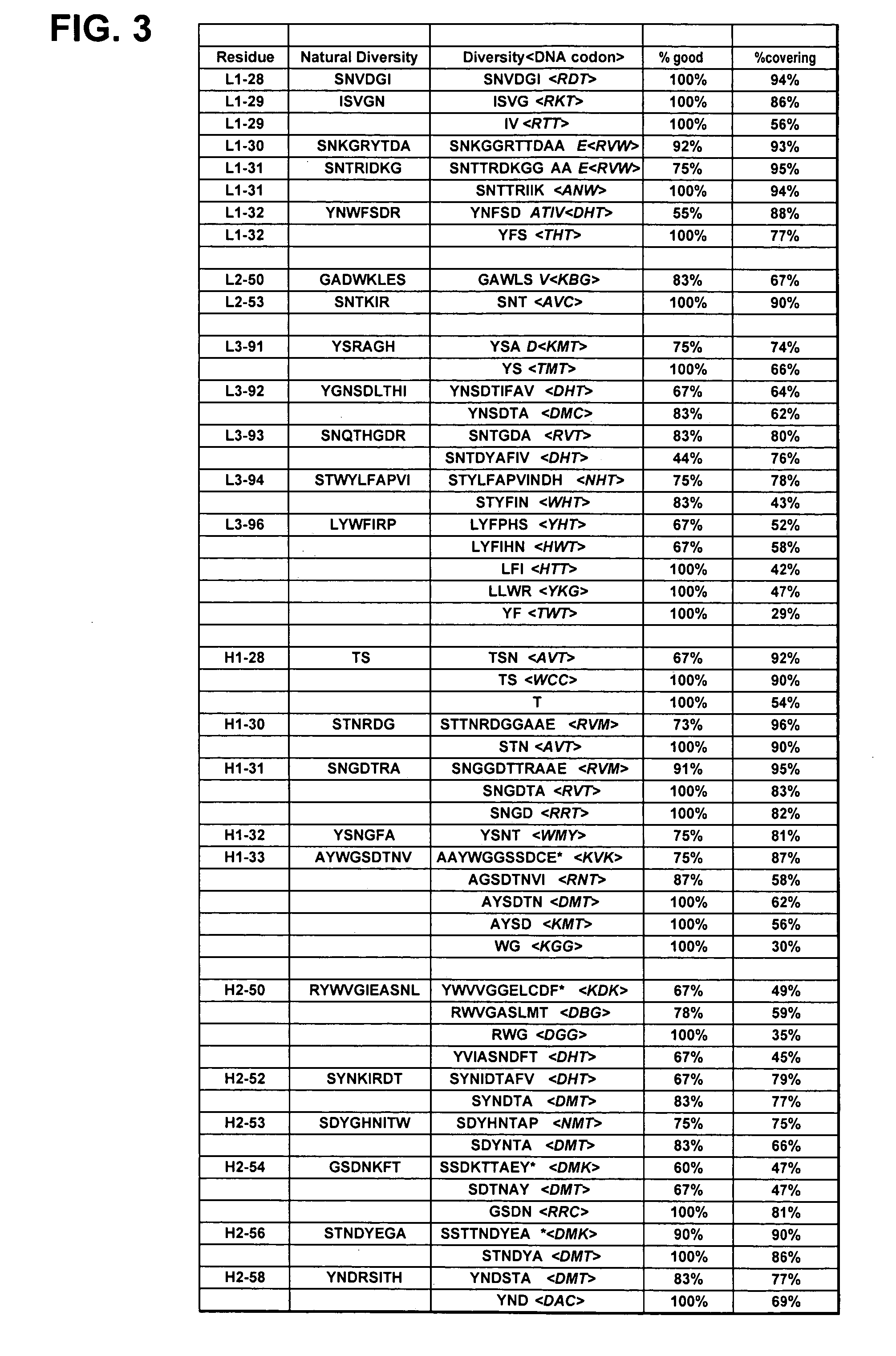
[0479] Libraries of antibody variable domains were designed to maximize diversity in the CDR regions while minimizing structural perturbations in the antibody variable domains. Structural perturbations in antibody variable domains are generally associated with improperly folded antibody domains resulting in low yield, for example when produced in bacterial cells. Low yields decrease the number of binders detected in screening. Diversity in the CDR regions was generated by identifying solvent accessible and highly diverse positions in each CDR for CDRs L1, L2, L3, H1 and H2, and designing an oligonucleotide comprising at least one tailored (i.e., non-random) codon set encoding variant amino acids for the amino acid position corresponding to the position of at least one solvent accessible residue at a highly diverse position in at least one CDR region. A tailored codon set is a degenerate nucleic acid sequence that preferably encodes the most commonly...
example 3
Library Design: H3
[0490] In comparison to other CDRs, heavy chain CDR3 (H3) regions exhibit the greatest diversity in sequences and lengths, although the sequence diversity is not completely random (i.e., some amino acids were found to occur more often than other in particular amino acid positions).
[0491] In a preliminary analysis to assess the amino acid preferences for each position in H3, a library with diverse H3 was generated using an NNK codon set for residues 95-100a of the humanized antibody 4D5 H3 region. The NNK codon set encodes all 20 amino acids and stop codons. This library was generated in a Fab phage display format and 400 clones that displayed functionally on phage were identified and sequenced. The amino acid sequence found in H3 regions in the NNK library were compared to those found in the Kabat database. A comparison of those amino acids is shown in FIG. 23. When the amino acid sequences in the NNK library and Kabat database were analyzed, it was determined ther...
example 4
Construction, Sorting and Analysis of scFv Libraries
[0498] Libraries with diversified CDRs were generated using vectors comprising 4D5 variable domains in the scFv or scFv-zip formats as described in Example 1. In total, five libraries were generated and the library characteristics were as follows:
3 Library name Format CDR Diversity ScFv-1 zipper H1, H2, H3 ScFv-2 Zipper H1, H2, H3, L3 ScFv-3 Zipper H3, L3 ScFv-4 No zipper H1, H2, H3 ScFv-5 No zipper H1, H2, H3, L3
[0499] Libraries were constructed using the method of Kunkel (Kunkel et al., Methods Enzymol. (1987), 154, 367-382) with previously described methods (Sidhu et al., Methods Enzymol.(2000), 328, 333-363). For each library a "stop template" version of a scFv or scFv-zip display vector was used; in each case, a stop template with TAA stop codons within each of the CDRs to be randomized was used. Mutagenic oligonucleotides with degenerate codons at the positions to be diversified were used to simultaneously introduce CDR diver...
PUM
| Property | Measurement | Unit |
|---|---|---|
| size | aaaaa | aaaaa |
| pH | aaaaa | aaaaa |
| temperature | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More - R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com